the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Impact of dust addition on the microbial food web under present and future conditions of pH and temperature
Estelle Bigeard
Frédéric Gazeau
Farooq Azam
Cécile Guieu
Emilio Marañón
Céline Ridame
France Van Wambeke
Ingrid Obernosterer
Anne-Claire Baudoux
In the oligotrophic waters of the Mediterranean Sea, during the stratification period, the microbial loop relies on pulsed inputs of nutrients through the atmospheric deposition of aerosols from both natural (e.g., Saharan dust), anthropogenic, or mixed origins. While the influence of dust deposition on microbial processes and community composition is still not fully constrained, the extent to which future environmental conditions will affect dust inputs and the microbial response is not known. The impact of atmospheric wet dust deposition was studied both under present and future environmental conditions (+3 ∘C warming and acidification of −0.3 pH units), through experiments in 300 L climate reactors. In total, three Saharan dust addition experiments were performed with surface seawater collected from the Tyrrhenian Sea, Ionian Sea, and Algerian basin in the western Mediterranean Sea during the PEACETIME (ProcEss studies at the Air–sEa Interface after dust deposition in the MEditerranean sea) cruise in May–June 2017. Top-down controls on bacteria, viral processes, and community, as well as microbial community structure (16S and 18S rDNA amplicon sequencing), were followed over the 3–4 d experiments. Different microbial and viral responses to dust were observed rapidly after addition and were, most of the time, more pronounced when combined with future environmental conditions. The dust input of nutrients and trace metals changed the microbial ecosystem from a bottom-up limited to a top-down controlled bacterial community, likely from grazing and induced lysogeny. The relative abundance of mixotrophic microeukaryotes and phototrophic prokaryotes also increased. Overall, these results suggest that the effect of dust deposition on the microbial loop is dependent on the initial microbial assemblage and metabolic state of the tested water and that predicted warming and acidification will intensify these responses, affecting food web processes and biogeochemical cycles.
Please read the corrigendum first before continuing.
-
Notice on corrigendum
The requested paper has a corresponding corrigendum published. Please read the corrigendum first before downloading the article.
-
Article
(978 KB)
- Corrigendum
-
Supplement
(1540 KB)
-
The requested paper has a corresponding corrigendum published. Please read the corrigendum first before downloading the article.
- Article
(978 KB) - Full-text XML
- Corrigendum
-
Supplement
(1540 KB) - BibTeX
- EndNote
Input of essential nutrients and trace metals through aerosol deposition is crucial to the ocean surface water biogeochemistry and productivity (at the global scale, e.g., Mahowald et al., 2017 and in the Mediterranean Sea, e.g., Guieu and Ridame, 2020), with episodic fertilization events driving microbial processes in oligotrophic regions such as the Pacific Ocean, the Southern Ocean, and the Mediterranean Sea.
The summer Mediterranean food web is characterized by low primary production (PP) and heterotrophic prokaryotic production (more classically abbreviated as BP for bacterial production) constrained by nutrient availability. Low BP further limits dissolved organic matter (DOM) utilization and export, resulting in DOM accumulation. Therefore, inputs of bioavailable nutrients through the deposition of atmospheric particles are essential to the Mediterranean Sea microbial ecosystem. Indeed, these nutrient pulses have been shown to support microbial processes, but the extent to which the microbial food web is affected might be dependent on the degree of the oligotrophy of the water (Marín-Beltrán et al., 2019; Marañon et al., 2010).
In the Mediterranean Sea, dust deposition may stimulate PP and N2 fixation (Guieu et al., 2014; Ridame et al., 2011, 2022) but also BP, bacterial respiration, virus production, and grazing activities and can alter the composition of the microbial community (e.g., Pulido-Villena et al., 2014; Tsiola et al., 2017; Guo et al., 2016; Pitta et al., 2017; Marín-Beltrán et al., 2019). Overall, in such an oligotrophic system, dust deposition appears to predominantly promote heterotrophic activity, which will increase respiration rates and CO2 release.
Anthropogenic CO2 emissions are projected to induce an increase in seawater temperature and an accumulation of CO2 in the ocean, leading to its acidification and an alteration of ocean carbonate chemistry (IPCC, 2014). In response to ocean warming and increased stratification, low-nutrient–low-chlorophyll (LNLC) regions, such as the Mediterranean Sea, are projected to expand in the future (Durrieu de Madron et al., 2011). Moreover, dust deposition is also expected to increase due to desertification (Moulin and Chiapello, 2006). For these reasons, in the future ocean, the microbial food web might become even more dependent on the atmospheric deposition of nutrients. Expected increased temperature and acidification might have complex effects on the microbial loop by modifying the microbial and viral and community (e.g., Highfield et al., 2017; Krause et al., 2012; Hu et al., 2021; Allen et al., 2020; Malits et al., 2021). While increasing temperature in combination with nutrient input might enhance heterotrophic bacterial growth (Degerman et al., 2012; Morán et al., 2020) more than PP (Marañón et al., 2018), future environmental conditions could push this microbial community even further towards heterotrophy. But, so far, the role of dust on the microbial food web in future climate scenarios is unknown.
Here, we studied the response of Mediterranean microbial and viral communities (i.e., viral strategies, microbial growth, and controls, as well as community composition) to simulated wet Saharan dust deposition during on-board minicosm experiments conducted in three different basins of the western and central Mediterranean Sea under present and future projected conditions of temperature and pH. To our knowledge, this is the first study assessing the effect of atmospheric deposition on the microbial food web under future environmental conditions.
2.1 Experimental setup
During the ProcEss studies at the Air–sEa Interface after dust deposition in the MEditerranean sea project cruise (PEACETIME), on board the research vessel (R/V) Pourquoi Pas? in May/June 2017, three experiments were conducted in 300 L climate reactors (minicosms) filled with surface seawater collected at three different stations (Table 1) in the Tyrrhenian Sea (TYR), Ionian Sea (ION), and in the Algerian basin (FAST). The experimental setup is described in detail in Gazeau et al. (2021a). Briefly, the experiments were conducted for 3 d (TYR and ION) and 4 d (FAST) in trace-metal-free conditions, under light-, temperature-, and pH-controlled conditions, following ambient or future projected conditions of temperature and pH. For each experiment, the biogeochemical evolution of the water, after dust deposition, under present and future environmental conditions was followed in three duplicate treatments: (i) CONTROL (C1 and C2) with no dust addition and under present pH and temperature conditions, (ii) DUST (D1 and D2) with dust addition under present environmental conditions, and (iii) GREENHOUSE (G1 and G2) with dust addition under projected temperature and pH for 2100 (IPCC, 2014; ca. +3 ∘C and −0.3 pH units). Water was acidified by addition of CO2-saturated 0.2 µm filtered seawater and slowly warmed overnight (Gazeau et al., 2021a). The same dust analog was used as during the DUNE 2009 experiments, as described in Desboeufs et al. (2014), and the same dust wet flux of 10 g m−2 was simulated (as described in Gazeau et al., 2021a). Briefly, the dust was derived from the < 20 µm fraction of soil collected in southern Tunisia (a major source for material transported and deposited in the northwestern Mediterranean), with most particles (99 %) smaller than 0.1 µm (Desboeufs et al., 2014). The collected material underwent an artificial chemical aging process by the addition of nitric and sulfuric acid (HNO3 and H2SO4, respectively) to mimic cloud processes during atmospheric transport of aerosol with anthropogenic acid gases (Guieu et al., 2010, and references therein). To mimic a realistic wet flux event for the Mediterranean of 10 g m−2, 3.6 g of this analog dust were quickly diluted in 2 L ultrahigh purity water and sprayed at the surface of the dust amended treatments (D1 and D2; G1 and G2; Gazeau et al., 2021a). Such a deposition event represents a high but realistic scenario, as several studies reported even higher short wet deposition events in this area of the Mediterranean Sea (Ternon et al., 2010; Bonnet and Guieu, 2006; Loÿe-Pilot and Martin, 1996), suggesting that wet deposition is the main pathway of dust input in the western Mediterranean Sea.
Samples for all parameters (except described below) were taken at t12h (while filling the tanks), t0 (just before dust addition), t1h, t6h, t12h, t24h, t48h, t72h, and t96h (after dust addition and t96h only for FAST).
Table 1Initial conditions (t12h) at the three stations sampled for the dust addition experiments. Other parameters are presented in more detail in Gazeau et al. (2021a, b).
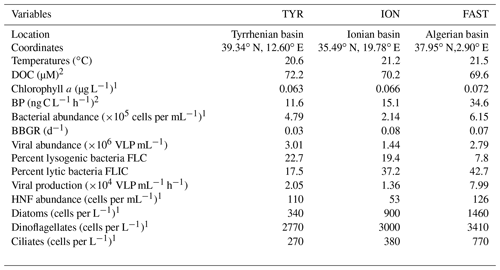
DOC – dissolved organic carbon; BP – heterotrophic prokaryotic production; BBGR – bacterial-biomass-specific growth rate; HNF – heterotrophic nanoflagellate. 1 Results presented in Gazeau et al. (2021a). 2 Results presented in Gazeau et al. (2021b).
2.2 Growth rates, mortality, and top-down controls
BP was estimated at all sampling points from rates of 3H-leucine incorporation (Kirchman et al., 1985; Smith and Azam, 1992), as described in Gazeau et al. (2021b). Briefly, triplicate 1.5 mL samples and one blank were incubated in the dark for 1–2 h after the addition of 20 nM of a mix of cold and 3H-leucine in two temperature-controlled incubators maintained, respectively, at ambient temperature for C1, C2, D1, and D2 and at ambient temperature +3 ∘C for G1 and G2. Heterotrophic prokaryotes (HB) and heterotrophic nanoflagellate (HNF) abundances were measured by flow cytometry, as described in Gazeau et al. (2021a). Briefly, samples (4.5 mL) were fixed with glutaraldehyde grade I (1 % final concentration) and stored at −80 ∘C until analysis. Counts were performed on a FACSCanto II flow cytometer (Becton, Dickinson, and Company). Cells were stained with SYBR Green I at 0.025 % (vol vol) final concentration (Gasol and DelGiorgio, 2000; Christaki et al., 2011). Bacterial biomass specific growth rates (BBGRs) were estimated following Kirchman (2002), BP/bacterial biomass, assuming a carbon-to-cell ratio of 20 fg C per cell (Lee and Fuhrman, 1987). Mortality was estimated as being the difference between HB present between two successive sampling points and those produced during that time.
2.3 Viral abundance, production, and life strategy
Virus abundances were determined on glutaraldehyde fixed samples (0.5 % final concentration; grade II; Sigma-Aldrich, St Louis, MO, USA) stored at −80 ∘C until analysis. Flow cytometry analysis was performed as described by Brussaard (2004). Briefly, samples were thawed at 37 ∘C, diluted in 0.2 µm filtered autoclaved TE buffer (10:1 Tris-EDTA; pH 8) and stained with SYBR Green I ( of the commercial stock; Life Technologies, Saint Aubin, France) for 10 min at 80 ∘C. Virus particles were discriminated based on their green fluorescence and suspended sediment concentration (SSC) during 1 min analyses (Fig. S1). All cytogram analyses were performed with the Flowing Software freeware (Turku Center for Biotechnology, Finland).
Viral production and bacterial losses due to phages were assessed by the virus reduction approach (Weinbauer et al., 2010) at t0 and t24h in all six minicosms. Briefly, 3 L of seawater were filtered through a 1.2 µm pore size polycarbonate filter (Whatman©), and HB (filtrate) were concentrated by ultrafiltration (0.22 µm pore size; Vivaflow 200© polyethersulfone – PES) down to a volume of 50 mL. Virus-free water was obtained by filtering 1 L of seawater through a 30 kDa pore size cartridge (Vivaflow 200©; PES). In total, six mixtures of HB concentrate (2 mL) diluted in virus-free water (23 mL) were prepared and distributed into 50 mL Falcon tubes. Then, three of the tubes were incubated as controls, while the other three were inoculated with mitomycin C (Sigma-Aldrich; 1 µg mL−1 final concentration) as an inducing agent of the lytic cycle in lysogenic bacteria. All tubes were incubated in darkness in two temperature-controlled incubators maintained, respectively, at an ambient temperature for C1, C2, D1, and D2 and at an ambient temperature +3 ∘C for G1 and G2. Samples for HB and viral abundances were collected every 6 h for a total incubation period of 18 h.
The estimation of virus-mediated mortality of HB was performed according to Weinbauer et al. (2002) and Winter et al. (2004). Briefly, an increase in virus abundance in the control tubes represents lytic viral production (VPL), and an increase in treatments with mitomycin C represents total viral production (VPT), i.e., lytic plus lysogenic viral production. The difference between VPT and VPL represents lysogenic production (VPLG). The frequency of lytically infected cells (FLIC) and the frequency of lysogenic cells (FLC) was calculated as follows:
where HBi is the initial HB abundance in the viral production experiment, and BS is a theoretical burst size of 20 viruses per infected cell (averaged BS in marine oligotrophic waters; Parada et al., 2006).
2.4 DNA sampling, sequencing, and sequence analysis
To study the temporal dynamics of the microbial diversity, water samples (3 L) were collected in acid-washed containers from each minicosm at t0, t24h, and at the end of the experiments (t72h at TYR and ION and t96h at FAST). Samples were filtered onto 0.2 µm PES filters (Sterivex©) and stored at −80 ∘C until DNA extraction. Nucleic acids were extracted from the filters using a phenol–chloroform method, and DNA was then purified using filter columns from NucleoSpin® Plant II kit (Macherey-Nagel GmbH & Co. KG ) following a modified protocol. DNA extracts were quantified and normalized at 5 ng µL−1 and used as templates for the polymerase chain reaction (PCR) amplification of the V4 region of the 18S rRNA (∼ 380 bp), using the primers TAReuk454FWD1 and TAReukREV3 (Stoeck et al., 2010), and the V4–V5 region of the 16S rRNA (∼ 411 bp), using the primers 515F-Y (5′-GTGYCAGCMGCCGCGGTAA) and 926R-R (5′-CCGYCAATTYMTTTRAGTTT; Parada et al., 2016). Following PCRs, DNA amplicons were purified, quantified and sent to Genotoul (Toulouse, France) for high throughput sequencing using paired-end 2×250 bp Illumina MiSeq. Note that although we used universal primer, archaea were mostly not detected, and the prokaryotic heterotrophic communities corresponded essentially to eubacteria; therefore, the taxonomic description referred to the general term bacterial communities.
All reads were processed using the Quantitative Insight Into Microbial Ecology 2 pipeline (QIIME 2 v2020.2; Bolyen et al., 2019). Reads were truncated at 350 bp based on sequencing quality, denoised, merged, and chimera checked using DADA2 software (Callahan et al., 2016). A total of 714 and 3070 amplicon sequence variants (ASVs) were obtained for 16S and 18S, respectively. Taxonomy assignments were made against the database SILVA 132 (Quast et al., 2013) for 16S and PR2 (Guillou et al., 2013) for 18S. All sequences associated with this study have been deposited under the BioProject ID no. PRJNA693966.
2.5 Statistics
Alpha and beta diversity indices for community composition were estimated after randomized subsampling to 26 000 reads for 16S rDNA and 19 000 reads for 18S rDNA. Analyses were run in QIIME 2 and in the Primer v.6 software package (Clarke and Warwick, 2001). Differences between the samples' richness and diversity were assessed using Kruskal–Wallis pairwise test. Beta diversity was run on the Bray–Curtis dissimilarity. Differences between the samples' beta diversity were tested using PERMANOVA (permutational multivariate analysis of variance) with a pairwise test and 999 permutations. The sequences contributing most to the dissimilarity between clusters were identified using SIMPER (similarity percentage). A linear mixed model was performed with R software (R Core Team, 2020), using the nlme package (Pinheiro et al., 2014) to test if the amended treatments differed from the controls at t24h and t72h or t96h.
3.1 Microbial growth, mortality, and top-down controls
Nutrients inputs were observed with dust addition (Fig. S2); in response, the autotrophic and heterotrophic microbial abundances and BP increased (Fig. S3), as described in more detail in Gazeau et al. (2021a, b). Already in the 24 h following the dust addition, significant increases in heterotrophic bacterial biomass specific growth rates (BBGRs; p≤0.016 at t24h) were observed in all experiments with dust under D and G, as seen in Figs. 1 (showing data normalized to C) and S4. The highest growth rates were already observed 24 h after dust seeding (up to 2.9 d−1 in G2 at FAST; Table S1; Fig. S4). At 24 h, in both D and G, the heterotrophic bacterial mortality rates were higher than in C (Fig. 1), especially at TYR in D (up to 0.5 d−1) and in G at ION (up to 0.6 d−1) and FAST (up to 0.7 d−1; Table S1). Over the course of the three experiments, the slope of the linear regression between log bacterial biomass and log bacterial production was below 0.4 in the three treatments, suggesting a weak bottom-up control (Fig. 2a; Ducklow, 1992). The slope decreased in D and G relative to C. Overall, the top-down index, as described by Morán et al. (2017), was higher in G (0.92) relative to C and D (0.80). The relationship between log-transformed HNF and log bacterial abundance (Fig. 3b), plotted according to the model in Gasol (1994), showed that HNF were below the MRA (mean realized HNF abundance) in all treatments, suggesting a top-down control of HNF abundance. HNF and bacteria were weakly coupled in all treatments. The relationship between total viruses and bacterial abundance was weaker in D and G relative to C (Fig. S5).
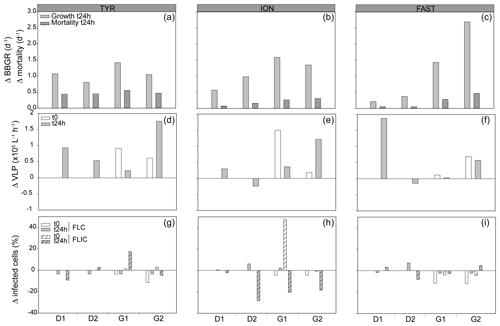
Figure 1Bacterial and viral parameters in the three experiments (TYR, ION, and FAST) in each minicosm (D1, D2, G1, and G2). The values are normalized to the controls, and the data are presented as the difference between the treatments and the mean value of the duplicate controls. The first row represents the bacterial biomass specific growth rates (BBGRs) and relative mortality rates at t24h after dust addition. The second raw represents the relative viral productions at t24h and at t0 for the G treatments. The last row represents the viral strategies, with the percentages of lytic (FLIC) or lysogenic (FLC) cells at t24h and at t0 for the G treatments.
3.2 Viral dynamics and processes
The initial abundance and production of virus-like particles (VLPs) was higher in the western stations (Table 1). Viral strategy (lysogenic vs. lytic replication) was also different between stations, with a higher frequency of lysogenic cells (FLC) at TYR and ION (23 % and 19 %, respectively; Table 1) and a higher frequency of lytically infected cells (FLIC) at FAST (43 %, Table 1).
During TYR and ION experiments, the relative contribution of VLP populations was similar and stable over time, with low DNA viruses representing over 80 % of the community (Figs. 3 and S5). The low DNA VLP abundance was, however, slightly higher in D and G relative to C after 24 h at TYR and significantly higher at ION after 48 h (p=0.037; Fig. 4). In contrast to the other two stations, at FAST, giruses (giant viruses, characterized by high DNA fluorescence and high SSC) were also present and increased in all treatments but especially in G, where they made up to 9 % of the viral community at the end of the experiment (Figs. 3 and 4). The abundance of high DNA viruses at FAST also increased independently of the treatments and accounted for 16 %–18 % of the community at the end of the experiment (Figs. 3 and 4).
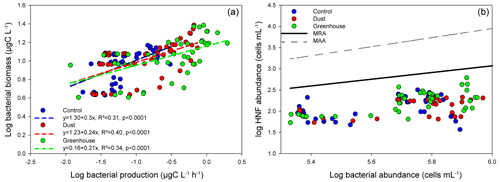
Figure 2(a) Log–log linear regression between bacterial biomass and bacterial production. Dotted lines represent the linear regressions for each treatment. (b) Relationships between log HNF abundance and log bacterial prey abundance. Solid black and dotted black lines correspond to the mean realized HNF abundance (MRA) and theoretical maximum attainable HNF abundance line (MAA), respectively. The samples are grouped per treatment.
The sampling strategy for the production and life strategies of HB viruses allowed us to independently discriminate the effect of (i) greenhouse conditions (sampling at t0 before dust addition), (ii) dust addition (sampling at t24h), and the combined effects of dust addition and greenhouse. Lytic viral production (VPL) increased significantly at t0 in G at TYR and ION compared to C (p≤0.036). The addition of dust induced higher VPL in D at TYR (normalized to C; Fig. 1). No significant impact of dust on VPL was observed in G compared to D after 24 h for any of the experiments. Changes in the viral infection strategy were observed with G conditions at t0, where FLC decreased relative to the non-G treatments at TYR and ION and especially at FAST (Fig. 1; p=0.047). FLIC increased slightly in G at TYR and ION already at t0. Dust addition had no detectable significant effect on this parameter for any experiment. Looking at the relative share between lytic and lysogenic infection, dust addition favored lytic infection at TYR (no lysogenic bacteria were observed after 24 h), but the contribution of both infection strategies remained unchanged compared to C at ION and FAST. Greenhouse conditions also favored replication through lytic cycle already at t0 for all three experiments, and this trend was not impacted by dust addition.
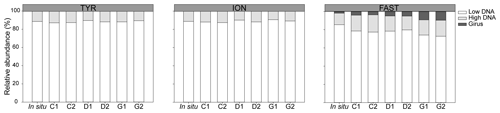
Figure 3Relative abundance of viral populations at the initial (in situ at t12h before dust addition) and final time points in all minicosms (C1, C2, D1, D2, G1, and G2) during the three experiments (TYR, ION, and FAST).
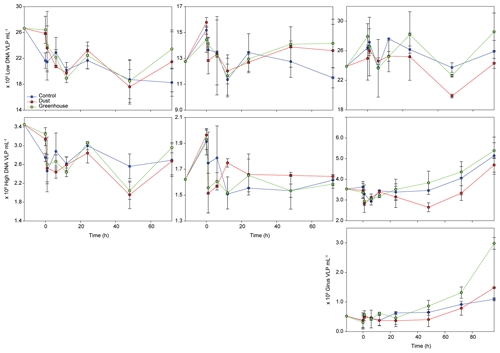
Figure 4Evolution of virus-like particle (VLP) abundances of three different viral populations over the course of the three experiments (TYR, ION, and FAST). The first row represents low DNA viruses or phages, the second row represents high DNA viruses, and the third row represents giant viruses (giruses).
3.3 Microbial community composition
Microbial community structure, bacteria, and microeukaryotes from 16S rDNA and 18S rDNA sequencing, respectively, responded to dust addition in all three experiments relative to C (Figs. 5 and 6). After quality controls, reads were assigned to 714 and 1443 ASVs for 16S and 18S, respectively.
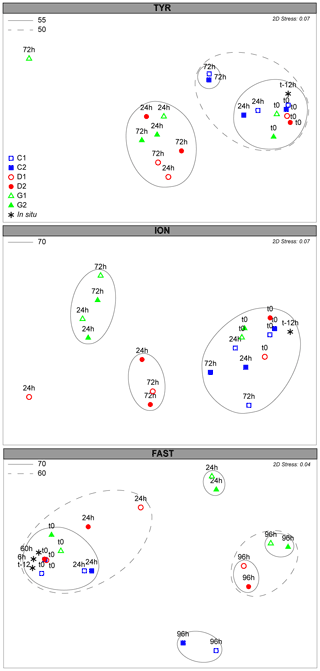
Figure 5Non-metric dimensional scaling (NMDS) plot of bacterial community composition over the course of the three experiments based on Bray–Curtis dissimilarities of 16S rDNA sequences. Samples clustering at different levels of similarity are circled together. All circles represent clusters which are significantly different from each other (p<0.05) based on a PERMANOVA test.
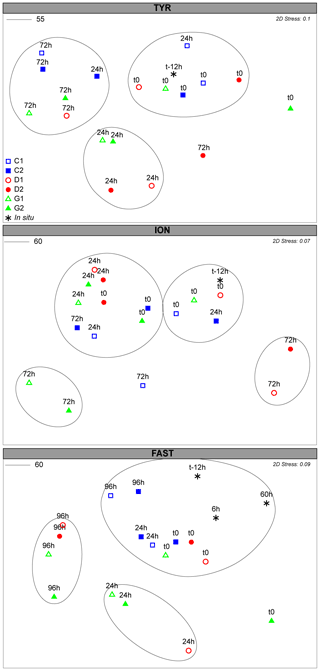
Figure 6NMDS plot of the microeukaryotes community composition over the course of the three experiments based on Bray–Curtis dissimilarities of 18S rDNA sequences. Samples clustering at different levels of similarity are circled together. All circles represent clusters which are significantly different (p<0.05) from each other based on a PERMANOVA test.
3.3.1 Bacterial community composition
The initial community composition (t12h) was significantly different at the three stations (PERMANOVA; p=0.001; Figs. S6a, S7). Rapid and significant changes in the bacterial community composition were already observed 24 h after dust addition (Fig. 5). Despite the initial different communities, the three stations appeared to converge towards a closer community composition in response to dust addition (Fig. S7). At TYR, communities in D and G significantly changed 24 h after dust addition (PERMANOVA; p=0.001). This cluster presented no significant differences between treatments (D and G) or time (24 and 72 h). The differences between C and D/G were attributed to a relative increase in ASVs related to different Alteromonas sp., OM60, and Pseudophaeobacter sp. and Erythrobacter sp., and the contribution of ASVs related to SAR11 and Verrucomicrobia and Synechococcus decreased (Table S2a). At ION, the bacterial community composition significantly changed 24 h after dust addition (PERMANOVA; p=0.001) and was significantly different between D and G (PERMANOVA; p=0.032). As observed at TYR, no further change occurred between 24 h and the end of the experiment (72 h; Fig. 5). The difference between the controls and dust-amended minicosms were assigned to an increase in ASVs related to different Alteromonas sp., Erythrobacter sp., Dokdonia sp., and OM60 and a decrease in ASVs related to SAR11, Synechococcus, Verrucomicrobia, Rhodospirillales, and some Flavobacteria (Table S2b). Several ASVs related to Alteromonas sp., Synechococcus sp., and Erythrobacter sp. were further enriched in G compared to D, while Dokdonia sp. was mainly present in D. At FAST, the bacterial community, after 24 h, only significantly changed in G (PERMANOVA; p=0.011; Fig. 5). However, after 96 h, the community in D and G were similar and appeared to transition back to the initial state at 96 h (PERMANOVA; p=0.077). The higher relative abundance in Erythrobacter sp., Synechococcus sp., different ASVs related to Alteromonas sp., and Flavobacteria appeared to contribute mainly to the difference between C and D/G (Table S2), while ASVs related to SAR11, Verrucomicrobia, Celeribacter sp., Thalassobius sp., and Rhodospirillales were mainly present in C (Table S2c).
3.3.2 Nano- and microeukaryotes community composition
The diversity of the initial community was large (Fig. S7) and significantly different at the three stations (PERMANOVA; p=0.001; Fig. S6b). At TYR, the nano- and microeukaryotes community responded rapidly (24 h) to the dust addition (PERMANOVA; p=0.003). This initial high diversity disappeared after 72 h, with similar communities in all minicosms (Fig. S7). They were significantly different from initial and t24h communities (p=0.002 and 0.03, respectively; Fig. 6) in D/G. The variations at t24h were attributed to changes in the dinoflagellate communities and, in particular, to an increase in ASVs related to Heterocapsa rotundata, Gymnodiniales and Gonyaulacales, as well as to an increase in Chlorophyta (Table S3a). At ION, no significant changes were observed between C and D/G after 24 h. However, after 72 h, the communities were significantly different in D (p=0.018) and G (p=0.05) compared to the communities at t24h in these treatments (Table S3b). In D, diversity was significantly higher at t72 h compared to t24 h and to C at the same sampling time (p=0.036). In contrast, diversity in G at t72h was lower than at t24h and lower to the one observed in C at the same sampling time (p=0.066; Fig. S8). These differences were mainly attributed to changes in ASVs related to dinoflagellates and to the increase at t72h in Emiliania huxleyi and Chlorophyta in D and G, respectively (Table S3b). At FAST, significant differences were observed between the controls and initial communities compared to the dust-amended (D and G) treatments at t24h (p=0.036). No major differences were observed between D/G at t24h and t96h (p=0.06). The differences were mainly attributed to changes in dinoflagellates ASVs and to an increase in Acantharea and Emiliania huxleyi in D and G treatments at t96h (Table S3c).
Pulsed inputs of essential nutrients and trace metals through aerosol deposition are crucial to surface microbial communities in LNLC regions such as the Mediterranean Sea (reviewed in Guieu and Ridame, 2020). Here we assessed the impact of dust deposition on the late spring microbial loop under present and future environmental conditions on the surface water of three different Mediterranean basins (Tyrrhenian – TYR; Ionian – ION; and Algerian – FAST). The initial conditions at the three sampled stations for the on-board experiments are described in more detail in Gazeau et al. (2021a). Briefly, very low levels of dissolved inorganic nutrients were measured at all three stations, highlighting the oligotrophic status of the waters. This is typical of the stratified conditions generally observed in the Mediterranean Sea in late spring/early summer (e.g., Bosc et al., 2004; D'Ortenzio et al., 2005). Despite similar total chl a concentrations at the three stations (Gazeau et al., 2021a), PP was higher at FAST (Table 1; Gazeau et al., 2021b; Marañón et al., 2021). The initial microbial communities differed substantially between the three stations, as shown by the pigments (Gazeau et al., 2021a) from the 18S and 16S rDNA sequencing (this study). DOC concentrations were slightly higher at TYR, where PP was the lowest (Gazeau et al., 2021b). HB, HNF abundances (Gazeau et al., 2021a), and viral abundance and production increased following the east-to-west gradient of the initial water conditions.
The dust addition induced a similar nitrate + nitrite (NOx) and dissolved inorganic phosphate (DIP) release during all three experiments. Rapid changes were observed on plankton stocks (autotroph and heterotroph abundances and chl a; Gazeau et al., 2021a) and metabolisms (BP and PP; Gazeau et al., 2021b), suggesting that the impact of dust deposition is constrained by the initial composition and metabolic state of the investigated community. While no direct effect of warming and acidification was observed on the amount of nutrient released from dust, Gazeau et al. (2021a, b) showed that biological processes were generally enhanced by these conditions and suggested that deposition may weaken the biological pump in future climate conditions. Here we are further investigating how the dust addition in present and future conditions affected, on a short-term scale (≤ 4 d), the microbial trophic interactions and community composition.
4.1 Trophic interactions after dust addition under present and future conditions
Parallel nutrient enrichment incubations conducted in darkness showed that in situ heterotrophic bacterioplankton communities (initial conditions of the present experiments) were N, P co-limited at TYR, mainly P limited at ION, and N limited at FAST (Van Wambeke et al., 2021). However, after incubation, the HB appeared to be weakly bottom-up controlled (Ducklow, 1992) especially in D and G (Fig. 2a) after the dust addition. Such a top-down control on the bacterioplankton has been previously observed in the Mediterranean Sea, where the bacterioplankton community lives in a dynamic equilibrium between grazing pressure and nutrients limitation (as reviewed by Siokou-Frangou et al., 2010). Moreover, a potential increase under future conditions, as suggested by the higher top-down index in G (G = 0.92 vs. C/D = 0.80; Morán et al., 2017), should be further assessed.
Bacterial mortality increased relative to controls in D and G at TYR and only in G at ION and FAST. The weak coupling between bacteria and viruses, as well as the increased virus production and relative abundance of lytic cells (see below), only explained a small fraction of the estimated bacterial mortality (max. 17 %), suggesting an additional grazing pressure on bacteria. Nanoflagellates bacterivory can account for up to 87 % of bacterial production in the Mediterranean Sea; however, the rates can be variable in space and time (Siokou-Frangou et al., 2010). Here, HNF abundances increased in D at TYR and at all stations in G (Gazeau et al., 2021a), which could explain the increased bacterial mortality. An increased grazing rate by HNF on bacteria with a dust addition has been previously reported in the eastern Mediterranean Sea (Tsiola et al., 2017). While our results suggest a strong grazing pressure on bacteria, no direct coupling between HNF and bacteria was observed, probably because the HNF appeared to be top-down controlled themselves (Gasol, 1994, Fig. 3b), potentially by the increasing populations of mixotrophic dinoflagellates and/or giruses (see below). This suggests an intensification of trophic cascades in the microbial loop with nutrient input. It is also possible that HB were grazed by mixotrophic nanoflagellates or by larger protozoans or that the HNF abundance was underestimated by flow cytometry. Towards the end of the experiment, bacterial growth and mortality may also have been linked to DIP depletion at TYR and ION.
Considering the seasonal impact of grazing and the viral mortality in the Mediterranean Sea, where higher grazing pressure and lysogeny were observed in the stratified nutrient-limited waters in summer (Sánchez et al., 2020), it will be important to further study the seasonal impact of dust deposition on trophic interactions and indirect cascading impact on microbial dynamics and community composition.
4.2 Viral processes and community during dust enrichment in present and future conditions
Viruses represent pivotal components of the marine food web, influencing genome evolution, community dynamics, and ecosystem biogeochemistry (Suttle, 2007). The impacts of marine viruses differ, depending on whether they establish lytic or lysogenic infections (Zimmerman et al., 2019; Howard-Varona et al., 2017). Understanding how viral infection processes are influenced by changes in environmental conditions is, thus, crucial to better constraining the microbial mortality and cascading effects on marine ecosystems. Aerosol deposition was already identified as a factor that stimulates virus production and the viral-induced mortality of bacteria in the Mediterranean Sea (Pulido-Villena et al., 2014; Tsiola et al., 2017), and the direct deposition of airborne viruses and viruses attached to dust particles may also affect microbial food webs (Sharoni et al., 2015; Rahav et al., 2020). However, the impact of future environmental conditions remains more controversial (Larsen et al., 2008; Brussaard et al., 2013; Maat et al., 2014; Vaqué et al., 2019; Malits et al., 2021). The combined effect of aerosol deposition and the future conditions of temperature and pH on the viral compartment has, to our knowledge, never been investigated.
The rapid changes in viral production and lifestyle observed in all three experiments support the idea that the viral component is sensitive to the environmental variability, even on short (hourly) timescales. The dynamics in viral activities were, however, impacted differently, depending on the treatments and the experiments. Viral production increased in D and G at TYR and only in G at ION and FAST. Regarding the G treatments, an increase in the viral production was detected before the dust addition for all three experiments and remained mostly unchanged for the remainder of the incubation. This suggests that water warming and acidification were responsible for most changes in viral activities, while dust had no detectable impact in such conditions, regardless of the studied station. Based on our results, the most likely explanation for the observed changes in viral production is an activation of a lysogenic to lytic switch. The factors that result in prophage induction are still not well constrained, but nutrient pulses and elevated temperatures have been identified as potential stressors (Danovaro et al., 2011 and references therein). Consistent with the observation of N and P co-limited bacterial community at TYR, it is likely that nutrients released from dust upon deposition to surface water activate the productive cycle of temperate viruses at this station. Such a mechanism was also speculated during another dust addition study (Pulido-Villena et al., 2014). Under future conditions (G), the low proportion of lysogens was associated to a higher frequency of lytically infected cells relative to C and D at TYR and ION. These trends probably reflect an indirect effect of enhanced bacterial growth with increased temperature, not only on prophage induction (Danovaro et al., 2011; Vaqué et al., 2019; Mojica and Brussaard, 2014) but also on the kinetics of lytic infections. Intriguingly, the enhanced viral production did not translate into marked changes in the viral abundance. The abundance of a low DNA virus population, which typically comprises the viruses of bacteria, actually decreased between t0 and t48h pointing to a possible viral decay, potentially related to an adsorption onto dust particles (Weinbauer et al., 2009; Yamada et al., 2020) and the potential export of viral particle to deeper water layers (Van Wambeke et al., 2021). While recurrent patterns emerged from this study, the amplitude of viral responses varied between the experiments. At TYR, where heterotrophic metabolism was higher, the dust addition induced a higher viral production relative to controls than at the two other sites, which suggests that viral processes, as with other microbial processes, are dependent on the initial metabolic status of the water.
Overall, no marked changes were observed for viral communities and abundances after dust addition under both present and future conditions relative to controls, except at FAST, where the abundance of the girus population increased significantly in G from t24h until the end of the experiment. Giruses typically comprise large double-stranded DNA viruses that infect nanoeukaryotes, including photosynthetic (microalgae) and heterotrophic (HNF, amoeba, and choanoflagellate) organisms (Brussaard and Martinez, 2008; Needham et al., 2019; Fischer et al., 2010; Martínez et al., 2014). The presence of giruses at FAST in this treatment might be explained by the increase in nano-eukaryote abundances at t72h and their decline after 96 h of incubation (Gazeau et al., 2021a). The coccolithophore Emiliania huxleyi appears as one of the potential host candidates for these giruses. The abundance of E. huxleyi increased in D and G at this station, and this phytoplankter is known to be infected by such giant viruses (Jacquet et al., 2002; Schroeder et al., 2002; Pagarete et al., 2011). It is not clear from our results whether increased girus abundance is due to the greenhouse effect only (as discussed above for viruses of HB) or the combination of dust addition and the greenhouse effect. While temperature warming was shown to accelerate viral production in several virus–phytoplankton systems (Mojica and Brussaard, 2014; Demory et al., 2017), a temperature-induced resistance to viral infection was specifically observed in E. huxleyi (Kendrick et al., 2014). Previous experiments have also reported a negative impact of acidification on E. huxleyi virus dynamics (Larsen et al., 2008). By contrast, the nutrient release following dust seeding could indirectly stimulate E. huxleyi virus production (Bratbak et al., 1993) or induce switching between the non-lethal temperate to lethal lytic stage (Knowles et al., 2020) under future conditions. Targeted analyses are, of course, required to identify the viral populations selected in G and the outcomes of their infection. Nonetheless, this is the first time, to our knowledge, that dust deposition and enhanced temperature and acidification have been shown to induce the proliferation of giruses. The impact of dust deposition under future environmental conditions on the viral infections processes could have significant consequences for microbial evolution, food web processes, biogeochemical cycles, and carbon sequestration.
4.3 Microbial community dynamic after dust addition under present and future conditions
While changes in bacterial community composition during various type of dust addition experiments have shown only minor transient responses (e.g., Marañon et al., 2010; Hill et al., 2010; Laghdass et al., 2011; Pulido-Villena et al., 2014; Marín-Beltrán et al., 2019), here the microbial community structure showed quick, significant, and sustained changes in response to dust addition in all three experiments. Similar to other parameters observed during these experiments (discussed above and in Gazeau et al., 2021a, b), the degree of response in terms of community composition was specific to the tested waters.
At TYR, where primary production was low, only transient changes after 24 h of incubation were observed before the microeukaryotes community converged back close to initial conditions. In contrast, the bacterial community significantly and rapidly changed after 24 h and remained different after 72 h. At FAST, where the addition of dust appeared to promote autotrophic processes, the microeukaryotes community responded quickly in the 24 h after dust addition, while minor and delayed changes, probably related to the lower growth rates compared to the other tested waters, were observed in the bacterial community. At ION, both eukaryotes and the bacterial community responded to the dust addition. The delayed response of microeukaryotes after 72 h, compared to the quick bacterial response at 24 h, suggests that HB were better at competing for nutrient inputs at this station and that autotrophic processes may be responding to bacterial nutrient regeneration after a lag phase, further suggesting the tight coupling between heterotrophic bacteria and phytoplankton at this station. The combined effect of decreased pH and elevated temperature on marine microbes is not yet well understood (reviewed in O'Brien et al., 2016). The absence of significant community changes at TYR and FAST, while changes were observed at ION, suggests that the response might be dependent on other environmental factors which need to be further studied.
Dust addition likely favors certain group of microorganisms, suggesting a quicker response of fast-growing/copiotrophic groups and the increase in specialized functional groups (Guo et al., 2016; Westrich et al., 2016; Maki et al., 2016). The potential toxicity effects of metals and biological particles released from dust/aerosols on certain microorganisms have also been reported (Paytan et al., 2009; Rahav et al., 2020). Here, the microeukaryotic community was dominated by a diverse group of dinoflagellates which were responsible for the main variations between treatments at all stations. The overwhelming abundance of dinoflagellates sequences over other microeukaryotes could be biased by the large genomes and multiple ribosomal gene copies per genome found in dinoflagellates (Zhu et al., 2005) or due to their preferential amplification. However, the dominance of dinoflagellates in surface water at this time of the year in the Mediterranean Sea is not uncommon (García-Gómez et al., 2020) and was also observed in surface waters of the three sampled stations by the Imaging Flow Cytobot (Marañón et al., 2021). While pigment data suggest an increase in haptophytes and pelagophytes in D (Gazeau et al., 2021a), the sequencing data only show the presence of E. huxleyi as being responsible for some of the community changes after dust addition at ION and FAST. These pigments could also indicate the presence of dinoflagellates through tertiary endosymbiosis, in particular for Karlodinium sp. (Yoon et al., 2002; Zapata et al., 2012), which is an important mixotrophic dinoflagellate (Calbet et al., 2011) observed in D and G at ION and FAST. The variations in dinoflagellate groups might have important trophic impacts due to their diverse mixotrophic states (Stoecker et al., 2017), and the effect of dust addition on mixotrophic interactions should be further studied to better understand the cascading impact of dust on food webs and the biological pump.
Positive to toxic impacts on cyanobacteria have been reported from atmospheric deposition experiments (e.g., Paytan et al., 2009; Zhou et al., 2021, Rahav et al., 2020). Here, Synechococcus appeared to be inhibited at TYR, while it was enhanced at ION and FAST, especially under future conditions (this study; Gazeau et al., 2021a). The same ASVs appeared to be inhibited at TYR and ION, while being promoted at FAST, and different ASVs increased at ION. Synechococcus has recently been shown to be stimulated by wet aerosol addition in P-limited conditions but inhibited in N-limited conditions in the South China Sea (Zhou et al., 2021). It was also shown to be repressed by dust addition in the nutrient-limited tropical Atlantic (Marañon et al., 2010). This suggests that different Synechococcus ecotypes (Sohm et al., 2016) might respond differently to dust addition, depending on the initial biogeochemical conditions of the water.
In the three experiments, the main bacterial ASVs responsible for the differences between the control and treatments were closely related to different Alteromonas strains. Alteromonas are ubiquitous in the marine environment and can respond rapidly to nutrient pulses (López-Pérez and Rodriguez-Valera, 2014). Some Alteromonas are capable of growing on a wide range of carbon compounds (Pedler et al., 2014). They can produce iron-binding ligands (Hogle et al., 2016) to rapidly assimilate Fe released from dust. Thus, they could have significant consequences for the marine carbon and Fe cycles during dust deposition events. Other copiotrophic Gammaproteobacteria, such as Vibrio, have been observed to bloom after dust deposition in the Atlantic Ocean (Westrich et al., 2016). Guo et al. (2016), using RNA sequencing, also show that Gammaproteobacteria quickly outcompete Alphaproteobacteria (mainly SAR11 and Rhodobacterales) that were initially more active. Here, while SAR11 relative abundance decreased in all experiments after 24 h, other Alphaproteobacteria related to the aerobic anoxygenic phototroph (AAP) Erythrobacter sp., increased in response to dust, particularly under future conditions. Other AAP, such as OM60, also responded to dust addition in our experiment and in the eastern Mediterranean Sea (Guo et al., 2016). Moreover, bacteriochlorophyll a, a light-harvesting pigment present in AAP, was generally higher in dust addition treatments, especially under future conditions compared to controls (Fig. S9). Fast-growing AAP might quickly outcompete other HB by supplementing their growth with light-derived energy (e.g., Koblížek, 2015). They have also been shown to be stimulated by higher temperature (Sato-Takabe et al., 2019). AAP response to dust and future conditions could have a significant role in marine biogeochemical cycles.
The microbial food web response to dust addition was dependent on the initial state of the microbial community in the tested waters. A different response in trophic interactions and community composition of the microbial food web to the wet dust addition was observed at each station. Generally, greater changes were observed in future conditions. A pulsed input of nutrients and trace metals changed the microbial ecosystem from a bottom-up limited to a top-down controlled bacterial community, likely from grazing and induced lysogeny. The composition of the mixotrophic microeukaryotes and phototrophic prokaryotes was also altered.
Overall, the impact of such a simulated pulsed nutrient deposition will depend on the initial biogeochemical conditions of the ecosystem, likely with a possible large impact on microbial trophic interactions, particularly viral processes and community structure. All effects might be generally enhanced in future climate scenarios. The impact of dust deposition on metabolic processes and the consequences for the carbon and nitrogen cycles and the biological pump based on these minicosm experiments are further discussed in Gazeau et al. (2021b) and Ridame et al. (2022), and the in situ effect of a wet dust deposition event is explored in Van Wambeke et al. (2021) in this special issue.
The PEACETIME dataset will be available at https://doi.org/10.17882/75747 (Guieu et al., 2020b) upon the publication of all papers in the special issue (2022).
The supplement related to this article is available online at: https://doi.org/10.5194/bg-19-1303-2022-supplement.
FG and CG designed the experiment. All authors participated in the sampling or sample processes. JD analyzed the data and wrote the paper, with contributions from all authors.
At least one of the (co-)authors is a member of the editorial board of Biogeosciences. The peer-review process was guided by an independent editor, and the authors also have no other competing interests to declare.
Publisher’s note: Copernicus Publications remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
This article is part of the special issue “Atmospheric deposition in the low-nutrient–low-chlorophyll (LNLC) ocean: effects on marine life today and in the future (ACP/BG inter-journal SI)”. It is not associated with a conference.
We gratefully acknowledge the onboard support from the captain and crew of the R/V Pourquoi Pas? and our chief scientists Cécile Guieu and Karine Desboeuf. We also thank Kahina Djaoudi, Julia Uitz, Céline Dimier, Philippe Catala, Barbara Marie, and María Perez-Lorenzo, for their assistance in the sampling and analysis of pigments, microbial abundance, DOC concentration, and primary production. We thank the two referees, for their input.
This study is a contribution to the PEACETIME project (http://peacetime-project.org, last access: 26 January 2022, https://doi.org/10.17600/17000300), a joint initiative of the MERMEX and ChArMEx components supported by CNRS-INSU, IFREMER, CEA, and Météo-France as part of the program MISTRALS, which is coordinated by INSU. PEACETIME was endorsed as a process study by GEOTRACES and SOLAS. Part of this research has been funded by the ANR CALYPSO attributed to ACB (grant no. ANR-15-CE01-0009). Emilio Marañón has been supported by the Spanish Ministry of Science, Innovation, and Universities (grant no. PGC2018-094553B-I00). Julie Dinasquet has been funded by a Marie Skłodowska-Curie Actions International Postdoctoral Fellowship (grant no. PIOF-GA-2013-629378).
This paper was edited by Christine Klaas and reviewed by two anonymous referees.
Allen, R., Hoffmann, L. J., Law, C. S., and Summerfield, T. C.: Subtle bacterioplankton community responses to elevated CO2 and warming in the oligotrophic South Pacific gyre, Env. Microbiol. Rep., 12, 377–386, https://doi.org/10.1111/1758-2229.12844, 2020.
Bolyen, E., Rideout, J. R., Dillon, M. R., Bokulich, N. A., Abnet, C. C., Al-Ghalith, G. A., Alexander, H., Alm, E. J., Arumugam, M., Asnicar, F., Bai, Y., Bisanz, J. E., Bittinger, K., Brejnrod, A., Brislawn, C. J., Brown, C. T., Callahan, B. J., Caraballo-Rodríguez, A. M., Chase, J., Cope, E. K., Da Silva, R., Diener, C., Dorrestein, P. C., Douglas, G. M., Durall, D. M., Duvallet, C., Edwardson, C. F., Ernst, M., Estaki, M., Fouquier, J., Gauglitz, J. M., Gibbons, S. M., Gibson, D. L., Gonzalez, A., Gorlick, K., Guo, J., Hillmann, B., Holmes, S., Holste, H., Huttenhower, C., Huttley, G. A., Janssen, S., Jarmusch, A. K., Jiang, L., Kaehler, B. D., Kang, K. B., Keefe, C. R., Keim, P., Kelley, S. T., Knights, D., Koester, I., Kosciolek, T., Kreps, J., Langille, M. G. I., Lee, J., Ley, R., Liu, Y.-X., Loftfield, E., Lozupone, C., Maher, M., Marotz, C., Martin, B. D., McDonald, D., McIver, L. J., Melnik, A. V., Metcalf, J. L., Morgan, S. C., Morton, J. T., Naimey, A. T., Navas-Molina, J. A., Nothias, L. F., Orchanian, S. B., Pearson, T., Peoples, S. L., Petras, D., Preuss, M. L., Pruesse, E., Rasmussen, L. B., Rivers, A., Robeson, M. S., Rosenthal, P., Segata, N., Shaffer, M., Shiffer, A., Sinha, R., Song, S. J., Spear, J. R., Swafford, A. D., Thompson, L. R., Torres, P. J., Trinh, P., Tripathi, A., Turnbaugh, P. J., Ul-Hasan, S., van der Hooft, J. J. J., Vargas, F., Vázquez-Baeza, Y., Vogtmann, E., von Hippel, M., Walters, W., Wan, Y., Wang, M., Warren, J., Weber, K. C., Williamson, C. H. D., Willis, A. D., Xu, Z. Z., Zaneveld, J. R., Zhang, Y., Zhu, Q., Knight, R., and Caporaso, J. G.: Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2, Nat. Biotechnol., 37, 852–857, https://doi.org/10.1038/s41587-019-0209-9, 2019.
Bonnet, S. and Guieu, C.: Atmospheric forcing on the annual iron cycle in the Western Mediterranean Sea: A 1-year survey, J. Geophys. Res.-Ocean., 111, C09010, https://doi.org/10.1029/2005JC003213, 2006.
Bosc, E., Bricaud, A., and Antoine, D.: Seasonal and interannual variability in algal biomass and primary production in the Mediterranean Sea, as derived from 4 years of SeaWiFS observations, Global Biogeochem. Cy., 18, GB002034, https://doi.org/10.1029/2003GB002034, 2004.
Bratbak, G., Egge, J. K., and Heldal, M.: Viral mortality of the marine alga Emiliania huxleyi (Haptophyceae) and termination of algal blooms, Mar. Ecol. Prog. Ser., 93, 39–48, 1993.
Brussaard, C. P. and Martinez, J. M.: Algal bloom viruses, Plant Viruses, 2, 1–13, 2008.
Brussaard, C. P. D.: Optimization of Procedures for Counting Viruses by Flow Cytometry, Appl. Environ. Microb., 70, 1506–1513, 10.1128/aem.70.3.1506-1513.2004, 2004.
Brussaard, C. P. D., Noordeloos, A. A. M., Witte, H., Collenteur, M. C. J., Schulz, K., Ludwig, A., and Riebesell, U.: Arctic microbial community dynamics influenced by elevated CO2 levels, Biogeosciences, 10, 719–731, https://doi.org/10.5194/bg-10-719-2013, 2013.
Calbet, A., Bertos, M., Fuentes-Grünewald, C., Alacid, E., Figueroa, R., Renom, B., and Garcés, E.: Intraspecific variability in Karlodinium veneficum: growth rates, mixotrophy, and lipid composition, Harmful Algae, 10, 654–667, 2011.
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., and Holmes, S. P.: DADA2: High-resolution sample inference from Illumina amplicon data, Nat. Methods, 13, 581–583, https://doi.org/10.1038/nmeth.3869, 2016.
Christaki, U., Courties, C., Massana, R., Catala, P., Lebaron, P., Gasol, J. M., and Zubkov, M. V.: Optimized routine flow cytometric enumeration of heterotrophic flagellates using SYBR Green I, Limnol. Oceanogr.-Meth., 9, 329–339, https://doi.org/10.4319/lom.2011.9.329, 2011.
Clarke, K. R. and Warwick, P. E.: Change in Marine Communities: An Approach to Statistical Analysis and Interpretation, Plymouth, Ltd Ed., 2001.
D'Ortenzio, F., Iudicone, D., de Boyer Montegut, C., Testor, P., Antoine, D., Marullo, S., Santoleri, R., and Madec, G.: Seasonal variability of the mixed layer depth in the Mediterranean Sea as derived from in situ profiles, Geophys. Res. Lett., 32, L12605, https://doi.org/10.1029/2005GL022463, 2005.
Danovaro, R., Corinaldesi, C., Dell'Anno, A., Fuhrman, J. A., Middelburg, J. J., Noble, R. T., and Suttle, C. A.: Marine viruses and global climate change, FEMS Microbiol. Rev., 35, 993–1034, https://doi.org/10.1111/j.1574-6976.2010.00258.x, 2011.
Degerman, R., Dinasquet, J., Riemann, L., Sjostedt de Luna, S., and Andersson, A.: Effect of resource availability on bacterial community responses to increased temperature, Aquat. Microb. Ecol., 68, 131–142, 2012.
Demory, D., Arsenieff, L., Simon, N., et al.: Temperature is a key factor in Micromonas–virus interactions, ISME J., 11, 601–612, https://doi.org/10.1038/ismej.2016.160, 2017.
Desboeufs, K., Leblond, N., Wagener, T., Bon Nguyen, E., and Guieu, C.: Chemical fate and settling of mineral dust in surface seawater after atmospheric deposition observed from dust seeding experiments in large mesocosms, Biogeosciences, 11, 5581–5594, https://doi.org/10.5194/bg-11-5581-2014, 2014.
Ducklow, H.: Factors regulating bottom-up control of bacteria biomass in open ocean plankton communities, Arch. Hydrobiol. Beih. Ergebn. Limnol., 37, 207–217, 1992.
Durrieu de Madron, X., Guieu, C., Sempéré, R., Conan, P., Cossa, D., D'Ortenzio, F., Estournel, C., Gazeau, F., Rabouille, C., Stemmann, L., Bonnet, S., Diaz, F., Koubbi, P., Radakovitch, O., Babin, M., Baklouti, M., Bancon-Montigny, C., Belviso, S., Bensoussan, N., Bonsang, B., Bouloubassi, I., Brunet, C., Cadiou, J. F., Carlotti, F., Chami, M., Charmasson, S., Charrière, B., Dachs, J., Doxaran, D., Dutay, J. C., Elbaz-Poulichet, F., Eléaume, M., Eyrolles, F., Fernandez, C., Fowler, S., Francour, P., Gaertner, J. C., Galzin, R., Gasparini, S., Ghiglione, J. F., Gonzalez, J. L., Goyet, C., Guidi, L., Guizien, K., Heimbürger, L. E., Jacquet, S. H. M., Jeffrey, W. H., Joux, F., Le Hir, P., Leblanc, K., Lefèvre, D., Lejeusne, C., Lemé, R., Loÿe-Pilot, M. D., Mallet, M., Méjanelle, L., Mélin, F., Mellon, C., Mérigot, B., Merle, P. L., Migon, C., Miller, W. L., Mortier, L., Mostajir, B., Mousseau, L., Moutin, T., Para, J., Pérez, T., Petrenko, A., Poggiale, J. C., Prieur, L., Pujo-Pay, M., Pulido, V., Raimbault, P., Rees, A. P., Ridame, C., Rontani, J. F., Ruiz Pino, D., Sicre, M. A., Taillandier, V., Tamburini, C., Tanaka, T., Taupier-Letage, I., Tedetti, M., Testor, P., Thébault, H., Thouvenin, B., Touratier, F., Tronczynski, J., Ulses, C., Van Wambeke, F., Vantrepotte, V., Vaz, S., and Verney, R.: Marine ecosystems' responses to climatic and anthropogenic forcings in the Mediterranean, Prog. Oceanogr., 91, 97–166, https://doi.org/10.1016/j.pocean.2011.02.003, 2011.
Fischer, M. G., Allen, M. J., Wilson, W. H., and Suttle, C. A.: Giant virus with a remarkable complement of genes infects marine zooplankton, P. Natl. Acad. Sci. USA, 107, 19508–19513, 2010.
García-Gómez, C., Yebra, L., Cortés, D., Sánchez, A., Alonso, A., Valcárcel-Pérez, N., Gómez-Jakobsen, F., Herrera, I., Johnstone, C., and Mercado, J. M.: Shifts in the protist community associated with an anticyclonic gyre in the Alboran Sea (Mediterranean Sea), FEMS Microbiol. Ecol., 96, 11, https://doi.org/10.1093/femsec/fiaa197, 2020.
Gasol, J. M.: A framework for the assessment of top-down vs bottom-up control of heterotrophic nanoflagellate abundance, Mar. Ecol. Prog. Ser., 113, 291–300, 1994.
Gasol, J. M. and del Giorgio, P. A.: Using flow cytometry for counting natural planktonic bacteria and understanding the structure of the plantkonic bacterial communties, Sci. Mar., 64, 197–224, 2000
Gazeau, F., Ridame, C., Van Wambeke, F., Alliouane, S., Stolpe, C., Irisson, J.-O., Marro, S., Grisoni, J.-M., De Liège, G., Nunige, S., Djaoudi, K., Pulido-Villena, E., Dinasquet, J., Obernosterer, I., Catala, P., and Guieu, C.: Impact of dust addition on Mediterranean plankton communities under present and future conditions of pH and temperature: an experimental overview, Biogeosciences, 18, 5011–5034, https://doi.org/10.5194/bg-18-5011-2021, 2021a.
Gazeau, F., Van Wambeke, F., Marañón, E., Pérez-Lorenzo, M., Alliouane, S., Stolpe, C., Blasco, T., Leblond, N., Zäncker, B., Engel, A., Marie, B., Dinasquet, J., and Guieu, C.: Impact of dust addition on the metabolism of Mediterranean plankton communities and carbon export under present and future conditions of pH and temperature, Biogeosciences, 18, 5423–5446, https://doi.org/10.5194/bg-18-5423-2021, 2021b.
Guieu, C. and Ridame, C.: Impact of atmospheric deposition on marine chemistry and biogeochemistry, in: Atmospheric Chemistry in the Mediterranean Region: Comprehensive Diagnosis and Impacts, edited by: Dulac, F., Sauvage, S., and Hamonou, E., Springer, Cham, Switzerland, 2020a.
Guieu, C., Desboeufs, K., Albani, S., Alliouane, S., Aumont, O., Barbieux, M., Barrillon, S., Baudoux, A.-C., Berline, L., Bhairy, N., Bigeard, E., Bloss, M., Bressac, M., Brito, J., Carlotti, F., de Liege, G., Dinasquet, J., Djaoudi, K., Doglioli, A., D’Ortenzio, F., Doussin, J.-F., Duforet, L., Dulac, F., Dutay, J.-C., Engel, A., Feliu-Brito, G., Ferre, H., Formenti, P., Fu, F., Garcia, D., Garel, M., Gazeau, F., Giorio, C., Gregori, G., Grisoni, J.-M., Guasco, S., Guittonneau, J., Haëntjens, N., Heimburger, L.-E., Helias, S., Jacquet, S., Laurent, B., Leblond, N., Lefevre, D., Mallet, M., Marañón, E., Nabat, P., Nicosia, A., Obernosterer, I., Perez, L. M., Petrenko, A., Pulido-Villena, E., Raimbault, P., Ridame, C., Riffault, V., Rougier, G., Rousselet, L., Roy-Barman, M., SaizLopez, A., Schmechtig, C., Sellegri, K., Siour, G., Taillandier, V., Tamburini, C., Thyssen, M., Tovar-Sanchez, A., Triquet, S., Uitz, J., Van Wambeke, F., Wagener, T., and Zaencker, B.: BIOGEOCHEMICAL dataset collected during the PEACETIME cruis, SEANOE [data set], https://doi.org/10.17882/75747, 2020b.
Guieu, C., Dulac, F., Desboeufs, K., Wagener, T., Pulido-Villena, E., Grisoni, J.-M., Louis, F., Ridame, C., Blain, S., Brunet, C., Bon Nguyen, E., Tran, S., Labiadh, M., and Dominici, J.-M.: Large clean mesocosms and simulated dust deposition: a new methodology to investigate responses of marine oligotrophic ecosystems to atmospheric inputs, Biogeosciences, 7, 2765–2784, https://doi.org/10.5194/bg-7-2765-2010, 2010.
Guieu, C., Aumont, O., Paytan, A., Bopp, L., Law, C. S., Mahowald, N., Achterberg, E. P., Marañón, E., Salihoglu, B., Crise, A., Wagener, T., Herut, B., Desboeufs, K., Kanakidou, M., Olgun, N., Peters, F., Pulido-Villena, E., Tovar-Sanchez, A., and Völker, C.: The significance of the episodic nature of atmospheric deposition to Low Nutrient Low Chlorophyll regions, Global Biogeochem. Cy., 28, 1179–1198, https://doi.org/10.1002/2014GB004852, 2014.
Guillou, L., Bachar, D., Audic, S., Bass, D., Berney, C., Bittner, L., Boutte, C., Burgaud, G., de Vargas, C., Decelle, J., del Campo, J., Dolan, J. R., Dunthorn, M., Edvardsen, B., Holzmann, M., Kooistra, W. H. C. F., Lara, E., Le Bescot, N., Logares, R., Mahé, F., Massana, R., Montresor, M., Morard, R., Not, F., Pawlowski, J., Probert, I., Sauvadet, A.-L., Siano, R., Stoeck, T., Vaulot, D., Zimmermann, P., and Christen, R.: The Protist Ribosomal Reference database (PR2): a catalog of unicellular eukaryote Small Sub-Unit rRNA sequences with curated taxonomy, Nucleic. Acids Res., 41, D597–D604, https://doi.org/10.1093/nar/gks1160, 2013.
Guo, C., Xia, X., Pitta, P., Herut, B., Rahav, E., Berman-Frank, I., Giannakourou, A., Tsiola, A., Tsagaraki, T. M., and Liu, H.: Shifts in Microbial Community Structure and Activity in the Ultra-Oligotrophic Eastern Mediterranean Sea Driven by the Deposition of Saharan Dust and European Aerosols, Front. Mar. Sci., 3, 170, https://doi.org/10.3389/fmars.2016.00170, 2016.
Highfield, A., Joint, I., Gilbert, J. A., Crawfurd, K. J., and Schroeder, D. C.: Change in Emiliania huxleyi Virus Assemblage Diversity but Not in Host Genetic Composition during an Ocean Acidification Mesocosm Experiment, Viruses, 9, 41, https://doi.org/10.3390/v9030041, 2017.
Hill, P. G., Zubkov, M. V., and Purdie, D. A.: Differential responses of Prochlorococcus and SAR11-dominated bacterioplankton groups to atmospheric dust inputs in the tropical Northeast Atlantic Ocean, FEMS Microbiol. Let., 306, 82–89, https://doi.org/10.1111/j.1574-6968.2010.01940.x, 2010.
Hogle, S. L., Bundy, R. M., Blanton, J. M., Allen, E. E., and Barbeau, K. A.: Copiotrophic marine bacteria are associated with strong iron-binding ligand production during phytoplankton blooms, Limnol. Oceanogr. Lett., 1, 36–43, https://doi.org/10.1002/lol2.10026, 2016.
Howard-Varona, C., Hargreaves, K., Abedon, S., and Sullivan, M.B.: Lysogeny in nature: mechanisms, impact and ecology of temperate phages, ISME J., 11, 1511–1520, https://doi.org/10.1038/ismej.2017.16, 2017.
Hu, C., Li, X., He, M., Jiang, P., Long, A., and Xu, J.: Effect of Ocean Acidification on Bacterial Metabolic Activity and Community Composition in Oligotrophic Oceans, Inferred From Short-Term Bioassays, Front. Microbiol., 12, 583982, https://doi.org/10.3389/fmicb.2021.583982, 2021.
IPCC: Climate Change 2013: The Physical Science Basis. Contribution of Working Group I to the Fifth Assessment Report of the Intergovernmental Panel on Climate Change, Cambridge University Press, Cambridge, United Kingdom and New York, NY, USA, 1535 pp., 2014.
Jacquet, S., Heldal, M., Iglesias-Rodriguez, D., Larsen, A., Wilson, W., and Bratbak, G.: Flow cytometric analysis of an Emiliana huxleyi bloom terminated by viral infection, Aquat. Microb. Ecol., 27, 111–124, 2002.
Kendrick, B. J., DiTullio, G. R., Cyronak, T. J., Fulton, J. M., Van Mooy, B. A., and Bidle, K. D.: Temperature-induced viral resistance in Emiliania huxleyi (Prymnesiophyceae), PLoS One, 9, e112134, https://doi.org/10.1371/journal.pone.0112134, 2014.
Kirchman, D.: Calculating microbial growth rates from data on production and standing stocks, Mar. Ecol. Prog. Ser., 233, 303–306, 2002.
Kirchman, D., Knees, E., and Hodson, R.: Leucine Incorporation and Its Potential As A Measure of Protein-Synthesis by Bacteria in Natural Aquatic Systems, Appl. Environ. Microbiol., 49, 599–607, 1985.
Knowles, B., Bonachela, J. A., Behrenfeld, M. J., Bondoc, K. G., Cael, B., Carlson, C. A., Cieslik, N., Diaz, B., Fuchs, H. L., and Graff, J. R.: Temperate infection in a virus–host system previously known for virulent dynamics, Nat. Commun., 11, 1–13, 2020.
Koblížek, M.: Ecology of aerobic anoxygenic phototrophs in aquatic environments, FEMS Microbiol. Rev., 39, 854–870, https://doi.org/10.1093/femsre/fuv032, 2015.
Krause, E., Wichels, A., Giménez, L., Lunau, M., Schilhabel, M. B., and Gerdts, G.: Small Changes in pH Have Direct Effects on Marine Bacterial Community Composition: A Microcosm Approach, PLOS ONE, 7, e47035, https://doi.org/10.1371/journal.pone.0047035, 2012.
Laghdass, M., Blain, S., Besseling, M., Catala, P., Guieu, C., and Obernosterer, I.: Effects of Saharan dust on the microbial community during a large in situ mesocosm experiment in the NW Mediterranean Sea, Aquat. Microb. Ecol., 62, 201–213, 2011.
Larsen, J. B., Larsen, A., Thyrhaug, R., Bratbak, G., and Sandaa, R.-A.: Response of marine viral populations to a nutrient induced phytoplankton bloom at different pCO2 levels, Biogeosciences, 5, 523–533, https://doi.org/10.5194/bg-5-523-2008, 2008.
Lee, S. H. and Fuhrman, J. A.: Relationships between biovolume and biomass of naturally derived marine bacterioplankton, Appl. Environ. Microbiol., 53, 1298–1303, 1987.
Loÿe-Pilot, M. and Martin, J.: Saharan dust input to the western Mediterranean: an eleven years record in Corsica, in: The impact of desert dust across the Mediterranean, Springer Netherlands, Dordrecht, 191–199, https://doi.org/10.1007/978- 94-017-3354-0_18, 1996.
López-Pérez, M. and Rodriguez-Valera, F.: The Family Alteromonadaceae, in: The Prokaryotes: Gammaproteobacteria, edited by: Rosenberg, E., DeLong, E. F., Lory, S., Stackebrandt, E., and Thompson, F., Springer Berlin Heidelberg, Berlin, Heidelberg, 69–92, https://doi.org/10.1007/978-3-642-38922-1_233, 2014.
Maat, D. S., Crawfurd, K. J., Timmermans, K. R., and Brussaard, C. P.: Elevated carbon dioxide and phosphorus limitation favor Micromonas pusilla through stimulated growth and reduced viral impact, aspects of algal host-virus interactions in a changing ocean, Appl. Environ. Microbiol., 80, 3119–3127, https://doi.org/10.1128/AEM.03639-13, 2014.
Mahowald, N. M., Scanza, R., Brahney, J., Goodale, C. L., Hess, P. G., Moore, J. K., and Neff, J.: Aerosol Deposition Impacts on Land and Ocean Carbon Cycles, Curr. Clim. Change Rep., 3, 16–31, https://doi.org/10.1007/s40641-017-0056-z, 2017.
Maki, T., Ishikawa, A., Mastunaga, T., Pointing, S. B., Saito, Y., Kasai, T., Watanabe, K., Aoki, K., Horiuchi, A., Lee, K. C., Hasegawa, H., and Iwasaka, Y.: Atmospheric aerosol deposition influences marine microbial communities in oligotrophic surface waters of the western Pacific Ocean, Deep-Sea Res. Pt. I, 118, 37–45, https://doi.org/10.1016/j.dsr.2016.10.002, 2016.
Malits, A., Boras, J. A., Balagué, V., Calvo, E., Gasol, J.M., Marrasé, C., Pelejero, C., Pinhassi, J., Sala, M. M., and Vaqué, D.: Viral-Mediated Microbe Mortality Modulated by Ocean Acidification and Eutrophication: Consequences for the Carbon Fluxes Through the Microbial Food Web, Front. Microbiol., 12, 635821, https://doi.org/10.3389/fmicb.2021.635821, 2021.
Marañon, E., Fernández, A., Mouriño-Carballido, B., MartÍnez-GarcÍa, S., Teira, E., Cermeño, P., Chouciño, P., Huete-Ortega, M., Fernández, E., Calvo-DÍaz, A., Morán, X. A. G., Bode, A., Moreno-Ostos, E., Varela, M. M., Patey, M. D., and Achterberg, E. P.: Degree of oligotrophy controls the response of microbial plankton to Saharan dust, Limnol. Oceanogr., 55, 2339–2352, https://doi.org/10.4319/lo.2010.55.6.2339, 2010.
Marañón, E., Lorenzo, M. P., Cermeño, P., and Mouriño-Carballido, B.: Nutrient limitation suppresses the temperature dependence of phytoplankton metabolic rates, ISME J., 12, 1836–1845, https://doi.org/10.1038/s41396-018-0105-1, 2018.
Marañón, E., Van Wambeke, F., Uitz, J., Boss, E. S., Dimier, C., Dinasquet, J., Engel, A., Haëntjens, N., Pérez-Lorenzo, M., Taillandier, V., and Zäncker, B.: Deep maxima of phytoplankton biomass, primary production and bacterial production in the Mediterranean Sea, Biogeosciences, 18, 1749–1767, https://doi.org/10.5194/bg-18-1749-2021, 2021.
Marín-Beltrán, I., Logue, J. B., Andersson, A. F., and Peters, F.: Atmospheric Deposition Impact on Bacterial Community Composition in the NW Mediterranean, Front. Microbiol., 10, 858, https://doi.org/10.3389/fmicb.2019.00858, 2019.
Martínez, J. M., Swan, B. K., and Wilson, W. H.: Marine viruses, a genetic reservoir revealed by targeted viromics, ISME J., 8, 1079–1088, https://doi.org/10.1038/ismej.2013.214, 2014.
Mojica, K. D. and Brussaard, C. P.: Factors affecting virus dynamics and microbial host–virus interactions in marine environments, FEMS Microbiol. Ecol., 89, 495–515, 2014.
Morán, X. A. G., Gasol, J. M., Pernice, M. C., Mangot, J.-F., Massana, R., Lara, E., Vaqué, D., and Duarte, C. M.: Temperature regulation of marine heterotrophic prokaryotes increases latitudinally as a breach between bottom-up and top-down controls, Glob. Change Biol., 23, 3956–3964, https://doi.org/10.1111/gcb.13730, 2017.
Morán, X. A. G., Baltar, F., Carreira, C., and Lønborg, C.: Responses of physiological groups of tropical heterotrophic bacteria to temperature and dissolved organic matter additions: food matters more than warming, Environ. Microbiol., 22, 1930–1943, https://doi.org/10.1111/1462-2920.15007, 2020.
Moulin, C. and Chiapello, I.: Impact of human-induced desertification on the intensification of Sahel dust emission and export over the last decades, Geophys. Res. Lett., 33, L18808, https://doi.org/10.1029/2006GL025923, 2006.
Needham, D. M., Yoshizawa, S., Hosaka, T., Poirier, C., Choi, C. J., Hehenberger, E., Irwin, N. A., Wilken, S., Yung, C.-M., and Bachy, C.: A distinct lineage of giant viruses brings a rhodopsin photosystem to unicellular marine predators, P. Natl. Acad. Sci. USA, 116, 20574–20583, 2019.
O'Brien, P. A., Morrow, K. M., Willis, B. L., and Bourne, D. G.: Implications of Ocean Acidification for Marine Microorganisms from the Free-Living to the Host-Associated, Front. Mar. Sci., 3, 47, https://doi.org/10.3389/fmars.2016.00047, 2016.
Pagarete, A., Le Corguillé, G., Tiwari, B., Ogata, H., de Vargas, C., Wilson, W. H., and Allen, M. J.: Unveiling the transcriptional features associated with coccolithovirus infection of natural Emiliania huxleyi blooms, FEMS Microbiol. Ecol., 78, 555–564, https://doi.org/10.1111/j.1574-6941.2011.01191.x, 2011.
Parada, A. E., Needham, D. M., and Fuhrman, J. A.: Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples, Environ. Microbiol., 18, 1403–1414, https://doi.org/10.1111/1462-2920.13023, 2016.
Parada, V., Herndl, G. J., and Weinbauer, M. G.: Viral burst size of heterotrophic prokaryotes in aquatic systems, JMBA – J. Mar. Biol. Assoc. UK, 86, 613–621, 2006.
Paytan, A., Mackey, K. R. M., Chen, Y., Lima, I. D., Doney, S. C., Mahowald, N., Labiosa, R., and Post, A. F.: Toxicity of atmospheric aerosols on marine phytoplankton, P. Natl. Acad. Sci. USA, 106, 4601–4605, https://doi.org/10.1073/pnas.0811486106, 2009.
Pedler, B. E., Aluwihare, L. I., and Azam, F.: Single bacterial strain capable of significant contribution to carbon cycling in the surface ocean, P. Natl. Acad. Sci. USA, 111, 7202–7207, 2014.
Pinheiro, J., Bates, D., DebRoy, S., and Sarkar, D.: R Core Team. nlme: linear and nonlinear mixed effects models, R package version 3.1-117, http://CRAN.R-project.org/package=nlme (last access: 2 October 2021), 2014.
Pitta, P., Kanakidou, M., Mihalopoulos, N., Christodoulaki, S., Dimitriou, P.D., Frangoulis, C., Giannakourou, A., Kagiorgi, M., Lagaria, A., Nikolaou, P., Papageorgiou, N., Psarra, S., Santi, I., Tsapakis, M., Tsiola, A., Violaki, K., and Petihakis, G.: Saharan Dust Deposition Effects on the Microbial Food Web in the Eastern Mediterranean: A Study Based on a Mesocosm Experiment, Front. Mar. Sci., 4, 117, https://doi.org/10.3389/fmars.2017.00117, 2017.
Pulido-Villena, E., Baudoux, A. C., Obernosterer, I., Landa, M., Caparros, J., Catala, P., Georges, C., Harmand, J., and Guieu, C.: Microbial food web dynamics in response to a Saharan dust event: results from a mesocosm study in the oligotrophic Mediterranean Sea, Biogeosciences, 11, 5607–5619, https://doi.org/10.5194/bg-11-5607-2014, 2014.
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., Peplies, J., and Glöckner, F. O.: The SILVA ribosomal RNA gene database project: improved data processing and web-based tools, Nucleic. Acids Res., 41, D590–D596, https://doi.org/10.1093/nar/gks1219, 2013.
R Core Team: R: A language and environment for statistical computing, R Foundation for Statistical Computing, Vienna, Austria, https://www.R-project.org/ (last access: 2 October 2021), 2020.
Rahav, E., Paytan, A., Mescioglu, E., Bar-Zeev, E., Martínez Ruiz, F., Xian, P., and Herut, B.: Bio-Aerosols Negatively Affect Prochlorococcus in Oligotrophic Aerosol-Rich Marine Regions, Atmosphere, 11, 540, https://doi.org/10.3390/atmos11050540, 2020.
Ridame, C., Le Moal, M., Guieu, C., Ternon, E., Biegala, I. C., L'Helguen, S., and Pujo-Pay, M.: Nutrient control of N2 fixation in the oligotrophic Mediterranean Sea and the impact of Saharan dust events, Biogeosciences, 8, 2773–2783, https://doi.org/10.5194/bg-8-2773-2011, 2011.
Ridame, C., Dinasquet, J., Hallstrøm, S., Bigeard, E., Riemann, L., Van Wambeke, F., Bressac, M., Pulido-Villena, E., Taillandier, V., Gazeau, F., Tovar-Sanchez, A., Baudoux, A.-C., and Guieu, C.: N2 fixation in the Mediterranean Sea related to the composition of the diazotrophic community and impact of dust under present and future environmental conditions, Biogeosciences, 19, 415–435, https://doi.org/10.5194/bg-19-415-2022, 2022.
Sánchez, O., Ferrera, I., Mabrito, I., Gazulla, C. R., Sebastián, M., Auladell, A., Marín-Vindas, C., Cardelús, C., Sanz-Sáez, I., Pernice, M. C., Marrasé, C., Sala, M. M., and Gasol, J. M.: Seasonal impact of grazing, viral mortality, resource availability and light on the group-specific growth rates of coastal Mediterranean bacterioplankton, Sci. Rep., 10, 19773, https://doi.org/10.1038/s41598-020-76590-5, 2020.
Sato-Takabe, Y., Hamasaki, K., and Suzuki, S.: High temperature accelerates growth of aerobic anoxygenic phototrophic bacteria in seawater, Microbiologyopen, 8, e00710–e00710, https://doi.org/10.1002/mbo3.710, 2019.
Schroeder, D., Oke, J., Malin, G., and Wilson, W.: Coccolithovirus (Phycodnaviridae): characterisation of a new large dsDNA algal virus that infects Emiliana huxleyi, Arch. Virol., 147, 1685–1698, 2002.
Sharoni, S., Trainic, M., Schatz, D., Lehahn, Y., Flores, M. J., Bidle, K. D., Ben-Dor, S., Rudich, Y., Koren, I., and Vardi, A.: Infection of phytoplankton by aerosolized marine viruses, P. Natl. Acad. Sci. USA, 112, 6643–6647, https://doi.org/10.1073/pnas.1423667112, 2015.
Siokou-Frangou, I., Christaki, U., Mazzocchi, M. G., Montresor, M., Ribera d'Alcalá, M., Vaqué, D., and Zingone, A.: Plankton in the open Mediterranean Sea: a review, Biogeosciences, 7, 1543–1586, https://doi.org/10.5194/bg-7-1543-2010, 2010.
Smith, D. C. and Azam, F.: A simple, economical method for measuring bacterial protein synthesis rates in seawater using 3H-leucine, Mar. Microb. Food Webs, 6, 102–114, 1992.
Sohm, J. A., Ahlgren, N. A., Thomson, Z. J., Williams, C., Moffett, J. W., Saito, M. A., Webb, E. A., and Rocap, G.: Co-occurring Synechococcus ecotypes occupy four major oceanic regimes defined by temperature, macronutrients and iron, ISME J., 10, 333–345, https://doi.org/10.1038/ismej.2015.115, 2016.
Stoeck, T., Bass, D., Nebel, M., Christen, R., Jones, M. D. M., Breiner, H. W., and Richards, T. A.: Multiple marker parallel tag environmental DNA sequencing reveals a highly complex eukaryotic community in marine anoxic water, Mol. Ecol., 19, 21–31, 2010.
Stoecker, D. K., Hansen, P. J., Caron, D. A., and Mitra, A.: Mixotrophy in the Marine Plankton, Ann. Rev. Mar. Sci., 9, 311–335, https://doi.org/10.1146/annurev-marine-010816-060617, 2017.
Suttle, C. A.: Marine viruses – major players in the global ecosystem, Nat. Rev. Microbiol., 5, 801–812, https://doi.org/10.1038/nrmicro1750, 2007.
Ternon, E., Guieu, C., Loÿe-Pilot, M. D., Leblond, N., Bosc, E., Gasser, B., Miquel, J. C., and Martín, J.: The impact of Saharan dust on the particulate export in the water column of the North Western Mediterranean Sea, Biogeosciences, 7, 809–826, https://doi.org/10.5194/bg-7-809-2010, 2010.
Tsiola, A., Tsagaraki, T. M., Giannakourou, A., Nikolioudakis, N., Yücel, N., Herut, B., and Pitta, P.: Bacterial Growth and Mortality after Deposition of Saharan Dust and Mixed Aerosols in the Eastern Mediterranean Sea: A Mesocosm Experiment, Front. Mar. Sci., 3, 281, https://doi.org/10.3389/fmars.2016.00281, 2017.
Van Wambeke, F., Taillandier, V., Desboeufs, K., Pulido-Villena, E., Dinasquet, J., Engel, A., Marañón, E., Ridame, C., and Guieu, C.: Influence of atmospheric deposition on biogeochemical cycles in an oligotrophic ocean system, Biogeosciences, 18, 5699–5717, https://doi.org/10.5194/bg-18-5699-2021, 2021.
Vaqué, D., Lara, E., Arrieta, J. M., Holding, J., Sà, E. L., Hendriks, I. E., Coello-Camba, A., Alvarez, M., Agustí, S., Wassmann, P. F., and Duarte, C. M.: Warming and CO2 Enhance Arctic Heterotrophic Microbial Activity, Front. Microbiol., 10, 494, https://doi.org/10.3389/fmicb.2019.00494, 2019.
Weinbauer, M., Bettarel, Y., Cattaneo, R., Luef, B., Maier, C., Motegi, C., Peduzzi, P., and Mari, X.: Viral ecology of organic and inorganic particles in aquatic systems: avenues for further research, Aquat. Microb. Ecol., 57, 321–341, 2009.
Weinbauer, M. G., Winter, C., and Höfle, M. G.: Reconsidering transmission electron microscopy based estimates of viral infection of bacterio-plankton using conversion factors derived from natural communities, Aquat. Microb. Ecol., 27, 103–110, 2002.
Weinbauer, M. G., Rowe, J. M., and Wilhelm, S.: Determining rates of virus production in aquatic systems by the virus reduction approach, in: Manual of Aquatic Virus Ecology, edited by: Wilhelm, S. W., Weinbauer, M. G., and Suttle, C. A., ASLO, 1–8, 2010.
Westrich, J. R., Ebling, A. M., Landing, W. M., Joyner, J. L., Kemp, K. M., Griffin, D. W., and Lipp, E. K.: Saharan dust nutrients promote Vibrio bloom formation in marine surface waters, P. Natl. Acad. Sci. USA, 113, 5964–5969, https://doi.org/10.1073/pnas.1518080113, 2016.
Winter, C., Herndl, G. J., and Weinbauer, M. G.: Diel cycles in viral infection of bacterioplankton in the North Sea, Aquat. Microb. Ecol., 35, 207–216, 2004.
Yamada, Y., Guillemette, R., Baudoux, A.-C., Patel, N., and Azam, F.: Viral Attachment to Biotic and Abiotic Surfaces in Seawater, Appl. Environ. Microbiol., 86, e01687–e01619, https://doi.org/10.1128/aem.01687-19, 2020.
Yoon, H. S., Hackett, J. D., and Bhattacharya, D.: A single origin of the peridinin- and fucoxanthin-containing plastids in dinoflagellates through tertiary endosymbiosis, P. Natl. Acad. Sci. USA, 99, 11724–11729, https://doi.org/10.1073/pnas.172234799, 2002.
Zapata, M., Fraga, S., Rodríguez, F., and Garrido, J. L.: Pigment-based chloroplast types in dinoflagellates, Mar. Ecol. Prog. Ser., 465, 33–52, 2012.
Zhou, W., Li, Q. P., and Wu, Z.: Coastal phytoplankton responses to atmospheric deposition during summer, Limnol. Oceanogr., 66, 1298–1315, 2021.
Zhu, F., Massana, R., Not, F., Marie, D., and Vaulot, D.: Mapping of picoeucaryotes in marine ecosystems with quantitative PCR of the 18S rRNA gene, FEMS Microbiol. Ecol., 52, 79–92, 2005.
Zimmerman, A. E., Howard-Varona, C., Needham, D. M., John, S. G., Worden, A. Z., Sullivan, M. B., Waldbauer, J. R., and Coleman, M. L.: Metabolic and biogeochemical consequences of viral infection in aquatic ecosystems, Nat. Rev. Microbiol., 18, 21–34, https://doi.org/10.1038/s41579-019-0270-x, 2020.
The requested paper has a corresponding corrigendum published. Please read the corrigendum first before downloading the article.
- Article
(978 KB) - Full-text XML
- Corrigendum
-
Supplement
(1540 KB) - BibTeX
- EndNote





