the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Basal thermal regime affects the biogeochemistry of subglacial systems
Martin Sharp
Brad Danielson
Alireza Saidi-Mehrabad
Joel Barker
Ice formed in the subglacial environment can contain some of the highest concentrations of solutes, nutrients, and microbes found in glacier systems, which can be released to downstream freshwater and marine ecosystems and glacier forefields. Despite the potential ecological importance of basal ice, our understanding of its spatial and temporal biogeochemical variability remains limited. We hypothesize that the basal thermal regime of glaciers is a dominant control on subglacial biogeochemistry because it influences the degree to which glaciers mobilize material from the underlying substrate and controls the nature and extent of biogeochemical activity that occurs at glacier beds. Here, we characterize the solutes, nutrients, and microbes found in the basal regions of a cold-based glacier and three polythermal glaciers and compare them to those found in overlying glacier ice of meteoric origin. Compared to meteoric glacier ice, basal ice from polythermal glaciers was consistently enriched in major ions, dissolved organic matter (including a specific fraction of humic-like fluorescent material), and microbes and was occasionally enriched in dissolved phosphorus and reduced nitrogen () and in a second dissolved component of humic-like fluorescent material. In contrast, the biogeochemistry of basal ice from the cold-based glacier was remarkably similar to that of meteoric glacier ice. These findings suggest that a glacier's basal thermal regime can play an important role in determining the mix of solutes, nutrients, and microbes that are acquired from subglacial substrates or produced in situ.
- Article
(1675 KB) - Full-text XML
- BibTeX
- EndNote
Glaciers form by the compression and metamorphism of snow and slowly deform and flow under their own weight. A considerable portion of a glacier's ice is of meteoric origin and receives chemical and biological inputs primarily from the atmosphere. However, subglacial processes, including melt–freeze events and erosion, can result in the production of basal ice near the bed. This basal ice is typically characterized by relatively high concentrations of solutes that are dominated by Ca2+, Mg2+, , and (Tranter, 2006). These solutes are often produced from reactions that involve carbonate and sulfide minerals (Tranter, 2006), which are trace components in most types of bedrock (Holland, 1978). Basal ice can also contain organic matter, nutrients (e.g. phosphorus, silica, potassium), and microbes from the underlying substrate (Montross et al., 2014; Sharp et al., 1999). Both basal ice and subglacial water are known to host populations of microbes that mediate redox reactions (e.g. Sharp et al., 1999; Wadham et al., 2004), play an active role in bedrock weathering (e.g. Tranter et al., 2002), and produce or consume ecologically important nutrients (e.g. Bottrell and Tranter, 2002; Boyd et al., 2011; Hodson, 2006b; Statham et al., 2008; Tranter et al., 2002; Wadham et al., 2012)
Subglacial processes and the composition of basal ice can dramatically impact the biogeochemistry of meltwater and sediments exported from glaciers in a warming world. For example, in glaciers where surface-derived meltwater drains through the subglacial environment and comes into contact with basal ice, subglacial water, and sediments, its geochemistry (Tranter et al., 2002), nutrient content (Hawkings et al., 2014; Wadham et al., 2016), and microbial community composition (Dubnick et al., 2017) are dramatically altered. Direct links have recently been established between subglacial biogeochemical signatures and impacts on downstream environments, including downstream freshwater (Sheik et al., 2015) and fjord ecosystems (Gutiérrez et al., 2015). Similarly, during glacial retreat, the biogeochemical material contained in basal ice is released to the terrestrial landscape. These materials have been directly linked to the nutrient dynamics of glacier forefields (Kazemi et al., 2016; Mindl et al., 2007; Sattin et al., 2010) and form the basis of the soils from which many postglacial landscapes evolve (Kastovská et al., 2005).
Despite the relatively high concentration and/or unique composition of solutes, nutrients, and microbes often found in subglacial systems, as well as their potential to impact glacier forefields and downstream ecosystems, our understanding of the controls on subglacial biogeochemical processes and products remains limited. We hypothesize that the basal thermal regime plays an important role in defining the physical and biogeochemical characteristics and variability of basal ice. Since warm ice deforms more easily than cold ice and subglacial water promotes basal sliding (Iken, 1981; Iken and Bindschadler, 1986), we expect basal ice that forms and persists in fast-flowing glaciers to experience relatively “warm” conditions and have distinct biogeochemistries from basal ice that forms and persists in the relatively “cold” conditions of slow-flowing glaciers. To evaluate how the basal thermal regime affects the biogeochemical materials that glaciers mobilize from the substrate or produce and cycle within subglacial environments, we explore the solutes, nutrients, and microbes found in the basal regions of three fast-flowing polythermal outlet glaciers and the slow-flowing western margin of the Devon Ice Cap (DIC, Devon Island, Nunavut, Canada).
2.1 Study site
The Devon Ice Cap (DIC) covers an area of approximately 14 400 km2 (Burgess and Sharp, 2004) and has been shrinking since 2005 (Sharp et al., 2011). The three fast-flowing glaciers from which warm basal ice was collected (Fig. 1) have surface velocities >20 m yr−1 and a temporally varying component of flow which peaks in summer and has been attributed to seasonal variation in the rate of basal motion (Burgess et al., 2005; Van Wychen et al., 2017) (Fig. 1). The occurrence of time-varying basal motion in these glaciers suggests that geothermal and frictional heat keep ice near the glacier bed at temperatures that are at or near the pressure-melting point over a considerable portion of their beds (Burgess et al., 2005; Cuffey and Paterson, 2010). Further, field observations of hydrologically active moulins on the surfaces of these glaciers suggests that surface-derived meltwater likely reaches at least some areas of their beds during the melt season. The presence of large open subglacial meltwater channels beneath the lateral margins of Sverdrup and Belcher glaciers suggests channelized subglacial water drainage. In May, prior to the initiation of surface melt, air temperatures at distances of >500 m into these subglacial channels were near 0 ∘C. We therefore assume that the basal ice in these fast-flowing glacier systems likely formed and persisted under relatively warm subglacial conditions. In contrast, ice at the western margin of the ice cap (Fig. 1) is relatively slow-flowing (with surface velocities generally <10 m a−1; Burgess et al., 2005) and is not constrained laterally by bedrock topography. We infer that glacier flow in this region occurs exclusively by internal deformation with ice frozen to the bed (Burgess et al., 2005). Therefore, basal ice at the western margin is likely below the pressure melting point and likely formed and persisted under relatively cold subglacial conditions. Although located up to ∼100 km apart, the three fast-flowing glaciers explored in this study are all underlain by metasedimentary rocks and gneiss, while ice at the western margin is largely underlain by sandstone, dolomite, and limestone bedrock (Harrison et al., 2016) (Fig. 1).
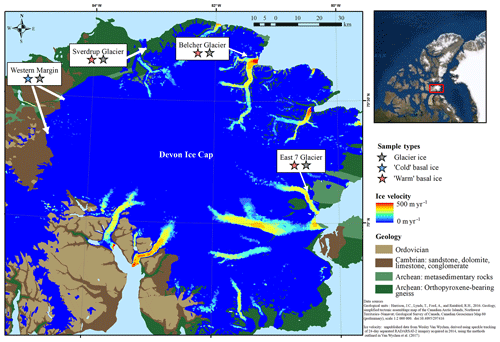
Figure 1Study site indicating the geology of the surrounding substrate and flow velocity of the Devon Ice Cap. Samples were collected from three polythermal glaciers with relatively fast-flowing ice that are surrounded by Archean bedrock and two locations along the relatively slow-flowing cold-based section of the western margin.
2.2 Field sampling
We sampled basal ice as well as overlying meteoric glacier ice from the Devon Ice Cap (Fig. 1). We identified basal ice near the glacier bed as an ice facies with high debris content and an anisotropic structure that incorporated features such as discontinuous layers, lenses, and pods of varying size (Hubbard and Sharp, 1989; Knight, 1997). In contrast, glacier ice is typically white, bubbly, and horizontally stratified. Ice samples were collected from marginal ice cliff faces, ice rubble at the base of cliff faces, and the walls of subglacial meltwater channels. All ice samples were collected using sterile (furnaced at 500 ∘C for 8 h) carbide chisels that were ethanol-bathed and flame-sterilized in the field before each use. One chisel was used for debris-poor samples and another was used for debris-rich samples. At least 10 cm of material was removed from exposed surfaces in the field before samples were collected. Samples were stored in 5 L Whirl-pak bags (Nasco, Fort Atkinson, USA) and kept frozen ( ∘C) until analysis.
2.3 Sample processing
Prior to analysis, samples were removed from the freezer and melted at 4 ∘C in Whirl-pak bags. For 16S rRNA gene sequencing, a glass filter tower and 0.2 µm Pall Supor® polysulfone 47 mm sterile filter papers were used to filter samples. Filter papers were collected in duplicate and stored in sterile Petri dishes at −80 ∘C until further processing. For analyses of soluble reactive phosphorus (SRP), , and water analyses, a sterile syringe (60 mL) with a 0.45 µm cellulose acetate luer-lock filter was used to fill two sterile 15 mL centrifuge tubes. One 120 mL Nalgene® bottle was also filled (with no headspace) for major ion analyses and stored at 4 ∘C for ∼2 weeks until analysis. For dissolved organic carbon (DOC) quantification and characterization, a 0.7 µm GF/F luer-lock filter was used to fill two sterile 45 mL universal glass vials (leaving headspace), and a piece of foil was placed beneath the cap for closure before freezing.
All filtration equipment was rinsed three times with sample, and a minimum of 5 mL of sample was passed through each filter paper before the sample was filtered for analysis. Glassware was acid-washed (10 % HCl for >48 h), and both glassware and foil were combusted (450 ∘C for 8 h) prior to use. All storage bottles, lids, and foil caps were rinsed three times with filtrate before a sample was collected for analysis.
2.4 Analytical methods
2.4.1 Nutrient concentrations (SRP/TDP///TDN/SiO2)
Determinations of soluble reactive phosphorus (SRP; ), total dissolved phosphorus (TDP), ammonium (), nitrate () + nitrite (), total dissolved nitrogen (TDN), and reactive silica (SiO2) were made with a Lachat QuickChem QC 8500 FIA Automated Ion Analyzer (Lachat Instruments, Loveland, CO, USA) using methods outlined by Rice et al. (2012) and O'Dell (1993) for + . TDP was digested with K2S2O8 to convert all dissolved P to . TDN was digested with K2S2O8 and NaOH to convert all dissolved N to . Analyses were conducted in an ISO17025 accredited laboratory and reference material and standards were applied according to those standards. Detection limits were based on instrument blanks (Shrivastava and Gupta, 2011) as follows: TDP = 1.8 ppb; SRP = 0.9 ppb; = 3 ppb; ppb; TDN = 7 ppb; and SiO2=0.02 ppm.
2.4.2 DOC concentrations
Dissolved organic carbon was quantified using a Shimadzu TOC-5000A Total Organic Carbon Analyzer (Shimadzu, Japan) equipped with a high-sensitivity platinum catalyst using US EPA method no. 415.1. Five standards between 0 and 2 ppm were used for calibration (R2=1.0). The detection limit for DOC was 0.06 ppm and was based on instrument blanks (Shrivastava and Gupta, 2011).
2.4.3 Dissolved organic matter characteristics
We used three-dimensional excitation emission matrices (EEMs) derived from total fluorescence scans to broadly characterize dissolved organic matter (DOM) into humic-like and protein-like fractions and to correlate specific fluorophores with those previously identified in the literature. DOM fluorescence was measured in ratio mode (S∕R) using an Agilent G1321B fluorescence detector (Agilent Technologies, Santa Clara, USA) and methods outlined by Cuss and Gueguen (2015). Prior to each analysis, the system was rinsed three times with deionized water and three times with sample at room temperature. EEMs were produced by measuring the fluorescence intensity every 1 nm at excitation wavelengths from 220 to 450 nm and every 5 nm at emission wavelengths from 280 to 545 nm.
2.4.4 Major ions (Cl−, , Na+, K+, Mg2+, Ca2+)
Anions were quantified using a Dionex DX-600 ion chromatograph (Dionex, USA) and methods outlined by US EPA method no. 300.1. Cations were measured using inductively coupled plasma – optical emission spectroscopy (ICP-OES; Thermo Scientific iCAP 6300, Cambridge, UK) and US EPA method no. 200.7. Detection limits were as follows: µeq L−1; µeq L−1; µeq L−1; µeq L−1; µeq L−1; and µeq L−1.
2.4.5 Sediment concentration
50 mL of unfiltered, melted ice was placed in a pre-weighed 50 mL dish and dried at 50 ∘C. The dish was then reweighed and the sediment in 50 mL was calculated as the change in mass from before to after the sample was added and dried.
2.4.6 The 16S rRNA gene sequencing
DNA was extracted from filter papers using MO BIO's PowerSoil® DNA isolation kit following the manufacturer's protocol but with several modifications to maximize the efficiency of the extraction, as follows: (1) at step 14, solution C4 was added for a total of 4 mL instead of 1200 µL and vortexing performed was for 20 s instead of 5 s; (2) prior to step 15, the samples were incubated for 30 min; and (3) at step 20, the DNA was eluted in 50 µL of solution C6 instead of 100 µL. Primers 515F and 806R were used to amplify the V4 region of the 16S rRNA gene. The Illumina libraries were prepared according to Illumina guidelines. An 8 pM library containing 7 % PhiX each was sequenced on a MiSeq instrument (Department of Biology, University of Waterloo) using a 2×250 cycle reagent kit v2.
2.5 Data processing and statistical analyses
2.5.1 Geochemistry and inorganic nutrients
The concentration of (µeq L−1) was calculated as the charge deficit between the sum of cations (µeq L−1) and the sum of anions (µeq L−1). To summarize the geochemical composition of the solute load in each sample, we (i) calculated the fractional contribution of each major ion to the total solute load by dividing the concentration of each major ion (µeq L−1) by either the sum of cations or the sum of the anions in the respective sample, (ii) normalized the fractional contributions of each ion species by their respective mean and variance (Iwamori et al., 2017), and (iii) conducted a principal component analysis (PCA) in MATLAB R2018a using these data. To summarize the nutrient composition of each sample and evaluate the concentrations of TDP and TDN relative to the total solute load, the concentrations of TDP (µg P L−1) and TDN (µg N L−1) were divided by the solute concentration (µeq L−1) of the corresponding sample. Spearman rank-order correlation coefficients (rs) were used to evaluate the significance of dependency between geochemical and nutrient variables.
2.5.2 DOM characterization
Parallel factor analysis (PARAFAC) was used to decompose the complex EEMs into discrete components using the drEEM toolbox in MATLAB 2018a and methods developed by Murphy et al. (2013). Corrections were applied for instrument spectral bias and for inner filter effects, and Raman scatter was normalized to daily Raman scans (Murphy et al., 2013). The scatter region for each EEM was excised and smoothed, and EEMs were normalized to unit variance. PARAFAC was completed using non-negativity constraints, and the EEM normalization was reversed after modeling. Although the modelled components cannot be identified as specific organic compounds, they were characterized using the OpenFluor database (Murphy et al., 2014) and comparisons with previous literature. To summarize the DOM composition of each sample, the fluorescent intensity of each component was normalized to its mean and variance across the dataset and a PCA was completed.
2.5.3 Microbial assemblage
Paired-end reads were assembled using PANDAseq (Masella et al., 2012) and analyzed using Quantitative Insights Into Microbial Ecology (QIIME, Caporaso et al., 2010), managed by the automated exploration of microbial diversity v1.5 (AXIOME, Lynch et al., 2013). Sequences were clustered with UPARSE (Edgar, 2013) into unique operational taxonomical units (OTUs) with 97 % similarity. OTUs were assigned to taxonomy via the Ribosomal Database Project (RDP) (Wang et al., 2007) with a confidence threshold of 0.8. The number of reads ranged from 7608 to 188 117 per sample so that reads were rarefied to the lowest read count (7608). The microbial assemblages in each sample were summarized by completing non-metric multidimensional scaling (NMDS) of Bray–Curtis distance measures, statistical significance between groups was determined using multi-response permutation procedure (MRPP) and multiple linear regressions to fit environmental vectors onto the NMDS (using the vegan toolbox in R).
3.1 Major ion chemistry
Glacier ice and cold basal ice had relatively low solute concentrations ( and 22 µeq L−1, respectively) that were dominated by atmospherically derived solutes, including Cl−, , and Na+ (Table 1). Warm basal ice samples contained significantly higher concentrations of solutes ( µeq L−1), including common rock-derived solutes such as K+, Ca2+, Mg2+, and , than glacier ice did (t test, p<0.05) (Table 1; Fig. 3). Therefore, while the composition and concentration of solutes in cold basal ice were very similar to those in glacier ice, it is likely that distinct solute sources exist in relatively warm subglacial systems (Fig. 2).
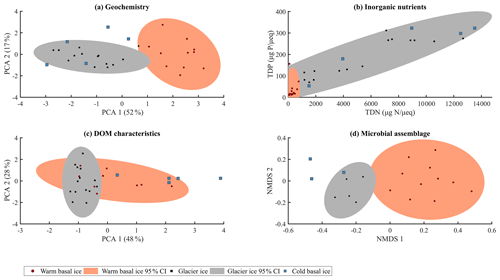
Figure 2Summary of (a) major ion composition produced by PCA using the contribution of each major ion to the solute load, with each major ion normalized to its mean and variance. (b) Inorganic nutrients relative to the solute load (TDP, µg P µeq−1; TDN, µgN µeq−1). (c) Character of dissolved organic matter determined by principle component analysis using the relative contributions of the five modelled fluorescent components, with each component normalized to its mean and variance. (d) Microbial assemblage structure determined by non-metric multidimensional scaling (NMDS) of Bray–Curtis distance measure using 16S rRNA gene sequencing (stress = 0.12).
Table 1Number, mean, and standard deviation of measures of major ions, inorganic nutrients, and DOM components in glacier ice, warm basal ice, and cold basal ice, and statistical tests between warm basal ice and cold basal and glacier ice. The p values that represent significant differences (p<0.05) are bold.
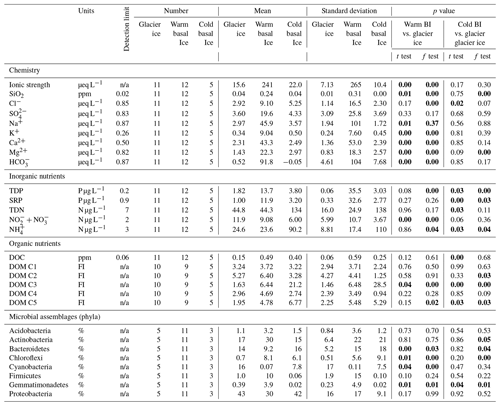
n/a – not applicable.
3.2 Nutrients (N and P)
While warm basal ice did not contain significantly more dissolved inorganic nutrients than glacier ice, including nitrogen (TDN, or ) and phosphorus (SRP, TDP) (t Test, p>0.05), it contained, on average, less (t test, p<0.05) and had higher inter-sample variability in the concentrations of , and , and TDP (f test, p<0.05; Table 1, Fig. 3). Therefore, while the subglacial system of warm-based glaciers may function as a sink of , the sources (and potentially sinks) of other inorganic nutrients may be spatially heterogeneous. Cold basal ice samples had significantly higher concentrations of , TDN, SRP, and TDP than samples of meteoric glacier ice (t test, p<0.05; Table 1), suggesting the existence of a relatively consistent subglacial source for these nutrients in this cold subglacial system.
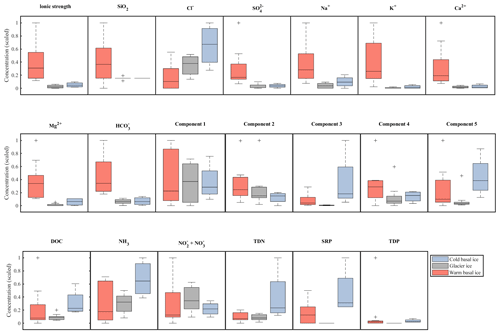
Figure 3Relative abundance and range in concentrations of major ions (top row), organic nutrients (middle row), and inorganic nutrients (bottom row) in basal ice and glacier ice. Data were scaled to the interval 0–1. Boxplots indicate the median, 25th, and 75th percentiles; whiskers indicate the most extreme datapoints that are not considered outliers; and outliers are indicated with a “+” symbol.
3.3 Dissolved organic matter
A five-component PARAFAC model explained 98.6 % of the variability in the spectrofluorescence dataset. Two of these components were similar to protein-like fluorescence and three were similar to humic-like fluorescence, as described in other studies (Table 2). DOC concentrations in warm basal ice were not significantly different from those in glacier ice (t test, p<0.05) but DOC concentrations in warm basal ice were positively correlated with the tyrosine-like C1 fluorescence (rs=0.61, p=0.01, n=9). Warm basal ice also contained significantly more humic-like C3 fluorescence (t test, p<0.05) and significantly more variable humic-like C3 and C5 fluorescence (f test, p<0.05) than glacier ice did (Table 1). Thus, a relatively consistent fraction of the DOC derived from these subglacial systems was probably in the form of proteinaceous and humic organic matter.
Table 2Excitation and emission maxima for the five-component PARAFAC model, including the identification of each component.
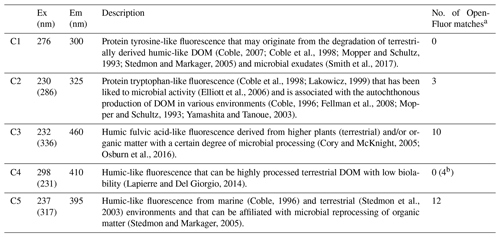

a >95 % certainty. b 90 %–95 % certainty.
Cold basal ice had significantly higher DOC concentrations ( ppm) than glacier ice ( ppm) (t test, p<0.05; Table 1), suggesting that ice can acquire DOC in cold basal environments. Relative to glacier ice, cold basal ice also contained significantly higher and more variable fluorescence of humic-like DOM components C3 and C5 (t test and f test, p<0.05) and exhibited significantly more variable fluorescence of protein-like DOM component C2 than glacier ice did (f test, p<0.05; Table 1). Since C3 and C5 humic-like DOM is typically associated with terrestrial soils and vegetation (Table 2), this DOM may have been acquired during past glacial advances over soils and sediments.
3.4 Microbial assemblages
A total of 3555 OTUs were identified across the rarefied dataset. Microbial assemblages in warm basal ice and glacier ice formed two distinct groups (Fig. 2). A total of 76 % of the OTUs observed in warm basal ice were absent from glacier ice (Fig. 4), suggesting that a large portion of the microbial assemblage in warm basal ice was sourced from the subglacial environment. Microbial assemblages in warm basal ice were also highly variable – <0.5 % of the OTUs found in warm basal ice were present in all warm basal ice samples. Geographic location influenced the structure of microbial assemblages in warm basal ice since warm basal ice samples from a given glacier had more OTUs in common with other basal ice samples from the same glacier (42 % of their OTUs and 26 % of their assemblages) than they did with basal ice samples from other warm-based glaciers (32 % of their OTUs and 15 % of their assemblages) (f test, p<0.001). Furthermore, although multiple linear regressions did not indicate any significant correlations between the major ion concentrations, major ion composition, nutrient concentrations or fluorescence index and the microbial assemblage structure of warm basal ice samples, sample location was significantly correlated with the structure of microbial assemblages in warm basal ice samples (p=0.01).
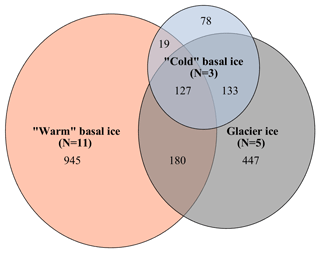
Figure 4Venn diagrams showing overlap in membership between the microbial assemblages observed in warm basal ice, glacier ice, and cold basal ice samples. Numbers represent the number of operational taxonomic units (OTUs) that are unique to each environment or shared between environments.
The microbial assemblages in cold basal ice were broadly similar to those in glacier ice (Fig. 2). Cold basal ice shared most (73 %) OTUs with glacier ice (Fig. 4). Of the shared OTUs between cold basal ice and glacier ice, many (37 %) were absent from the warm basal ice samples. Thus, the microbial assemblages in cold basal ice remained remarkably similar to those in meteoric glacier ice, despite the potential for interactions with the substrate.
4.1 Basal ice formation
Glacier ice originates as snow in the accumulation zones of glaciers/ice caps. This ice is of meteoric origin and receives chemical and biological inputs primarily from the atmosphere, experiences consistently sub-freezing temperatures and is likely to host low rates of biogeochemical activity in the englacial system (Price and Sowers, 2004). Surface melt routed through the subglacial system, or subglacially, produced meltwater formed by geothermal and frictional heat sources may refreeze to form basal ice beneath temperate and polythermal glaciers. The interactions between ice or water and the overridden substrate can mobilize particulates and solutes and incorporate them into the base of the glacier during the formation of warm basal ice (Hubbard et al., 2009). Relatively warm temperatures (i.e. near the pressure melting point) beneath the glacier may also promote biogeochemical activity by increasing both the availability of liquid water, the rates of chemical weathering, and the metabolic rates of micro-organisms (Price and Sowers, 2004).
The subglacial conditions in cold-based glaciers differ considerably from those in temperate and polythermal glaciers because temperatures are below the pressure melting point. The modes of formation of cold basal ice can vary between glaciers and are generally poorly understood. Thus, interpretations of the environments in (and processes by) which such ice is formed are often ambiguous, making our understanding of the biogeochemistry of cold basal ice even more limited. The formation of cold basal ice is often described by the “apron entrainment model” that invokes the production of basal ice by the overriding and reworking of apron material (snow, ice blocks, refrozen melt water, and debris) along an advancing margin (Shaw, 1977). However, the dark, largely bubble-free ice and absence of coarse-grained debris in the western margin basal ice facies suggests that the apron entrainment model may not describe its mode of formation. Case studies have demonstrated that basal ice in cold-based systems can also be produced by subglacial processes including the deformation and entrainment of subglacial permafrost (Fitzsimons et al., 2008), the overriding of ice marginal lakes (Lorrain et al., 1999), and the refreezing of water produced in warm thermal zones or high-pressure zones at the glacier bed that then flows into cold thermal zones or low-pressure zones downstream, where it refreezes (Knight, 1997; Wettlaufer et al., 1996) and entrains debris and excludes gases as it accretes to the glacier sole (Gilpin, 1979; Walder, 1986).
4.2 Chemistry
Warm basal ice from fast-flowing glaciers was consistently enriched in rock-derived solutes, including SiO2, , K+, Ca2+, Mg2+, and , compared to overlying glacier ice (Table 1). These solutes were likely derived from weathering of reactive minerals such as carbonates, sulfides, and aluminosilicates in the underlying bedrock (Tranter et al., 1996) that are commonly present in trace amounts (Holland, 1978). Furthermore, the presence of water during the formation of warm basal ice would support reactions involving acid hydrolysis, which are usually the most important subglacial weathering processes (Raiswell, 1984). In contrast, limited rock–water contact during the formation and persistence of cold basal ice likely limited chemical weathering, resulting in low mean solute concentrations (22 µeq L−1), similar to those in glacier ice (15.6 µeq L−1). Also like glacier ice, the solutes in cold basal ice were dominated by atmospherically derived components (Cl−, , and Na+), rather than by solutes likely to be derived from the local sandstone, dolomite, limestone, and conglomerate rocks (Ca2+, Mg2+, and ). The cold basal ice therefore contained solute concentrations and compositions more similar to those in meteoric glacier ice than warm basal ice, even though the substrate beneath the cold-based ice (local sandstone, dolomite, limestone and conglomerate substrate) is likely far more reactive than the substrate beneath the warm-based glaciers (metasedimentary rocks and gneiss). Therefore, the differences in chemistry between these basal environments cannot be explained by differences in substrate alone.
4.3 Inorganic nutrients
Both warm and cold basal ice showed some evidence of inorganic nutrient acquisition in the subglacial environment. The cold basal ice had relatively high concentrations of TDP and TDN (particularly ) (Table 1), while the warm basal ice samples were only occasionally enriched in reduced nitrogen () and dissolved phosphorus (TDP) (Table 2). The substrate surrounding the western margin sample sites is composed largely of Cambrian and Ordovician sandstone, dolomite, limestone, and conglomerate (Harrison et al., 2016), which likely contain higher phosphorus concentrations than the metasedimentary rocks (Porder and Ramachandran, 2013) that underly the polythermal glaciers in this study. Phosphorus in rocks is almost exclusively found in apatite mineral groups (Taylor and McClennan, 1985), while nitrogen can occur in recalcitrant organic matter or in silicate minerals (Honma, 1996; Honma and Schwarcz, 1979). The distribution of these components, and thus P and N, in shield rocks, volcaniclastics, and the Canadian Shield can be spatially heterogeneous (Honma, 1996; Honma and Schwarcz, 1979). Warm basal ice may also have variable inorganic nutrient concentrations if the location of subglacial biogeochemical activity is temporally or spatially heterogeneous. Subglacial microbial communities may function as a source of via catabolic processes and the degradation of organic matter. Excess would be particularly prevalent during the degradation of nitrogen-rich organic matter as has been identified in basal ice from other sites (O'Donnell et al., 2016) and observed in this study (protein-like DOM described by PARAFAC C1 and C2).
4.4 DOM
Both warm and cold basal ice contained higher average DOC concentrations (0.49 and 0.40 ppm, respectively) than glacier ice (0.15 ppm) (Table 1) suggesting a potential source of DOC in subglacial systems, as observed in Greenland (O'Donnell et al., 2016) and Antarctica (Wadham et al., 2012). Compared to glacier ice, the DOM in warm and cold basal ice had higher and more variable proportions of humic-like fluorescent material (C3 and C5) but no significant differences in the presence of C1, C2 or C4 protein-like fluorescent material (Tables 1, 2, Fig. 3). Humic DOM, and humic-like C3 and C5 fluorescence are commonly associated with soils and vegetation (Cory and McKnight, 2005; Osburn et al., 2016; Stedmon et al., 2003), thus it is possible that both the fast- and slow-flowing glaciers acquired these compounds by direct (via abiotic leaching) and indirect (via microbial cycling) of material from the substrate. Similar observations were made for low molecular weight DOC compounds in previous studies of basal ice from Greenland (O'Donnell et al., 2016). In polythermal glaciers, high rates of mechanical weathering and meltwater contact with the underlying substrate could facilitate the acquisition of humic-like DOM from the substrate. While this is unlikely the case for cold basal ice where mechanical weathering and meltwater is limited, the sedimentary rocks near or underlying the western margin support well-developed soils and vegetation. Therefore, even limited interaction with the substrate could have resulted in the acquisition of significant humic-like DOM in this cold-based system if this material was abundant in the substrate. Previous studies have also associated humic-like C3 and C5 fluorescence with microbial processing of organic matter (Table 2), suggesting basal ice may have acquired these components via heterotrophic microbial activity in subglacial environments or in supraglacial or ice marginal material that was transported into the subglacial system by meltwater. The positive correlation between DOC concentrations in warm basal ice and tyrosine-like C1 fluorescence (rs=0.61, p=0.02, n=9) indicates that a relatively consistent fraction of the DOC derived from these subglacial systems was proteinaceous in character and suggests the presence of in situ subglacial microbial activity. The production of tyrosine-like fluorescence has been widely linked to the degradation of terrestrially derived humic-like DOM and microbial exudates (Table 2). Since tyrosine-like fluorophores are considered to be highly biodegradable (Yamashita and Tanoue, 2003), it is likely that tyrosine-like fluorescence was produced in situ within the subglacial environment from the degradation of allochthonous organic matter or the production of autochthonous organic matter. This is consistent with other studies that suggest subglacial environments contain both allochthonous organic matter (i.e. from bedrock, paleosols, overridden soils, and vegetation) and autochthonous organic matter that may be produced in situ from microbial metabolism (e.g. Hodson, 2006a; O'Donnell et al., 2016).
4.5 Microbial assemblages
The microbial assemblages contained in the cold basal ice were remarkably similar to those in meteoric glacier ice; cold basal ice shared most (i.e. 73 %) OTUs with glacier ice, of which many (37 %) were unique to only cold basal ice and meteoric glacier ice (Fig. 4). Like glacier ice, the microbial assemblages observed in cold basal ice included Proteobacteria ( %), Bacteroidetes ( %), Actinobacteria ( %), and Cyanobacteria ( %) (Table 1). Proteobacteria, Bacteroidetes, Actinobacteria, and Cyanobacteria commonly dominate the microbiome of surface environments such as cryoconite holes (e.g. Cameron et al., 2012), glacier ice (e.g. Christner et al., 2005), and snow (e.g. Harding et al., 2011), and Cyanobacteria, Proteobacteria, and Actinobacteria contain organisms with the potential to photosynthesize (Cameron et al., 2012). Since microbial assemblages in soils and sediment typically differ considerably from those in the atmosphere and meteoric glacier ice, it is very unlikely that the substrate surrounding the western margin, composed of sandstone, dolomite, limestone, and conglomerate rocks and relatively well-developed soils, is characterized by the same microbial assemblages as were observed in glacier ice. Therefore, the remarkable similarity between the microbial assemblages in cold basal ice and glacier ice suggest that either the cold basal ice acquired few microbes from the underlying substrate during its formation or that most microbes acquired from the underlying substrate during its formation did not remain active in situ (and their DNA not preserved).
In contrast, the microbial assemblages in warm basal ice were distinct from those in glacier ice (Figs. 2, 4, MRPP, p<0.05). The warm basal ice therefore probably acquired distinct microbes from the underlying substrate during its formation and may have sustained in situ microbial activity, as has been reported in laboratory incubation experiments of basal material (e.g. Stibal et al., 2012a). The microbial assemblages observed in warm basal ice were dominated by Proteobacteria ( %) and Actinobacteria ( %), which are commonly observed in cold ecosystems (Amato et al., 2007; Møller et al., 2013) including those in glacier ice (this study), basal ice (Stibal et al., 2012b; Yde et al., 2010), and subglacial waters (Christner et al., 2006; Rondón et al., 2016). Additionally, the warm basal ice contained a large proportion of Chloroflexi ( %) and Gemmatimonadetes ( %), which are common and active in permafrost soils (Tuorto et al., 2014) and other basal ice environments (Yde et al., 2010) but were significantly less dominant in glacier ice samples in this study (Table 1; t test, p<0.05). Warm basal ice also contained relatively few Bacteroidetes ( %) and Cyanobacteria ( %) which were more abundant in glacier ice (Table 1) and other surface environments (Cameron et al., 2012; Harding et al., 2011). The warm basal ice yielded a very small portion of “ubiquitous” OTUs, with less than 1 % of the OTUs found in all warm basal ice samples. The abundances of these ubiquitous subglacial organisms were also highly variable between samples, ranging between a minimum of <0.01 % of the assemblage, to a maximum of 2 %–40 % of the assemblages, suggesting that the distributions of either the source(s) of these microbes and/or their in situ activity (i.e. reproduction) are spatially heterogeneous. Thus, even though there is evidence for globally distributed microbial species that are capable of survival across the range of extreme subglacial environments (Bhatia et al., 2006; Lanoil et al., 2009; Skidmore et al., 2005), these species appear to be few, and their local abundance may be highly dependent on site-specific conditions.
Although sample location did not affect the major ion chemistry, nutrient, or organic composition of basal ice, it was an important influence on the composition of the microbial assemblages in warm basal ice. We observed that microbial assemblages in basal ice samples from the same glacier were more similar to each other than were assemblages in basal ice from different glaciers. Inter-glacier differences in basal microbial assemblages were resolved over relatively small distances (less than 100 km) and between glaciers with similar basal thermal regimes and underlying substrates (Fig. 1). Geographic location has previously been identified as an important determinant of microbial assemblages across various spatial scales, from metres (Lear et al., 2014) to global (Fuhrman et al., 2008), and within other polar environments including Antarctic and Arctic terrestrial and aquatic habitats (Comte et al., 2016; Yergeau et al., 2007), as well as on glacier surfaces (Cameron et al., 2016) and in subglacial discharge (Zarsky et al., 2018). Basal ice in different glaciers can be particularly isolated from each other, so microbial dispersal between systems is probably very limited. Furthermore, although residence times of warm basal ice within a system are difficult to estimate, they may be long enough to allow stochastic processes, such as random extinction, chance colonization, drift, and priority effects (Chase and Myers, 2011; Vellend and Agrawal, 2010), to play important roles in shaping the structure of microbial assemblages in basal ice. In contrast, basal processes within a system, including ice deformation and melt–freeze effects, would provide some degree of intra-glacial mixing of microbial assemblages and may explain the higher degree of similarity between assemblages in basal ice from the same system.
We investigated the biogeochemical properties of warm basal ice from three polythermal glaciers that drain a region of the Devon Ice Cap that is underlain by metasedimentary rocks and gneiss. We found samples of basal ice from their subglacial environments to be consistently enriched in solutes (i.e. SiO2, Na+, K+, Ca2+, Mg2+, and ), a specific fraction of humic-like fluorescent DOM (C3), and microbes compared to glacier ice of meteoric origin. Although these basal ice samples were not enriched in nitrate, they were occasionally enriched in dissolved phosphorus (TDP), reduced nitrogen (), and a second component of humic-like fluorescent DOM (C5), compared to meteoric glacier ice. The sources and/or sinks of these nutrients can therefore be spatially heterogeneous in the relatively warm subglacial systems of polythermal glaciers. Large fractions of the solutes, microbes, and nutrients derived from these subglacial systems were probably acquired directly from the underlying substrate.
While basal ice in warm subglacial systems appear to have acquired abundant solutes, microbes, and nutrients from the underlying substrate, basal ice produced in cold-based systems acquired few biogeochemical characteristics from the underlying substrate. The cold basal ice explored in this study may have acquired some inorganic and organic nutrients from the substrate, but acquisition of other solutes or microbes appears to be limited. This cold basal ice acquired few solutes and microbes even though the local substrate, composed of sandstone, dolomite, limestone, and relatively well-developed soils, would have been more reactive than the metasedimentary and gneiss substrate beneath the warm-based systems. It remains unknown whether the intricacies of the biogeochemical characteristics that were observed in the cold basal ice in this study result from (i) specific characteristics of the underlying or surrounding substrate, (ii) specific glaciological and hydrological processes that occurred during the formation of the cold basal ice, or (iii) the effects of biogeochemical processes that occur in situ in cold basal ice. Further research is required to define how the cold basal ice at the western margin of the DIC developed and to better characterize the biogeochemical processes that occur in subglacial environments where liquid water is limited. Nevertheless, findings from this study suggest that basal temperature play important roles in controlling subglacial biogeochemistry and the suite of solutes, nutrients, and microbes that are either mobilized from the substrate or produced within subglacial systems.
Sequences were submitted to the National Center for Biotechnology Information Sequence Read Archive (accession number PRJNA564341). Other data from this paper are available upon request to the corresponding author.
AD conceptualized the study, AD and BD designed and completed the fieldwork, AM completed the DNA extractions in the laboratory, and AD completed the formal analysis and wrote the manuscript with reviews and edits from all authors.
The authors declare that they have no conflict of interest.
We thank the Nunavut Research Institute and the communities of Grise Fjord and Resolute Bay for permission to conduct research on Devon Island. We also thank the staff at the BASL Laboratory for chemical analyses, Chad Cuss at the SWAMP Laboratory for assistance with DOM/PARAFAC analyses, Brian Lanoil for providing laboratory facilities for completing DNA extractions, the Neufeld lab at the University of Waterloo for DNA sequencing, and past field crews on the Devon Ice Cap for accumulating such a valuable wealth of knowledge about the ice cap. We sincerely thank Maria Cavaco, Maya Bhatia, Dave Burgess, and Andy Hodson for comments and conversation that helped develop this paper and the three reviewers for providing comments that improved this paper.
This research has been supported by the Natural Sciences and Engineering Research Council of Canada (Discovery grant no. 05234-2015 and Graduate Scholarship CGSD3-475559-2015), Polar Knowledge Canada (the Northern Scientific Training Program), The W. Garfield Weston Foundation (award for Northern Research), and the Polar Continental Shelf Program (grant nos. 604-16 and 620-17).
This paper was edited by Tom J. Battin and reviewed by Marek Stibal and two anonymous referees.
Amato, P., Hennebelle, R., Magand, O., Sancelme, M., Delort, A.-M. M., Barbante, C., Boutron, C., and Ferrari, C.: Bacterial characterization of the snow cover at Spitzberg, Svalbard, FEMS Microbiol. Ecol., 59, 255–264, https://doi.org/10.1111/j.1574-6941.2006.00198.x, 2007.
Bhatia, M. P., Sharp, M. J., and Foght, J. M.: Distinct bacterial communities exist beneath a high Arctic polythermal glacier, Appl. Environ. Microb., 72, 5838–5845, https://doi.org/10.1128/AEM.00595-06, 2006.
Bottrell, S. H. and Tranter, M.: Sulphide oxidation under partially anoxic conditions at the bed of the Haut Glacier d'Arolla, Switzerland, Hydrol. Process., 16, 2363–2368, https://doi.org/10.1002/hyp.1012, 2002.
Boyd, E. S., Lange, R. K., Mitchell, A. C., Havig, J. R., Hamilton, T. L., Lafrenière, M. J., Shock, E. L., Peters, J. W. and Skidmore, M.: Diversity, abundance, and potential activity of nitrifying and nitrate-reducing microbial assemblages in a subglacial ecosystem, Appl. Environ. Microb., 77, 4778–4787, https://doi.org/10.1128/AEM.00376-11, 2011.
Burgess, D. and Sharp, M. J.: Recent Changes in Areal Extent of the Devon Ice Cap, Nunavut, Canada, Arct. Antarct. Alp. Res., 36, 261–271, 2004.
Burgess, D. O., Sharp, M. J., Mair, D. W. F., Dowdeswell, J. A., and Benham, T. J.: Flow dynamics and iceberg calving rates of Devon Ice Cap, Nunavut, Canada, J. Glaciol., 51, 219–230, https://doi.org/10.3189/172756505781829430, 2005.
Cameron, K. A., Hodson, A. J., and Osborn, A. M.: Carbon and nitrogen biogeochemical cycling potentials of supraglacial cryoconite communities, Polar Biol., 35, 1375–1393, https://doi.org/10.1007/s00300-012-1178-3, 2012.
Cameron, K. A., Stibal, M., Hawkings, J. R., Mikkelsen, A. B., Telling, J. P., Kohler, T. J., Gözdereliler, E., Zarsky, J. D., Wadham, J., and Jacobsen, C. S.: Meltwater export of prokaryotic cells from the Greenland Ice Sheet, Environ. Microbiol., 19, 524–534, https://doi.org/10.1111/1462-2920.13483, 2016.
Caporaso, J. G., Kuczynski, J., Stombaugh, J., Bittinger, K., Bushman, F. D., Costello, E. K., Fierer, N., Peña, A. G., Goodrich, J. K., Gordon, J. I., Huttley, G. A., Kelley, S. T., Knights, D., Koenig, J. E., Ley, R. E., Lozupone, C. A., Mcdonald, D., Muegge, B. D., Pirrung, M., Reeder, J., Sevinsky, J. R., Turnbaugh, P. J., Walters, W. A., Widmann, J., Yatsunenko, T., Zaneveld, J., and Knight, R.: correspondence QIIME allows analysis of high- throughput community sequencing data Intensity normalization improves color calling in SOLiD sequencing, Nat. Publ. Gr., 7, 335–336, https://doi.org/10.1038/nmeth.f.303, 2010.
Chase, J. M. and Myers, J. A.: Disentangling the importance of ecological niches from stochastic processes across scales, Philos. T. R. Soc. B, 366, 2351–2363, https://doi.org/10.1098/rstb.2011.0063, 2011.
Christner, B. C., Mosley-thompson, E., Thompson, L. G., and Reeve, J. N.: Classification of Bacteria from Polar and Nonpolar Glacial Ice, in: Life in Ancient Ice, 227–239, Princeton University Press, Princeton, NJ, USA, 2005.
Christner, B. C., Royston-Bishop, G., Foreman, C. M., Arnold, B. R., Tranter, M., Welch, K. A., Lyons, W. B., Tsapin, A. I., Studinger, M., and Priscu, J. C.: Limnological conditions in Subglacial Lake Vostok, Antarctica, Limnol. Oceanogr., 51, 2485–2501, https://doi.org/10.4319/lo.2006.51.6.2485, 2006.
Coble, P. G.: Characterization of marine and terrestrial DOM in seawater using excitiation-emission spectroscopy, Mar. Chem., 51, 325–346, 1996.
Coble, P. G.: Marine optical biogeochemistry: the chemistry of ocean color, Chem. Rev., 107, 402–418, https://doi.org/10.1021/cr050350+, 2007.
Coble, P. G., Del Castillo, C. E., and Avril, B.: Distribution and optical properties of CDOM in the Arabian Sea during the 1995 Southwest Monsoon, Deep-Sea Res. Pt. II, 45, 2195–2223, https://doi.org/10.1016/S0967-0645(98)00068-X, 1998.
Comte, J., Monier, A., Crevecoeur, S., Lovejoy, C., and Vincent, W. F.: Microbial biogeography of permafrost thaw ponds across the changing northern landscape, Ecography, 39, 609–618, https://doi.org/10.1111/ecog.01667, 2016.
Cory, R. M. and McKnight, D. M.: Fluorescence Spectroscopy Reveals Ubiquitous Presence of Oxidized and Reduced Quinones in Dissolved Organic Matter, Environ. Sci. Technol., 39, 8142–8149, https://doi.org/10.1021/es0506962, 2005.
Cuffey, K. M. and Paterson, W. S. B.: The Physics of Glaciers, 4th Edn., Butterworth-Heinemann/Elsevier, Burlington, MA, 2010.
Cuss, C. W. and Gueguen, C.: Relationships between molecular weight and fluorescence properties for size-fractionated dissolved organic matter from fresh and aged sources, Water Res., 68, 487–497, https://doi.org/10.1016/j.watres.2014.10.013, 2015.
Dubnick, A., Kazemi, S., Sharp, M., Wadham, J., Hawkings, J., Beaton, A., and Lanoil, B.: Hydrological controls on glacially exported microbial assemblages, J. Geophys. Res.-Biogeo., 1–13, https://doi.org/10.1002/2016JG003685, 2017.
Edgar, R. C.: UPARSE: Highly accurate OTU sequences from microbial amplicon reads, Nat. Methods, 10, 996–998, https://doi.org/10.1038/nmeth.2604, 2013.
Elliott, S., Lead, J. R., and Baker, A.: Characterisation of the fluorescence from freshwater, planktonic bacteria, Water Res., 40, 2075–2083, https://doi.org/10.1016/j.watres.2006.03.017, 2006.
Fellman, J. B., D'Amore, D. V., Hood, E. W., and Boone, R. D.: Fluorescence characteristics and biodegradability of dissolved organic matter in forest and wetland soils from coastal temperate watersheds in southeast Alaska, Biogeochemistry, 88, 169–184, https://doi.org/10.1007/s10533-008-9203-x, 2008.
Fitzsimons, S., Webb, N., Mager, S., MacDonell, S., Lorrain, R., and Samyn, D.: Mechanisms of basal ice formation in polar glaciers: An evaluation of the apron entrainment model, J. Geophys. Res.-Earth, 113, 1–10, https://doi.org/10.1029/2006JF000698, 2008.
Fuhrman, J. A., Steele, J. A., Hewson, I., Schwalbach, M. S., Brown, M. V., Green, J. L., and Brown, J. H.: A latitudinal diversity gradient in planktonic marine bacteria, P. Natl. Acad. Sci. USA, 105, 7774–7778, https://doi.org/10.1073/pnas.0803070105, 2008.
Gilpin, R. R.: A model of the “liquid-like” layer between ice and a substrate with applications to wire regelation and particle migration, J. Colloid Interf. Sci., 68, 235–251, https://doi.org/10.1016/0021-9797(79)90277-7, 1979.
Gutiérrez, M., Galand, P., Moffat, C., and Pantoja, S.: Melting glacier impacts the community structure of Bacteria, Fungi and Archaea in Chilean Patagonia fjord system, Environ. Microbiol., 16, 11110515, https://doi.org/10.1111/1462-2920.12872, 2015.
Harding, T., Jungblut, A. D., Lovejoy, C., and Vincent, W. F.: Microbes in high arctic snow and implications for the cold biosphere, Appl. Environ. Microb., 77, 3234–3243, https://doi.org/10.1128/AEM.02611-10, 2011.
Harrison, J. C., Lynds, T., Ford, A., and Rainbird, R. H.: Simplified tectonic assemblage map of the Canadian Arctic Islands, Geol. Surv. Canada, Can. Geosci. Map 80, scale 12 000 000, https://doi.org/10.4095/297416, 2016.
Hawkings, J. R., Wadham, J. L., Tranter, M., Raiswell, R., Benning, L. G., Statham, P. J., Tedstone, A., Nienow, P., Lee, K., and Telling, J. P.: Ice sheets as a significant source of highly reactive nanoparticulate iron to the oceans, Nat. Commun., 5, 1–8, https://doi.org/10.1038/ncomms4929, 2014.
Hodson, A. J.: Biogeochemistry of snowmelt in an Antarctic glacial ecosystem, Water Resour. Res., 42, 1–15, https://doi.org/10.1029/2005WR004311, 2006a.
Hodson, A. J.: Phosphorus in Glacial Meltwaters, in: Glacier Science and Environmental Change, edited by: Knight, P. G., 81–82, Blackwell Science Ltd, Malden, MA, 2006b.
Holland, H. D.: The chemistry of atmospheres and oceans, Wiley-Interscience, New York, 1978.
Honma, H.: High ammonium contents in the 3800 Ma Isua supracrustal rocks, central West Greenland, Geochim. Cosmochim. Ac., 60, 2173–2178, https://doi.org/10.1016/0016-7037(96)00083-X, 1996.
Honma, H. and Schwarcz, H. P.: Ammonium content of Archaean rocks of the Superior Province, Therm. Spring Res., Okayama Univ., 49, 7–13, 1979.
Hubbard, B. and Sharp, M.: Basal ice formation and deformation: a review, Prog. Phys. Geogr. Earth Environ., 13, 529–558, https://doi.org/10.1177/030913338901300403, 1989.
Hubbard, B., Cook, S., and Coulson, H.: Basal ice facies: a review and unifying approach, Quaternary Sci. Rev., 28, 1956–1969, https://doi.org/10.1016/j.quascirev.2009.03.005, 2009.
Iken, A.: The effect of the subglacial water pressure on the sliding velocity of a glacier in an idealized numerical model, J. Glaciol., 27, 407–421, 1981.
Iken, A. and Bindschadler, R. A.: Combined measurements of subglacial water pressure and surface velocity of Findelengletscher, Switzerland, J. Glaciol., 32, 101–119, 1986.
Iwamori, H., Yoshida, K., Nakamura, H., Kuwatani, T., Hamada, M., Haraguchi, S., and Ueki, K.: Classification of geochemical data based onmultivariate statistical analyses: Complementary roles of cluster, principal component, and independent component analyses, Geochem. Geophy. Geosy., 18, 994–1012, https://doi.org/10.1002/2016GC006663, 2017.
Kastovská, K., Elster, J., Stibal, M., and Santrůcková, H.: Microbial assemblages in soil microbial succession after glacial retreat in Svalbard (high arctic), Microb. Ecol., 50, 396–407, https://doi.org/10.1007/s00248-005-0246-4, 2005.
Kazemi, S., Hatam, I., and Lanoil, B.: Bacterial community succession in a high-altitude subarctic glacier foreland is a three-stage process, Mol. Ecol., 25, 5557–5567, https://doi.org/10.1111/mec.13835, 2016.
Knight, P. G.: The basal ice layer of glaciers and ice sheets, Quaternary Sci. Rev., 16, 975–993, https://doi.org/10.1016/S0277-3791(97)00033-4, 1997.
Lakowicz, R. J.: Principles of Fluorescence Spectroscopy, 2nd Edn., Kluwer Academic/Plenum Publishers, New York, 1999.
Lanoil, B., Skidmore, M., Priscu, J. C., Han, S., Foo, W., Vogel, S. W., Tulaczyk, S., and Engelhardt, H.: Bacteria beneath the West Antarctic ice sheet, Environ. Microbiol., 11, 609–615, https://doi.org/10.1111/j.1462-2920.2008.01831.x, 2009.
Lapierre, J.-F. and del Giorgio, P. A.: Partial coupling and differential regulation of biologically and photochemically labile dissolved organic carbon across boreal aquatic networks, Biogeosciences, 11, 5969–5985, https://doi.org/10.5194/bg-11-5969-2014, 2014.
Lear, G., Bellamy, J., Case, B. S., Lee, J. E., and Buckley, H. L.: Fine-scale spatial patterns in bacterial community composition and function within freshwater ponds, ISME J., 8, 1715–1726, https://doi.org/10.1038/ismej.2014.21, 2014.
Lorrain, R. D., Fitzsimons, S. J., Vandergoes, M. J., and Stievenard, M.: Ice composition evidence for the formation of basal ice from lake water beneath a cold-based Antarctic glacier, Ann. Glaciol., 28, 277–281, 1999.
Lynch, M. D. J., Masella, A. P., Hall, M. W., Bartram, A. K., and Neufeld, J. D.: AXIOME: Automated exploration of microbial diversity, Gigascience, 2, 3–7, https://doi.org/10.1186/2047-217X-2-3, 2013.
Masella, A. P., Bartram, A. K., Truszkowski, J. M., Brown, D. G., and Neufeld, J. D.: PANDAseq: Paired-end assembler for illumina sequences, BMC Bioinformatics, 13, 1–7, https://doi.org/10.1186/1471-2105-13-31, 2012.
Mindl, B., Anesio, A. M., Meirer, K., Hodson, A. J., Laybourn-Parry, J., Sommaruga, R., and Sattler, B.: Factors influencing bacterial dynamics along a transect from supraglacial runoff to proglacial lakes of a high Arctic glacieri, FEMS Microbiol. Ecol., 59, 307–317, https://doi.org/10.1111/j.1574-6941.2006.00262.x, 2007.
Møller, A. K., Søborg, D. A., Al-soud, W. A., and Sørensen, S. J.: Bacterial community structure in High-Arctic snow and freshwater as revealed by pyrosequencing of 16S rRNA genes and cultivation, Polar Res., 32, 8369, https://doi.org/10.3402/polar.v32i0.17390, 2013.
Montross, S., Skidmore, M., Christner, B., Samyn, D., Tison, J.-L., Lorrain, R., Doyle, S., and Fitzsimons, S.: Debris-Rich Basal Ice as a Microbial Habitat, Taylor Glacier, Antarctica, Geomicrobiol. J., 31, 76–81, https://doi.org/10.1080/01490451.2013.811316, 2014.
Mopper, K. and Schultz, C. A.: Fluorescence as a possible tool for studying the nature and water column distribution of DOC components, Mar. Chem., 41, 229–238, https://doi.org/10.1016/0304-4203(93)90124-7, 1993.
Murphy, K. R., Stedmon, C. A., Graeber, D., and Bro, R.: Fluorescence spectroscopy and multi-way techniques. PARAFAC, Anal. Methods, 5, 6557, https://doi.org/10.1039/c3ay41160e, 2013.
Murphy, K. R., Stedmon, C. A., Wenig, P., and Bro, R.: OpenFluor – an online spectral library of auto-fluorescence by organic compounds in the environment, Anal. Methods, 6, 658–661, https://doi.org/10.1039/C3AY41935E, 2014.
O'Dell, J. W.: Determination of Nitrate-Nitrite Nitrogen by Automated Colorimetry, United States Environmental protection Agency, Cincinnati, Ohio, 1993.
O'Donnell, E. C., Wadham, J. L., Lis, G. P., Tranter, M., Pickard, A. E., Stibal, M., Dewsbury, P., and Fitzsimons, S.: Identification and analysis of low-molecular-weight dissolved organic carbon in subglacial basal ice ecosystems by ion chromatography, Biogeosciences, 13, 3833–3846, https://doi.org/10.5194/bg-13-3833-2016, 2016.
Osburn, C. L., Boyd, T. J., Montgomery, M. T., Bianchi, T. S., Coffin, R. B., and Paerl, H. W.: Optical Proxies for Terrestrial Dissolved Organic Matter in Estuaries and Coastal Waters, Front. Mar. Sci., 2, 127, https://doi.org/10.3389/fmars.2015.00127, 2016.
Porder, S. and Ramachandran, S.: The phosphorus concentration of common rocks-a potential driver of ecosystem P status, Plant Soil, 367, 41–55, https://doi.org/10.1007/s11104-012-1490-2, 2013.
Price, P. B. and Sowers, T.: Temperature dependence of metabolic rates for microbial growth , maintenance, and survival, P. Natl. Acad. Sci. USA, 101, 4631–4636, 2004.
Raiswell, R.: Chemical models of solute acquisition in glacial melt waters, J. Glaciol., 30, 49–57, 1984.
Rice, E. W., Baird, R. B., Eaton, A. D., and Clesceri, L. S. (Eds.): Standard methods for the examination of water and wastewater, 22nd Edn., American Public Health Association, American Water Works Association, Water Environment Federation, Washington, DC, 2012.
Rondón, J., Gómez, W., Ball, M. M., Melfo, A., Rengifo, M., Balcazar, W., Dávila-Vera, D., Balza-Quintero, A., Mendoza-Briceño, R. V., and Yarzábal, L. A.: Diversity of culturable bacteria recovered from Pico Bolívar's glacial and subglacial environments, at 4,950 m, in Venezuelan Tropical Andes, Can. J. Microbiol., 62, 904–917, https://doi.org/10.1139/cjm-2016-0172, 2016.
Sattin, S. R., Cleveland, C. C., Hood, E., Reed, S. C., King, A. J., Schmidt, S. K., Robeson, M. S., Ascarrunz, N., and Nemergut, D. R.: Functional shifts in unvegetated, perhumid, recently-deglaciated soils do not correlate with shifts in soil bacterial community composition, J. Microbiol., 47, 673–681, https://doi.org/10.1007/s12275-009-0194-7, 2010.
Sharp, M., Burgess, D. O., Cogley, J. G., Ecclestone, M., Labine, C., and Wolken, G. J.: Extreme melt on Canada's Arctic ice caps in the 21st century, Geophys. Res. Lett., 38, 3–7, https://doi.org/10.1029/2011GL047381, 2011.
Sharp, M. J., Parkes, J., Cragg, B., Fairchild, I. J., Lamb, H., and Tranter, M.: Widespread bacterial populations at glacier beds and their relationship to rock weathering and carbon cycling, Geology, 27, 107–110, https://doi.org/10.1130/0091-7613(1999)027<0107:WBPAGB>2.3.CO;2, 1999.
Shaw, J.: Till body morphology and structure related, Boreas, 6, 189–201, 1977.
Sheik, C. S., Stevenson, E. I., Den Uyl, P. A., Arendt, C. A., Aciego, S. M., and Dick, G. J.: Microbial communities of the Lemon Creek Glacier show subtle structural variation yet stable phylogenetic composition over space and time, Front. Microbiol., 6, 495, https://doi.org/10.3389/fmicb.2015.00495, 2015.
Shrivastava, A. and Gupta, V. B.: Methods for the determination of limit of detection and limit of quantitation of the analytical methods, Chronicles Young Sci., 2, 21–25, https://doi.org/10.4103/2229-5186.79345, 2011.
Skidmore, M., Anderson, S. P., Sharp, M. J., Foght, J. M., and Lanoil, B.: Comparison of microbial community compositions of two subglacial environments reveals a possible role for microbes in chemical weathering processes, Appl. Environ. Microb., 71, 6986–6997, https://doi.org/10.1128/AEM.71.11.6986-6997.2005, 2005.
Smith, H. J., Foster, R. A., McKnight, D. M., Lisle, J. T., Littmann, S., Kuypers, M. M. M., and Foreman, C. M.: Microbial formation of labile organic carbon in Antarctic glacial environments, Nat. Geosci., 10, 356–359, https://doi.org/10.1038/ngeo2925, 2017.
Statham, P. J., Skidmore, M., and Tranter, M.: Inputs of glacially derived dissolved and colloidal iron to the coastal ocean and implications for primary productivity, Global Biogeochem. Cy., 22, GB3013, https://doi.org/10.1029/2007GB003106, 2008.
Stedmon, C. A. and Markager, S.: Tracing the production and degradation of autochthonous fractions of dissolved organic matter using fluorescence analysis, Limnol. Oceanogr., 50, 1415–1426, https://doi.org/10.4319/lo.2005.50.5.1415, 2005.
Stedmon, C. A., Markager, S., and Bro, R.: Tracing dissolved organic matter in aquatic environments using a new approach to fluorescence spectroscopy, Mar. Chem., 82, 239–254, https://doi.org/10.1016/S0304-4203(03)00072-0, 2003.
Stibal, M., Wadham, J. L., Lis, G. P., Telling, J. P., Pancost, R. D., Dubnick, A., Sharp, M. J., Lawson, E. C., Butler, C. E. H., Hasan, F., Tranter, M., and Anesio, A. M.: Methanogenic potential of Arctic and Antarctic subglacial environments with contrasting organic carbon sources, Glob. Change Biol., 18, 3332–3345, https://doi.org/10.1111/j.1365-2486.2012.02763.x, 2012a.
Stibal, M., Hasan, F., Wadham, J., Sharp, M. J., and Anesio, A. M.: Prokaryotic diversity in sediments beneath two polar glaciers with contrasting organic carbon substrates, Extremophiles, 16, 255–265, https://doi.org/10.1007/s00792-011-0426-8, 2012b.
Taylor, S. R. and McClennan, M. S.: The Continental Crust: its composition and evolution, Blackwell Scientific Publications, Oxford, 1985.
Tranter, M.: Glacial Chemical Weathering, Runoff Composition and Solute Fluxes, in: Glacier Science and Environmental Change, 71–75, Blackwell Science Ltd, Malden, MA, 2006.
Tranter, M., Brown, G. H., Hodson, A. J., and Gurnell, A. M.: Hydrochemistry as an indicator of subglacial drainage system structure: A comparison of alpine and sub-polar environments, Hydrologica, 10, 541–556, 1996.
Tranter, M., Sharp, M. J., Lamb, H. R., Brown, G. H., Hubbard, B. P., and Willis, I. C.: Geochemical weathering at the bed of Haut Glacier d'Arolla, Switzerland – a new model, Hydrol. Process., 16, 959–993, https://doi.org/10.1002/hyp.309, 2002.
Tuorto, S. J., Darias, P., Mcguinness, L. R., Panikov, N., and Zhang, T.: Bacterial genome replication at subzero temperatures in permafrost, ISME J., 8, 139–149, https://doi.org/10.1038/ismej.2013.140, 2014.
Van Wychen, W., Davis, J., Copland, L., Burgess, D. O., Gray, L., Sharp, M., Dowdeswell, J. A., and Benham, T. J.: Variability in ice motion and dynamic discharge from Devon Ice Cap, Nunavut, Canada, J. Glaciol., 63, 436–449, https://doi.org/10.1017/jog.2017.2, 2017.
Vellend, M. and Agrawal, A.: Conceptual Synthesis in Community Ecology, Q. Rev. Biol., 85, 183–206, https://doi.org/10.1086/652373, 2010.
Wadham, J., Arndt, S., Tulaczyk, S., Stibal, M., Tranter, M., Telling, J. P., Lis, G. P., Lawson, E. C., Ridgwell, A., Dubnick, A., Sharp, M. J., Anesio, A. M., and Butler, C. E. H.: Potential methane reservoirs beneath Antarctica, Nature, 488, 633–637, https://doi.org/10.1038/nature11374, 2012.
Wadham, J. L., Bottrell, S. H., Tranter, M., and Raiswell, R.: Stable isotope evidence for microbial sulphate reduction at the bed of a polythermal high Arctic glacier, Earth Planet. Sc. Lett., 219, 341–355, https://doi.org/10.1016/S0012-821X(03)00683-6, 2004.
Wadham, J. L., Hawkings, J., Telling, J., Chandler, D., Alcock, J., O'Donnell, E., Kaur, P., Bagshaw, E., Tranter, M., Tedstone, A., and Nienow, P.: Sources, cycling and export of nitrogen on the Greenland Ice Sheet, Biogeosciences, 13, 6339–6352, https://doi.org/10.5194/bg-13-6339-2016, 2016.
Walder, B. J. S.: Motion of sub-freezing ice past particles, with applications to wire regelation and frozen soils, J. Glaciol., 32, 1986.
Wang, Q., Garrity, G. M., Tiedje, J. M., and Cole, J. R.: Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy, Appl. Environ. Microb., 73, 5261–5267, https://doi.org/10.1128/AEM.00062-07, 2007.
Wettlaufer, J. S., Worster, M. G., Wilen, L. A., and Dash, J. G.: A Theory of Premelting Dynamics for all Power Law Forces, Phys. Rev. Lett., 76, 3602–3605, https://doi.org/10.1103/PhysRevLett.76.3602, 1996.
Yamashita, Y. and Tanoue, E.: Chemical characterization of protein-like fluorophores in DOM in relation to aromatic amino acids, Mar. Chem., 82, 255–271, https://doi.org/10.1016/S0304-4203(03)00073-2, 2003.
Yde, J. C., Finster, K. W., Raiswell, R., Steffensen, J. P., Heinemeier, J., Olsen, J., Gunnlaugsson, H. P., and Nielsen, O. B.: Basal ice microbiology at the margin of the Greenland Ice Sheet, Ann. Glaciol., 51, 71–79, https://doi.org/10.3189/172756411795931976, 2010.
Yergeau, E., Newsham, K. K., Pearce, D. A., and Kowalchuk, G. A.: Patterns of bacterial diversity across a range of Antarctic terrestrial habitats, Environ. Microbiol., 9, 2670–2682, https://doi.org/10.1111/j.1462-2920.2007.01379.x, 2007.
Zarsky, J. D., Kohler, T. J., Yde, J. C., Lamarche-gagnon, G., Hawkings, J. R., Hatton, J. E., and Stibal, M.: Prokaryotic assemblages in suspended and subglacial sediments within a glacierized catchment on Qeqertarsuaq (Disko Island), west Greenland, FEMS Microbiol. Ecol., 94, fiy100, https://doi.org/10.1093/femsec/fiy100, 2018.





