the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Unexpected high abyssal ophiuroid diversity in polymetallic nodule fields of the northeast Pacific Ocean and implications for conservation
Magdalini Christodoulou
Timothy O'Hara
Andrew F. Hugall
Sahar Khodami
Clara F. Rodrigues
Ana Hilario
Annemiek Vink
Pedro Martinez Arbizu
The largest and commercially appealing mineral deposits can be found in the abyssal sea floor of the Clarion-Clipperton Zone (CCZ), a polymetallic nodule province, in the NE Pacific Ocean, where experimental mining is due to take place. In anticipation of deep-sea mining impacts, it has become essential to rapidly and accurately assess biodiversity. For this reason, ophiuroid material collected during eight scientific cruises from five exploration licence areas within CCZ, one area being protected from mining (APEI3, Area of Particular Environmental Interest) in the periphery of CCZ and the DISturbance and re-COLonisation (DISCOL) Experimental Area (DEA), in the SE Pacific Ocean, was examined. Specimens were genetically analysed using a fragment of the mitochondrial cytochrome c oxidase subunit I (COI). Maximum-likelihood and neighbour-joining trees were constructed, while four tree-based and distance-based methods of species delineation (automatic barcode gap discovery, ABGD; barcode index numbers, BINs; general mixed Yule–coalescent, GMYC; multi-rate Poisson tree process, mPTP) were employed to propose secondary species hypotheses (SSHs) within the ophiuroids collected. The species delimitation analyses' concordant results revealed the presence of 43 deep-sea brittle star SSHs, revealing an unexpectedly high diversity and showing that the most conspicuous invertebrates in abyssal plains have been so far considerably underestimated. The number of SSHs found in each area varied from five (IFREMER area) to 24 (BGR (Federal Institute for Geosciences and Natural Resources, Germany) area) while 13 SSHs were represented by singletons. None of the SSHs were found to be present in all seven areas while the majority of species (44.2 %) had a single-area presence (19 SSHs). The most common species were Ophioleucidae sp. (Species 29), Amphioplus daleus (Species 2) and Ophiosphalma glabrum (Species 3), present in all areas except APEI3. The biodiversity patterns could be mainly attributed to particulate organic carbon (POC) fluxes that could explain the highest species numbers found in BGR (German contractor area) and UKSRL (UK Seabed Resources Ltd, UK contractor area) areas. The five exploration contract areas belong to a mesotrophic province, while conversely the APEI3 is located in an oligotrophic province, which could explain the lowest diversity as well as very low similarity with the other six study areas. Based on these results the representativeness and the appropriateness of APEI3 to meet its purpose of preserving the biodiversity of the CCZ fauna are questioned. Finally, this study provides the foundation for biogeographic and functional analyses that will provide insight into the drivers of species diversity and its role in ecosystem function.
- Article
(64523 KB) - Full-text XML
-
Supplement
(687 KB) - BibTeX
- EndNote
The deep sea holds the vastest and least explored ecosystems on Earth and has justifiably being characterized as “Earth's last frontier” since research and exploration in these areas is still incomplete at the very best (Ramirez-Llodra et al., 2010; Danovaro et al., 2017). Deep-sea habitats cover more than 65 % of the Earth's surface and can plunge from water depths of 200 m (below the continental shelf) to as deep as 11 km in the Mariana Trench (Gage and Tyler, 1991; Carney, 2005; Jamieson et al., 2009; Ramirez-Llodra, et al., 2011). Abyssal ecosystems, found between 3000 and 6000 m, cover 54 % of the Earth's surface and constitute a network of plains and hills and seamounts, segmented by mid-ocean ridges, island arcs and ocean trenches (Gage and Tyler, 1991; Carney, 2005; Smith et al., 2008). The abyssal plains represent perhaps the single largest contiguous ecosystem of our planet; nevertheless, because of its enormous size and seclusion it has been the least studied (Smith et al., 2008; Ramirez-Llodra et al., 2010). The sea floor of the abyssal plains is mostly covered by fine sediments, while hard substrates often occur in the form of polymetallic nodules (Smith et al., 2008; Ramirez-Llodra et al., 2011).
Metal-rich (polymetallic) nodules from the deep-sea floor were described and their potential economic importance acknowledged as early as 1873, during the HMS Challenger expedition (Murray and Renard, 1891; Lusty and Murton, 2018). However, it was in the 1960s that economic interest in these deposits was ignited after polymetallic nodule resources in the Pacific Ocean were estimated to be so abundant as to be an essentially endless supply of metals such as Mn, Cu, Ni and Co (Mero, 1965; Lusty and Murton, 2018). Despite the optimism in the 1970s and 1980s and the widely held belief that deep-sea mining would commence before the end of 2000, subsequent progress has been slow and unsteady. The adequate supply of metals from land-based mines, unfavourable economic conditions (e.g. rising energy costs, lower metal prices), technological challenges, increasing environmental awareness and legal obligations to international organizations (e.g. lack of a mining legislation for the deep sea) were some of the reasons slowing down deep-sea mining (Lusty and Murton, 2018). However, the growing global demand for these metals coupled with the increasing challenges of land-based mining (Calas, 2017), and the advances in mining technology, drove a renewed interest in the exploitation of deep-sea mineral deposits (Ramirez-Llodra et al., 2011; Lusty and Murton, 2018; Miller et al., 2018).
The greatest known accumulations of economically interesting Ni and Cu and Co-rich polymetallic (Fe–Mn) nodules occur in the Clarion-Clipperton Zone (CCZ), extending over an area of approximately 6×106 km2 in size, between Hawaii and Mexico from 120∘ W to approximately 160∘ W and from 20∘ N to 6∘ S. Additional important occurrences have been found in the Central Indian Ocean Basin, the Cook Islands area and the Peru Basin off South America (e.g. the DISCOL Experimental Area, DEA) (Mukhopadhyay et al., 2008; Hein et al., 2013; Miller et al., 2018). The CCZ lies in the Areas Beyond National Jurisdiction and thus falls under the legal mandate of the International Seabed Authority (ISA; Wedding et al., 2013). So far, 16 licence areas for the exploration of polymetallic nodules have been approved by the ISA within the CCZ, each up to 75 000 km2 in size (Wedding et al., 2013). In its environmental management plan for the CCZ (Lodge et al., 2014), the ISA adopted nine large protection areas defined as Areas of Particular Environmental Interest (APEIs), where mining will not be permitted (Lodge et al., 2014). These APEIs are large enough (each of them 400 km × 400 km) and far enough away from potential mining areas that they will not be affected by deep-sea mining (Wedding et al., 2013). In order to be effective as source populations for the recolonization of impacted areas, however, APEIs should harbour a representative subset of the fauna found in the potential fields. In addition to these protection measures, the ISA has stipulated that prior to exploitation, a benthic biological baseline study must be undertaken for each exploration contract area, and the possible environmental impacts arising from exploration should be assessed. Nodule mining carries significant environmental concerns, including negative direct and indirect impacts on the biodiversity (Ramirez-Llodra et al., 2011; Vanreusel et al., 2016; Van Dover et al., 2017; Niner et al., 2018). The removal of the nodules and associated organisms could result in habitat loss, fragmentation or modification while the generation of sediment plumes may bury the organisms or clog their feeding apparatuses and thus disrupt the food webs (Ramirez-Llodra et al., 2011; Vanreusel et al., 2016; Van Dover et al., 2017; Niner et al., 2018; Stratmann et al., 2018). Unfortunately, accurate documentation of species diversity, which comprises the first step in understanding patterns and structures in different levels of biodiversity and biogeographical and ecological processes and is essential for marine ecosystems' management, remains poor across the CCZ (Amon et al., 2016). To date, Taboada et al. (2018), although dealing with a single hexactinellid sponge species, are the only ones who have assessed the effectiveness of an APEI (no. 6) or investigated connectivity with the adjacent potential mining areas. Thus, prior to exploitation, there is an urgent need to obtain baseline data on faunal biodiversity at local and regional scales in order to assess and predict the effects of mining on deep-sea organisms.
The Ophiuroidea (brittle stars and basket stars) are amongst the most emblematic mobile megafaunal inhabitants of the deep sea regarding species diversity and individuals' numbers (Gage and Tyler, 1991; Rex and Etter, 2010; Vanreusel et al., 2016). They constitute the most diverse echinoderm class, numbering more than 2064 species found in all oceans, from intertidal to hadal depths (Stöhr et al., 2012; Jamieson, 2015). Since then, at least 1412 species have been recorded from the deep sea of which only 109 are from abyssal depths, despite abyssal plains being the most extensive ecosystem in the world (Stöhr et al., 2012). Studies describing the tropical abyssal northeast Pacific ophiuroid fauna are scarce but include a few historical studies resulting from the great expeditions of the late nineteenth and early twentieth centuries such as the HMS Challenger (Lyman, 1878, 1879, 1882; Ludwig, 1898, 1899) and the Albatross (Clark, 1911, 1949). Limited recent studies exist (Amon et al., 2016, 2017; Glover et al., 2016), reporting only a small number of species. Consequently, the diversity of the deep-sea ophiuroid fauna in the CCZ is only poorly known. Thus the main objectives of this study were to (1) ensure the future molecular species identification for all different life-cycle stages by matching morphology-based species identifications of adult ophiuroids with molecular species assignments using c oxidase subunit I (COI) sequences and consequently compiling a comprehensive reference library; (2) determine species ranges; (3) describe the ophiuroid biodiversity patterns of the CCZ and the DEA; and (4) explore the usefulness of APEI3 for the preservation of the ophiuroid nodule fauna in the CCZ in the case of deep-sea mining.
2.1 Study areas
The study areas are located within the Clarion and Clipperton Fracture Zone (CCZ) in the northeast equatorial Pacific Ocean and at the DISCOL Experimental Area (DEA) in the Peru Basin (Fig. 1), at depths varying from 4050 to 4933 m (Hein et al., 2013). The CCZ is characterized by gradual changes in environmental conditions (e.g. differences in surface-water productivity, depth and sediment characteristics) across an east–west and a north–south axis that correspond to a variation in nodule size and coverage, as well as variations in faunal composition along these gradients (Wedding et al., 2013). Ophiuroid samples were collected from CCZ during six scientific cruises from five different exploration licence areas and one area protected from mining (APEI3). Specifically ophiuroid samples were collected during the following cruises: BioNod on R/V L'Atalante (29 March–10 May 2012) to the eastern IFREMER (Institut Français de Recherche pour l'Exploitation de la Mer, France) licence area and to the eastern BGR (Federal Institute for Geosciences and Natural Resources, Germany) licence area; two ABYSSLINE research cruises, AB01 on the R/V Melville (3–27 October 2013) and the AB02 cruise on the R/V Thompson (12 February–25 March 2015) to the UKSRL licence area (UK Seabed Resources Ltd, United Kingdom); two MANGAN cruises, MANGAN 2013 on R/V Kilo Moana (1 April–13 May 2013) and MANGAN 2014 on R/V Kilo Moana (15 April–3 June 2014) to the eastern BGR licence area; the scientific cruise EcoResponse on R/V Sonne (SO239) (11 March–30 April 2015) to the licence areas of BGR, GSR (G-TEC Sea Mineral Resources NV, Belgium), IOM (Interoceanmetal Joint Organization, a country consortium of Bulgaria, Cuba, the Czech Republic, Poland, the Russian Federation, and Slovakia), IFREMER and APEI3. Furthermore, the DISCOL Experimental Area (DEA) in the Peru Basin, in which the German project DISCOL (DISturbance and re-COLonisation experiment) was performed in the late 1980s (Thiel and Schriever, 1990; Thiel et al., 2001), was recently revisited in the framework of the JPIO pilot action “Ecological Aspects of Deep-Sea Mining”. Ophiuroid samples were collected from the DEA during two cruises, SO242/1 and SO242/2 on the R/V Sonne from 28 July to 25 August 2015 and 28 August to 1 October 2015, respectively.
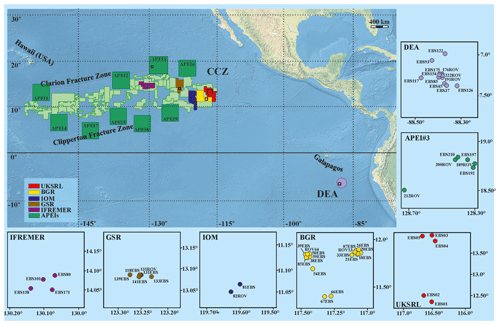
Figure 1Compilation of study areas in the Clarion-Clipperton Zone (CCZ) and in the DISCOL Experimental Area (DEA, Peru Basin). Insets represent detailed maps of sampling locations in the IFREMER, GSR, IOM, BGR and UKSRL exploration licence areas for polymetallic nodules as well as in APEI3 (ISA protected area) and the DEA. Copyright for shapefiles (for CCZ area): © International Seabed Authority 2009–2019; copyright for raster file (bathymetry): © Natural Earth 2009–2020.
2.2 Specimen sampling and processing
Small-sized ophiuroid samples were collected using a Brenke-type epibenthic sledge (EBS; Brenke, 2005) from the UKSRL (five deployments), BGR (15 deployments), IFREMER (four deployments), GSR (four deployments) and IOM (one deployment) licence areas and the APEI3 (three deployments) and the DEA (nine deployments) (Fig. 1), following standard deployment procedures (Brenke, 2005). The cod ends of the supra- and epi-net were sieved through a 500 and 300 µm mesh with cold (+10 ∘C) seawater and immediately transferred to pre-cooled (−20 ∘C) 96 % EtOH. Large-sized ophiuroid samples were collected with a remotely operated vehicle (ROV Kiel 6000, GEOMAR) using either the ROV's suction sampler or the ROV's manipulator arm by direct picking, manipulating scoops, shovels and nets. Large specimens were also preserved in pre-cooled 96 % EtOH. For all specimens the ethanol was decanted after 24 h and replaced with new 96 % EtOH to guarantee high ethanol concentration for preservation of high-quality DNA and subsequently stored at −20 ∘C. In the laboratory at Senckenberg am Meer, Germany, an integrative molecular-morphological approach was implemented for the identification of the ophiuroid specimens. In total, DNA was extracted from 525 specimens. For species delimitation analyses 300 sequences were selected (see below). All the ROV-collected specimens were photographed on board, while the EBS-collected specimens were photographed in the lab using a Leica binocular stereo-microscope or a Keyence digital microscope, VHX-5000. The voucher specimens are stored in Senckenberg am Meer, DZMB, Wilhelmshaven, Germany.
2.3 Morphological species identification
All ophiuroid individuals collected were morphologically identified to the lowest possible taxonomic level (primary species hypotheses, PSHs; Puillandre et al., 2012; Boissin et al., 2017). Where possible, individuals were assigned to named species; however, in many cases because of their very small size, their early developmental stage (post-larval individuals) or their unique morphology assignment in a morphological operational taxonomic unit was possible only at a higher taxonomic level, i.e. genus or family level. For a small number of damaged specimens, morphological identification beyond class was not possible. Following the DNA analyses (see below), all individuals within the same morphospecies that appeared to be genetically distinct from one another were re-examined and if necessary reassigned to different morphospecies, while some were considered to be true cryptic species in which clear morphological differences were not identified. Finally, the integrated approach allowed the assignment of damaged specimens into different operational taxonomic units. Taxonomic and systematic remarks for each SSH are given in the Supplement.
2.4 Barcoding data collection
2.4.1 DNA extraction, amplification and sequencing
For the mtDNA COI analyses genomic DNA was extracted from arm tissue in individuals larger than 1–2 mm or from whole individuals when smaller than 1–2 mm. DNA extractions were carried out using 30 µL Chelex (InstaGene Matrix, Bio-Rad) according to the protocol of Estoup et al. (1996) and directly used as a DNA template for polymerase chain reaction (PCR). All DNA samples were stored at −20 ∘C. In the cases where the whole individual was used, 20–25 µL of the supernatant was first separated from the ophiuroid's voucher specimen, while the individual, which was generally intact, was transferred to 96 % ethanol and stored as a voucher for morphological identifications. A fragment of 657 bp of the mitochondrial cytochrome c oxidase subunit (COI) was amplified by polymerase chain reaction (PCR). Amplifications were performed using Illustra PuReTaq Ready-To-Go PCR Beads (GE Healthcare) in a 25 µL volume containing 22 µL ddH2O, 0.5 µL of each primer (10 pmol µL−1) and 2 µL of DNA template or AccuStart PCR SuperMix (Thermo Fisher Scientific) in a 25 µL volume containing PCR SuperMix (9.5 µL ddH2O, 12.5 µL AccuStart), 0.5 µL of each primer (10 pmol µL−1) and 2 µL of DNA template. For the COI amplification the forward primer LCOech1aF1 and the reverse primer HCO2198, tailed with M13F and M13R-pUC, respectively (Folmer et al., 1994; Layton et al., 2016), were used. The amplification conditions consisted of an initial denaturation step of 3 min at 94 ∘C, 35 cycles of 30 s at 94 ∘C, 60 s at 42–47 ∘C and 1 min at 72 ∘C, followed by a final extension step of 5 min at 72 ∘C. All PCR products were purified using ExoSAP-IT (Thermo Fisher Scientific). The amplified fragments were sequenced in both directions at Macrogen Europe Laboratory (Amsterdam, the Netherlands).
2.4.2 Alignment, genetic divergence
The obtained COI sequences were searched against the GenBank nucleotide database using BLASTN (Altschul et al., 1990). Forward and reverse sequences for each individual were assembled and edited using Geneious v.9.1.7 (https://www.geneious.com, last access: 30 March 2020; Kearse et al., 2012). The edited COI sequences were aligned using MAFFT v7.308 under the E-INS-i and G-INS-I algorithms (Katoh et al., 2002), while alignments were further manually edited. Our dataset was supplemented with 18 COI ophiuroid sequences from the study of Glover et al. (2016). Sequence data are available in GenBank (accession numbers MN088035–MN088083, MT103664–MT103870, MT160426–MT160451). The sequences, trace files, collection data and photos for each specimen are listed in the datasets CCZ_Ophiuroidea (https://doi.org/10.5883/DS-CCZ1, Christodoulou et al., 2020a) and DEA_Ophiuroidea in BOLD (https://doi.org/10.5883/DS-DEA1, Christodoulou et al., 2020b).
2.5 Putative species delimitation
Congruent support across a range of species delimitation approaches assumedly provides more reliable results than a single method (Carstens et al., 2013; Fontaneto et al., 2015). Therefore, five different species delimitation analyses, including both distance- and tree-based approaches, were performed on the COI dataset, to allocate sequences into genetic species (secondary species hypotheses, SSHs; Puillandre et al., 2012; Boissin et al., 2017). Distance-based approaches detect the distance at which the “barcode gap” occurs and sort the sequences into putative species based on this distance, whereas tree-based approaches use a phylogenetic tree from which the fit of speciation and coalescent processes is modelled to delineate species based on the branching rate of the tree (Carstens et al., 2013; Tang et al., 2014).
2.5.1 Distance-based approaches
A neighbour-joining tree was constructed in MEGA7 using a p-distance substitution model, treating gaps and missing data with “pairwise deletion” and by running 1000 bootstrap replicates. Automatic barcode gap discovery (ABGD) analysis was implemented on the web interface http://wwwabi.snv.jussieu.fr/public/abgd/ (last access: 26 July 2019) with default parameters, under the p-distance model (Puillandre et al., 2012). Barcode index numbers (BINs) were assigned to the registered DNA dataset automatically using the BOLD v.4 workbench (http://www.boldsystems.org, last access: 14 October 2019; Ratnasingham and Hebert, 2013).
2.5.2 Maximum-likelihood tree
The COI barcode data available for all 300 samples were adequate to show genetic diversity patterns within and among closely related species but were not sufficient to accurately reconstruct relationships and genetic distances among the many divergent lineages in this biota. Hence, to provide an all-barcode-sample maximum-likelihood tree better reflecting these divergences, the barcode samples were appended to a powerful phylogenetic framework: the 48 475-site exon-28SrDNA-COI super-matrix dataset used by Christodoulou et al. (2019). This dataset comprised 200 species that outlined the ophiuroid family-level phylogeny (O'Hara et al., 2017) and 49 CCZ-DEA barcode samples with both COI and 28S sequences included (Christodoulou et al., 2019). The COI and 28S allowed the barcode-only samples to be linked to the phylogenomic exon data. A maximum-likelihood tree was constructed by IQ-TREE 1.6.9 (Nguyen et al., 2015; Hoang et al., 2018) using a five partition (exon codon positions, 28S, COI) HKY+G model and 1000 ultra-fast bootstrap replicates (with nearest-neighbour interchange (NNI) optimization). Then the 200 “super-matrix backbone” samples were pruned out to leave only the 300 barcode samples, node support bootstrap values were recalculated and the tree was rooted according to O'Hara et al. (2017).
2.5.3 Tree-based approaches
The general mixed Yule–coalescent (GMYC; Pons et al., 2006) method was implemented using the R package SPLITS (Fujisawa and Barraclough, 2013), under the single-threshold model (stGMYC), and with the required ultrametric tree being produced in BEAST v.2.5. Settings were as follows: strict clock, Yule speciation model, GTR+G substitution site model, two independent Markov chain Monte Carlo (MCMC) runs for 50 000 000 generations, sampling every 1000 steps (10 % was discarded as burn-in period). The multi-rate Poisson tree processes (mPTPs; Kapli et al., 2017) analysis used the rooted super-matrix backbone IQ-TREE phylogeny (see above). The mPTP was implemented on the web server https://mptp.h-its.org (last access: 6 August 2019) using the multi-rate Poisson tree process model and following default settings.
2.6 Genetic distances
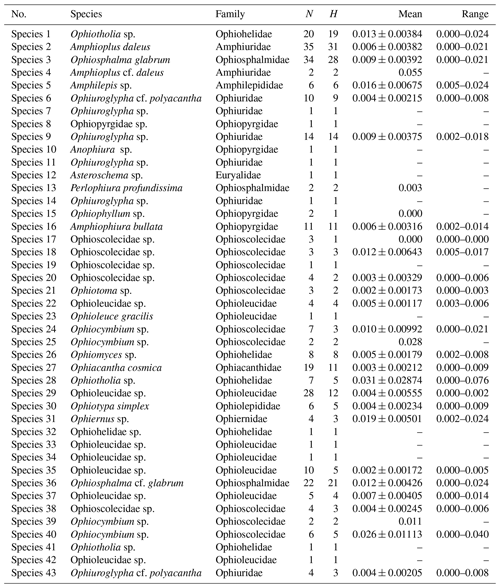
Sequence divergences (Tables 1 and S1–S2 in the Supplement) were estimated using uncorrected p distances and under the K2P model using MEGA7 according to the secondary species hypotheses.
2.7 Assemblage structure and diversity analyses
Comparison of the ophiuroid assemblages between areas was performed in R using the package “vegan” (Oksanen et al., 2008). As the sampling effort was very different between areas, the species composition table (Table 2), including the specimens of each species found in each area, was subjected to “chord” transformation to explore differences in relative abundance and “presence–absence” transformation related to faunistic differences. After transformation non-metric multidimensional scaling (nMDS) ordination was achieved with Euclidean distance (Legendre and Gallagher, 2001). As the number of specimens found differs greatly between areas, diversity comparison was achieved using rarefaction curves, together with standard Shannon (H′), Simpson (D) and Jaccard's evenness (J) diversity indices. The expected number of species per area was inferred using the extrapolation methods Chao1 (Chao, 1984; Colwell and Coddington, 1994) and ACE (Chazdon et al., 1998). Chao1 uses the proportions of singletons and doubletons in the sample to estimate expected species richness, while ACE is an abundance-based coverage estimator. For the analysis of beta (regional) diversity, the total multiple-site beta diversity βSOR was calculated using the modified Sørensen index (Sørensen, 1948; Baselga and Orme, 2012), and βSOR was decomposed into its additive components “multiple-site species turnover” βSIM (Simpson index: Simpson, 1943) and “multiple-site nestedness” βSNE using the R package “betapart” (Baselga, 2010; Baselga and Orme, 2012). In order to explore the relative contribution of every area to species turnover and nestedness, these values were calculated taking one area out each time in a jackknife approach. Changes in turnover and nestedness were then attributable to the area that was excluded from the analysis.
3.1 Species delineation
The species delineation dataset was comprised of 300 barcode sequences (Fig. 2), out of which 287 were novel sequences (BOLD datasets: CCZ_Ophiuroidea, DEA_Ophiuroidea), ranging from 547 to 657 bp in length (92 % have a length of 657 bp).
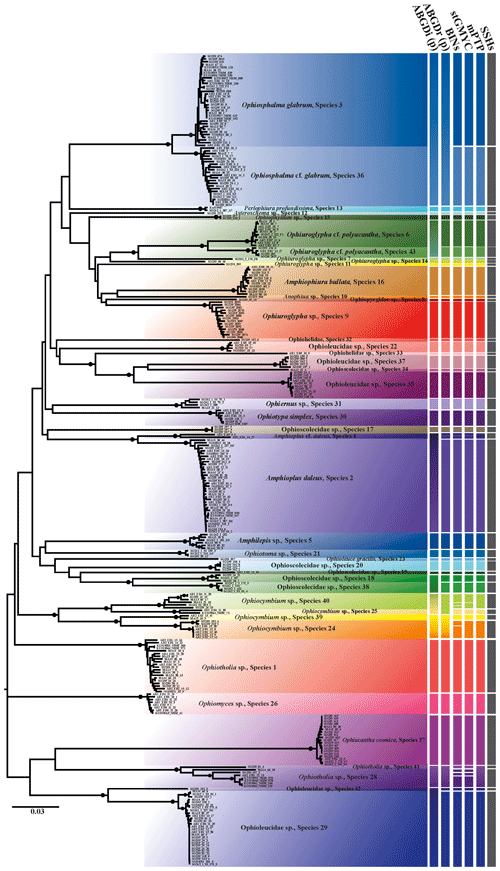
Figure 2Neighbour-joining tree (p distance) based on 300 brittle star COI DNA barcodes. Black circles on branches represent bootstrap supports ≥90 %. The results of species delimitation analyses (ABGD, BINs, stGMYC and mPTP) are shown on the right-hand margin of the tree.
Trees produced by both neighbour-joining (NJ, Fig. 2) and maximum likelihood (ML, Fig. 3) showed a broad pattern in which SSHs were separated by long branches, while branches within species were shorter. The 300 DNA barcodes clustered into 42 monophyletic clades in NJ and into 40 in ML, supported by high bootstrap values (>90).
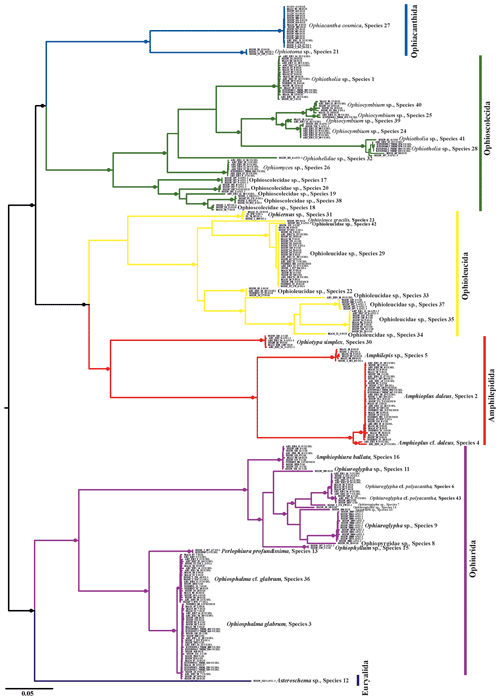
Figure 3Maximum-likelihood phylogenetic tree based on 300 brittle star COI DNA barcodes calculated using an IQ tree. Black circles on branches represent bootstrap support (≥90 %).
The ABGD analysis yielded a total of 35 SSHs based on initial partitioning over the range of prior values for maximum intraspecific divergence (Figs. 2 and S1 in the Supplement). Identical results were produced based on JC69 and K80 corrected distances. The number of SSHs varied between 37 and 50 after the application of recursive partitioning. Low threshold values of 0.0010–0.0028 and 0.0046–0.0077 prompted 50 and 47 SSHs, respectively (Fig. S1). Moderate threshold values of 0.0129 and 0.0215 resulted in 43 and 42 SSHs, respectively (Fig. S1). Finally, higher prior threshold values of 0.0359–0.0599, and 0.1000 provided 40 and 37 SSHs, respectively (Fig. S1). To be conservative, we focus primarily on the results of initial partitioning (35 SSHs) as they were consistent across the parameter settings and congruent with other species delimitation methods (Puillandre et al., 2012; Kekkonen and Hebert, 2014). Nevertheless, for comparative reasons, the results of the recursive partition with prior divergence 0.0359–0.0599 and which suggested 40 SSHs are also presented here (Figs. 2, S1).
In BOLD, the 300 barcodes were assigned to 49 BINs (Fig. 2), of which 22 BINs had a single record and three BINs had two records (CCZ_Ophiuroidea DEA and Ophiuroidea datasets, BOLD).
Single-threshold general mixed Yule–coalescent calculations (stGMYC) yielded 47 SSHs (entities) with a confidence interval ranging from 46 to 49 (Supplement, Result of GMYC).
The mPTP model produced a more conservative number of clusters (42 SSHs) compared to the GMYC method (Supplement, Results of mPTP).
Depending on the applied method, the numbers of different putative species ranged from 35 to 49. Arranging the implemented methods by increasing conservativeness gives the following: BINs (49) < stGMYC (47) < mPTP (42) < ABGD (35). In the present study a consensus dataset of species that were delineated by at least three of the four above-mentioned approaches was selected, as species delineation methods tend to overestimate the number of species present in a dataset. In the few cases that the methods were inconsistent, the most conservative approach was adopted after taking into account the genetic distance between the potential species. The results were cross-referenced with the topology produced by both the NJ and ML trees. It is worth mentioning that 27 SSHs were congruent throughout all methods and 34 SSHs were consistent when excluding ABGDi (initial), which was the most conservative method. In total 43 SSHs were recovered from the CCZ and the DEA, of which some were PSHs split from two up to five SSHs each. Noticeably, the PSHs Amphioplus daleus, Ophiuroglypha cf. polyacantha, Ophiosphalma glabrum and Ophiocymbium sp. revealed cryptic lineages between their populations in the CCZ and the DEA. The 43 SSHs (Figs. 5–16) are grouped in 11 families, Amphilepididae, Amphiuridae, Euryalidae, Ophiernidae, Ophiohelidae, Ophiolepididae, Ophioleucidae, Ophiopyrgidae, Ophioscolecidae, Ophiosphalmidae and Ophiotomidae, attributed to all the current ophiuroid orders (Fig. 3), Amphilepidida, Euryalida, Ophiacanthida, Ophioscolecida and Ophiurida (see also Taxonomic and systematic remarks, Supplement).
3.2 Genetic distances
Summaries of uncorrected pairwise distances for the ophiuroid species (SSHs) are shown in Table 1 and Fig. 4, with the full data available in the Supplement (Tables S1–S2). Mean interspecific genetic distances ranged from 0.050 to 0.370 (p distance) and 0.052 to 0.512 (K2P distance) with the lowest divergence value observed between Ophiosphalma glabrum and Ophiosphalma cf. glabrum (Species 3 vs. 36) and the highest between Ophiacantha cosmica (Species 27) and Ophiohelidae sp. (Species 32). Mean intraspecific variability ranged from 0.00 to 0.055 (p distance) and 0.00 to 0.057 (K2P distance), with the highest values observed in the ophiuroid Amphioplus cf. daleus. It should be mentioned that there were 13 SSHs represented by only one sample (singletons).
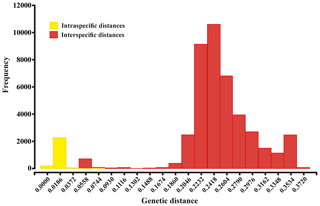
Figure 4Histogram showing the percentage of genetic p distances within and between brittle star species based on the 657 bp “barcode” fragment of the COI gene. Intraspecific and interspecific variations are shown in yellow and red, respectively.
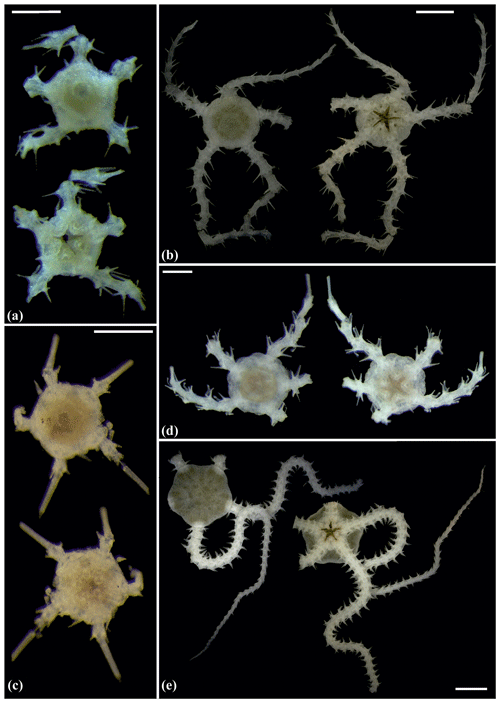
Figure 5Amphilepis sp. (sp5): (a) dorsal and ventral view, MA13_85_32; (b) dorsal and ventral view, MA14_39_9. Amphioplus (Unioplus) daleus (sp2): (c) dorsal and ventral view, AB2_EB1_16_13; (d) dorsal and ventral view, AB1_EB5_10_4; (e) dorsal and ventral view, SO239_81_07. Scale bars: 0.5 mm (a, c–d); 1 mm (b); 2 mm (e).
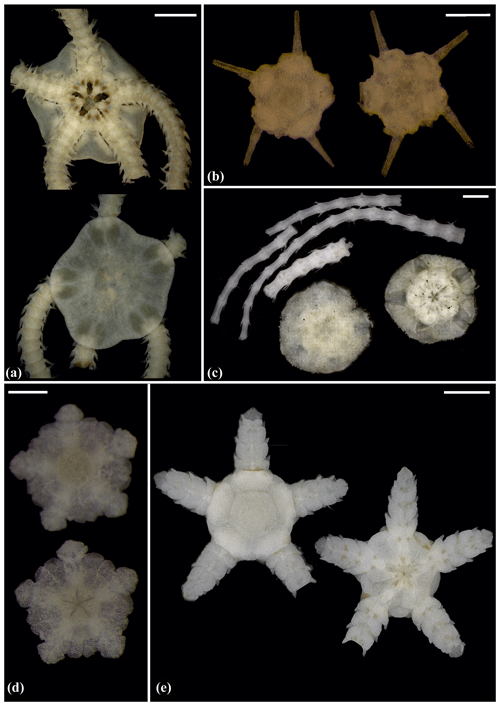

Figure 6Amphioplus (Unioplus) daleus (sp2): (a) dorsal and ventral view, MA14_38_01. Amphioplus (Unioplus) cf. daleus (sp4): (b) dorsal and ventral view, AB2_EB1_14_27. Ophiernus sp. (sp31): (c) dorsal and ventral view of disc and detached arms, MA14_21_12. Ophiotypa simplex (sp30): (d) dorsal and ventral view, AB2_EB2_12_3; (e) dorsal and ventral view, AB1_EB5_4. Scale bars: 0.5 mm (a, d); 2 mm (b); 1 mm (c, e).
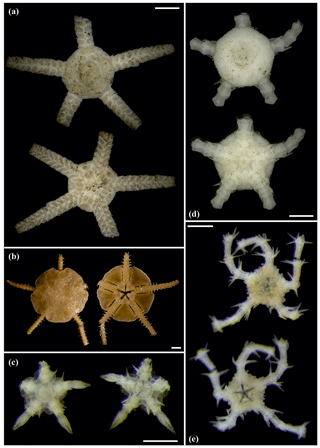

Figure 7Ophiotypa simplex (sp30): (a) dorsal and ventral view, SO239_118_1. Ophioleuce gracilis (sp23): (b) dorsal and ventral view, SO239_397. Ophioleucidae sp. (sp22): (c) dorsal and ventral view, SO239_24_17; (d) dorsal and ventral view, SO239_59_1. Ophioleucidae sp. (sp29): (e) dorsal and ventral view, SO239_24_12. Scale bars: 2 mm (a, b); 1 mm (d); 0.5 mm (e, c).
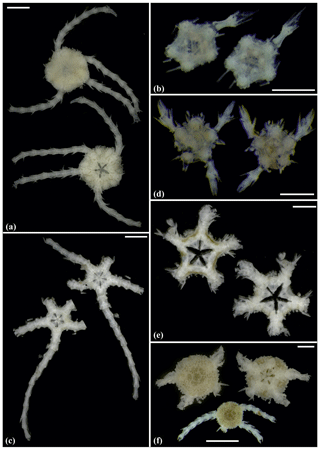
Figure 8Ophioleucidae sp. (sp29): (a) dorsal and ventral view, SO239_24_3. Ophioleucidae sp. (sp33): (b) dorsal and ventral view, AB1_EB5_10_9. Ophioleucidae sp. (sp34): (c) dorsal and ventral view, MA14_21_3. Ophioleucidae sp. (sp35): (d) dorsal and ventral view, SO239_118_14; (e) dorsal and ventral view, SO239_24_5; (f) dorsal and ventral view, SO239_133_2. Scale bars: 1 mm (a, c, e, f); 0.5 mm (b, d).
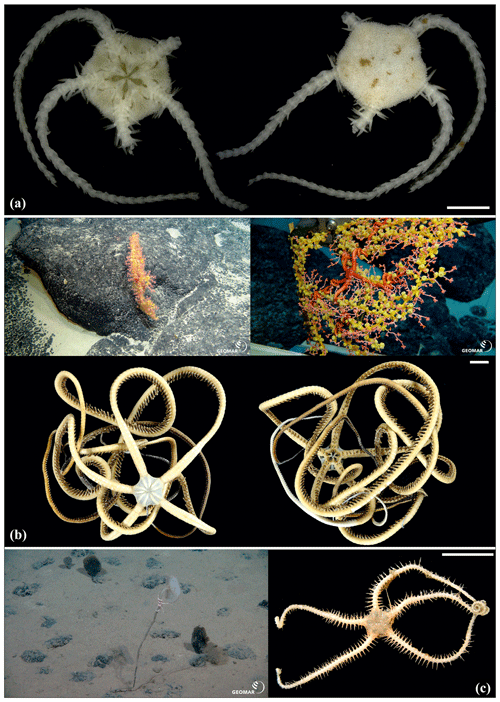
Figure 9Ophioleucidae sp. (sp37): (a) dorsal and ventral view, SO239_139_2. Asteroschema sp. (sp12): (b) in situ (upper left and right), specimen collected with the ROV KIEL 6000 in dorsal (lower left) and ventral (lower right) view, SO239_2113. Ophiocantha cosmica: (c) in situ (left), specimen collected with the ROV KIEL 6000 in dorsal view (right), SO239_130. Scale bars: 2 mm (a); 1 cm (b, c). Copyright (for in situ photos) ROV KIEL 6000 Team/GEOMAR Kiel.
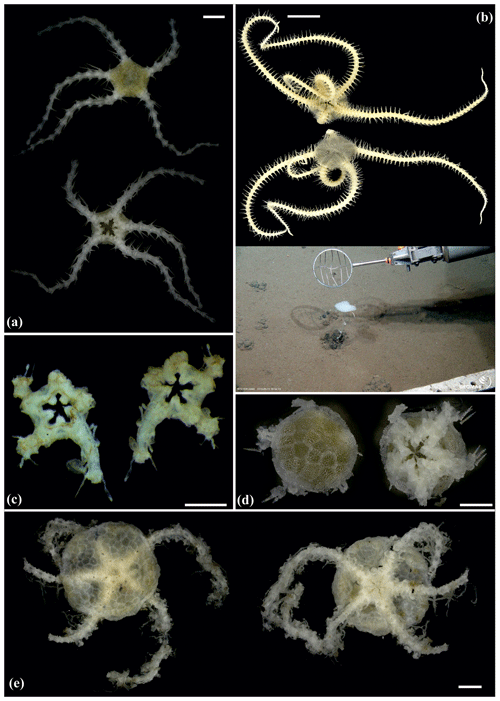
Figure 10Ophiocantha cosmica (sp27): (a) dorsal and ventral view, MA14_20_4; (b) specimen collected with the ROV KIEL 6000 in dorsal and ventral view (up), in situ (down), SO242-2_191_F5. Ophiotoma sp. (sp21): (c) dorsal and ventral view, SO239_20_12. Ophiocymbium sp. (sp39): (d) dorsal and ventral view, AB2_EB1_13_41. Ophiocymbium sp. (sp24): (e) dorsal and ventral view, AB2_EB1_13_8. Scale bars: 0.5 cm (d); 1 mm (a, c); 1 cm (b); 2 mm (e). Copyright (for in situ photos) ROV KIEL 6000 Team/GEOMAR Kiel.
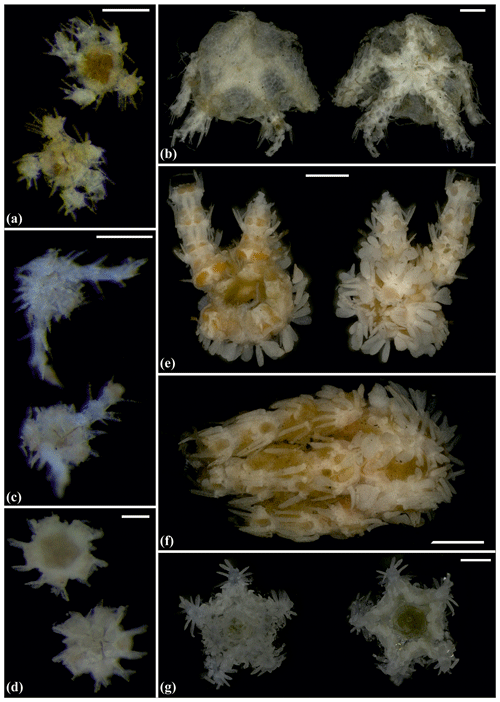
Figure 11Ophiocymbium sp. (sp40): (a) dorsal and ventral view, SO239_24_19; (b) dorsal and ventral view, MA14_21_10. Ophiohelidae sp. (sp32): (c) dorsal and ventral view, SO239_192_06. Ophiomyces sp. (sp26): (d) dorsal and ventral view, AB1_EB5_ 10_3; (e) dorsal and ventral view, AB1_EB4_11_24; (f) lateral view, AB1_EB4_11_22. Ophiotholia sp. (sp1): (g) dorsal and ventral view, MA14_38_13. Scale bars: 0.5 mm (a, c); 1 mm (b, e, f); 0.2 mm (d).
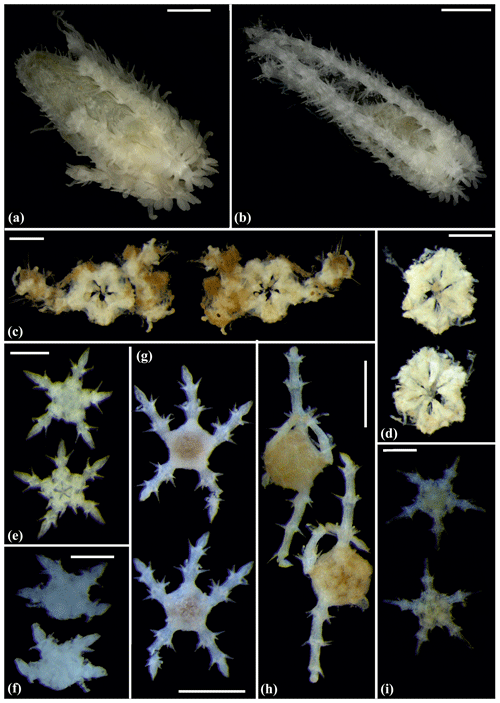
Figure 12Ophiotholia sp. (sp1): (a) lateral view, MA13_85_3; (b) lateral view, MA13_90_18. Ophiotholia sp. (sp28): (c) dorsal and ventral view, MA14_66_10. Ophioscolecidae sp. (sp17): (d) dorsal and ventral view, SO239_197_ 4. Ophioscolecidae sp. (sp18): (e) dorsal and ventral view, SO239_24_21. Ophioscolecidae sp. (sp19): (f) dorsal and ventral view, AB2_EB2_ 12_ 10. Ophioscolecidae sp. (sp20): (g) dorsal and ventral view, SO239_192_2; (h) dorsal and ventral view, SO239_192_8. Amphiophiura bullata (sp16): (i) dorsal and ventral view, SO239_118_13. Scale bars: 1 mm (a–d, g, h); 0.5 mm (e–f, i).
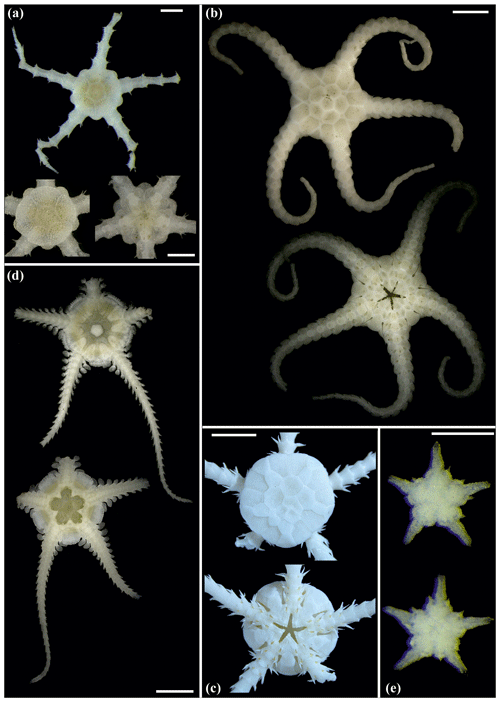
Figure 13Amphiophiura bullata (sp16): (a) dorsal view (upper) and dorsal (lower left) and ventral view (lower right) of disc, SO239_133_3; (b) dorsal and ventral view, MA13_90_16. Anophiura sp. (sp10): (c) dorsal view and detailed dorsal and ventral view of disc, SO239_396. Ophiophyllum sp. (sp15): (d) dorsal and ventral view, SO239_139_1. Ophiopyrgidae sp. (sp8): (e) dorsal and ventral view, SO239_59_10. Scale bars: 0.5 mm (a, d); 2 mm (b); 1 mm (c, e).
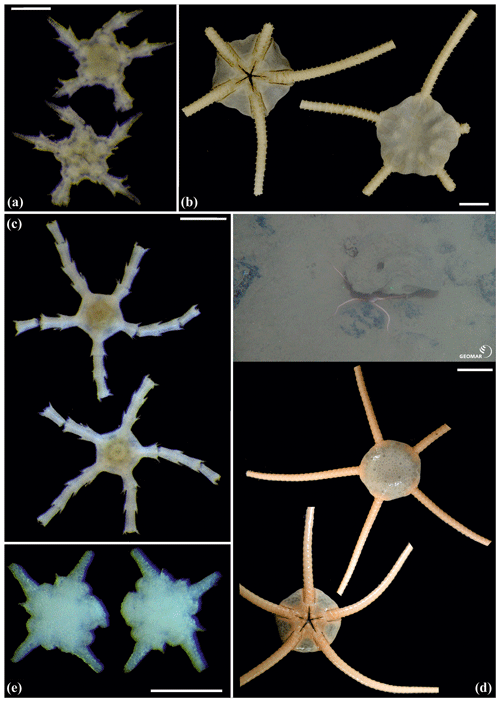
Figure 14Ophiuroglypha cf. polyacantha (sp6): (a) dorsal and ventral view, MA14_66_7; (b) dorsal and ventral view, SO242-2_222_F1. Ophiuroglypha cf polyacantha (sp43): (c) dorsal and ventral view, SO239_118_6; (d) in situ (right), dorsal and ventral view (left), SO239_2037. Ophiuroglypha sp. (sp14): (e) dorsal and ventral view, SO239_59_18. Scale bars: 0.5 mm (a, e); 1 mm (c); 1 cm (b, d). Copyright (for in situ photos) ROV KIEL 6000 Team/GEOMAR Kiel.
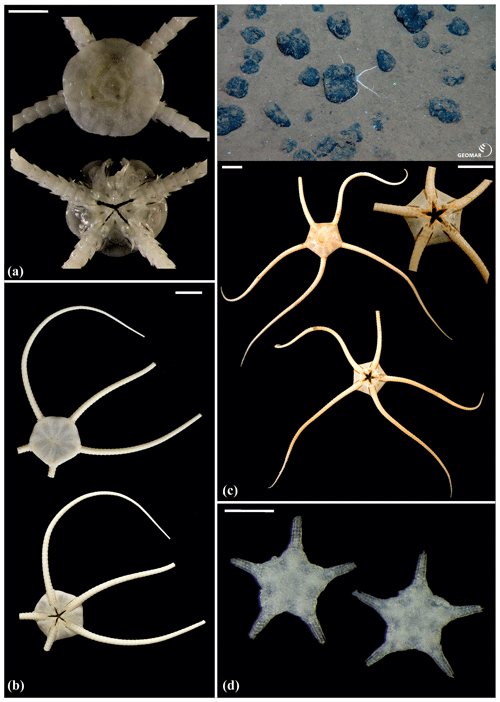
Figure 15Ophiuroglypha sp. (sp11): (a) dorsal and ventral view, SO239_395. Ophiuroglypha sp. (sp7): (b) dorsal and ventral view, SO242-2_176_F8. Ophiuroglypha sp. (sp9): (c) in situ (upper right), dorsal and ventral view, detailed ventral view of disc (lower right), SO239_2059. Ophiosphalma glabrum (sp3): (d) dorsal and ventral view, AB1_EB5_10_7. Scale bars: 1 cm (a–e); 0.5 mm (d). Copyright (for in situ photos) ROV KIEL 6000 Team/GEOMAR Kiel.
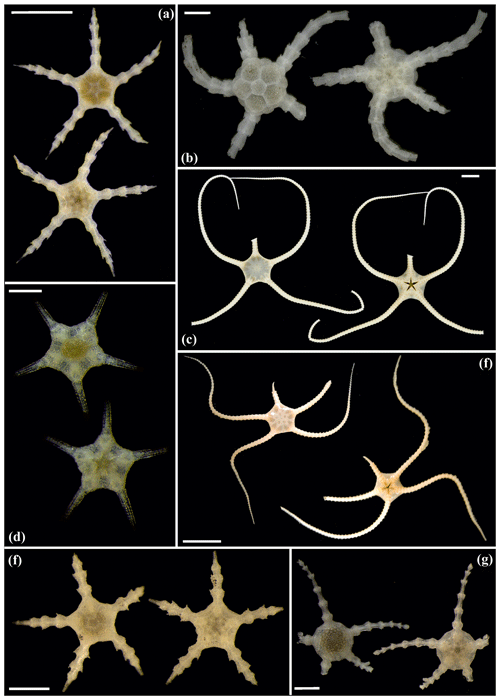
Figure 16Ophiosphalma glabrum (sp3): (a) dorsal and ventral view, SO239_24_4; (b) dorsal and ventral view, SO239_50_2; (c) dorsal and ventral view, SO242-2_222_F2. Ophiosphalma cf. glabrum (sp36): (d) dorsal and ventral view, SO239_24_18; (e) dorsal and ventral view, AB2_EB1_14_2; (f) dorsal and ventral view, SO239_2014. Perlophiura profundissima: (g) dorsal and ventral view, SO242-1_387_A7. Scale bars: 0.5 mm (d); 1 mm (b, e, g); 2 mm (a, e); 1 cm (c, f).
3.3 Ophiuroid assemblages and diversity
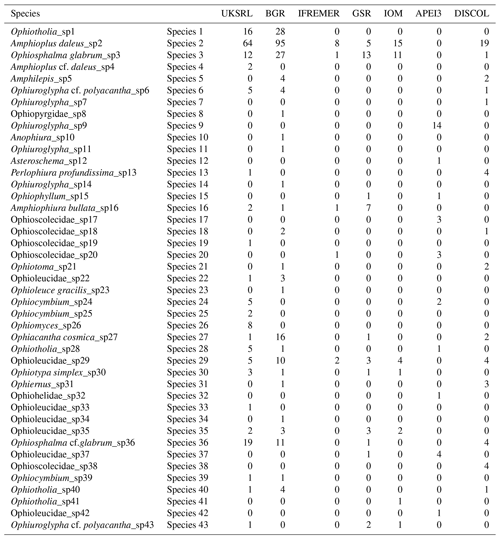
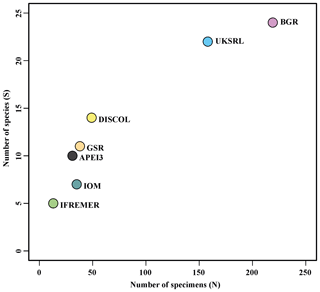
The species composition table (Table 2) shows the counts of each species by area. The diversity values are summarized in Table 3. A total of 55 sites were sampled in seven areas. Sampling effort was uneven, with most samples deriving from the BGR area (18) and the DEA (14) in the Peru Basin. For all other areas, three to six sites were sampled. A total of 543 specimens were assigned to the 43 species. None of the species was recorded in all seven areas, while the most common species were Species 29 (Ophioleucidae), Species 2 (Amphioplus daleus) and Species 3 (Ophiosphalma glabrum), which were found in six areas, with all of them absent in APEI3. It is worth mentioning that the majority of species (44.2 %) were present only in one of the areas (19 SSHs). The highest species numbers were found in the BGR and UKSRL areas (24 and 22, respectively), where the highest number of specimens was also recorded (219 and 158, respectively). The lowest values were found in the IFREMER area, with 13 specimens being attributable to five species. While the number of species was a function of the number of specimens, fewer species were recorded in the IFREMER and IOM areas than would be expected if they were to follow the same pattern as at other sites (Fig. 17). This was corroborated by the rarefaction analysis (Fig. 18), which shows that for the same number of specimens, the IFREMER and IOM areas have fewer species. The rarefaction curves of all other areas were very similar. Low diversity in the IFREMER and IOM areas was also indicated by the lowest Shannon diversity, Simpson diversity and evenness values, while the highest diversity values were recorded in the UKSRL, BGR and DISCOL areas (Table 3). The extrapolation analyses predicted a total of 57 species (Chao1 index) and 53.5 species (ACE index) for all areas together. The lowest extrapolated numbers of species were again obtained for the IFREMER and IOM areas (6.5–12 and 8.5–11.5, respectively), whereas the highest numbers were obtained for the BGR and UKSRL areas (57–51.1 and 27.2–30.5, respectively). The highest number of unique species (species found only in one area) was found in the BGR (six species), UKSRL (five species) and APEI3 (five species) areas, while no unique species were observed in the IFREMER and GSR areas.
Table 3Summary of diversity parameters per sampled area. Sites: number of collection sites; N: number of specimens; S: number of species; Usp: number of unique species; Chao ± SE: Chao estimated number of species with standard error; ACE ± SE: ACE estimated number of species with standard error; H′: Shannon diversity; 1-D: Simpson diversity; J: Jaccard's evenness. βSOR, βSIM and βSNE express multiple-site total beta diversity, multiple-site species turnover and multiple-site nestedness, respectively. Note that in the rows of each area the value is the result of excluding this area, except for the row “Total”, which includes all areas.
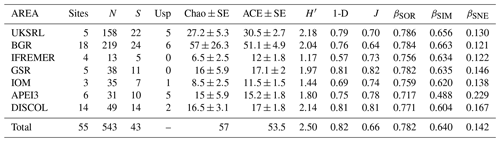
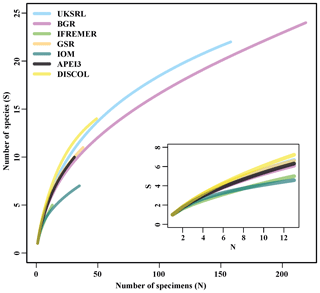
Figure 18Sample-based rarefaction curves of the examined areas. The inset shows a close-up for the minimum shared number of specimens (12).
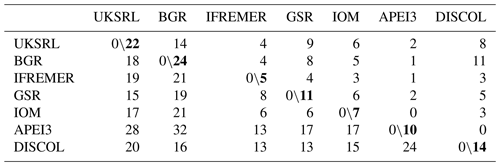
The faunistic similarity is summarized in Table 4, showing the number of shared and unshared species between areas. APEI3 showed the lowest numbers of shared species (zero to two) and the highest number of unshared species (13–32) compared with other areas. The most distant area, DISCOL in the Peru Basin, shared 3–11 species with CCZ exploration areas, but none with APEI3.
Table 4Faunistic similarity between areas. Left of slash: number of unshared species; right of slash: number of shared species. Bold numbers indicate the number of species in each area.
Beta diversity decomposition is shown in Fig. 19. The total multiple-site beta diversity was high (βSOR=0.782), with a higher component of turnover (βSIM=0.640) versus nestedness (βSNE=0.142). To explore the relative contribution of each area to total beta diversity, each area was taken out once and beta diversity was recalculated. The relative change in turnover and nestedness was then attributable to the omitted area. Results of this exercise are shown graphically in Fig. 19 and numerically in Table 3. Removing most of the areas one by one (excluding APEI3) did not result in a drastic change in turnover and nestedness (βSIM = 0.604–0.663; βSNE = 0.121–0.167). Only the exclusion of APEI3 resulted in a substantial reduction of turnover and increase in nestedness (βSIM=0.488; βSNE=0.229).
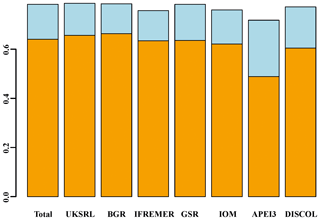
Figure 19Resulting beta diversity, when each area is excluded from calculation, decomposed into its additive component species turnover (light blue) and nestedness (orange). Excluding APEI3 has the greatest impact on beta diversity. “Total” shows the values including all areas.
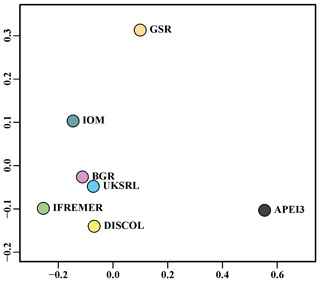
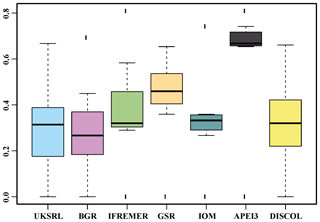
The nMDS plot in Fig. 20 shows the quantitative assemblage analysis using chord distance (relative abundance). The BGR and UKSRL areas were close together but also close to the areas of DISCOL, IFREMER and IOM, while greater dissimilarity occurs with the GSR and APEI3 areas. The box plot in Fig. 21 shows the variation in chord distance of each area to other areas, evidencing that APEI3 was most different from any other area (see median and extent of whiskers) than other areas were among each other.
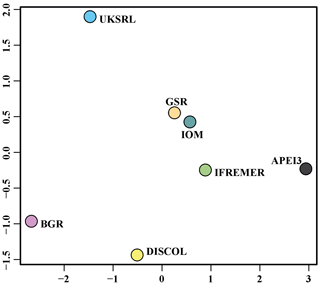
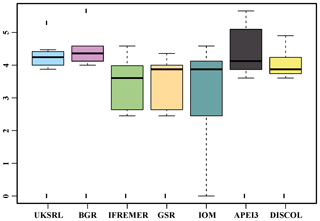
The ordination using presence–absence-transformed data placed the areas with fewer unique species (IFREMER, GSR and IOM) in the middle of the plot and spread the areas with the highest number of unique species at the outer margins and apart from each other (Fig. 22). The box plot in Fig. 23 shows that APEI3 was the most dissimilar in terms of presence–absence of species, but the median value (black horizontal bar inside the boxes) was as high as that for the UKSRL and BGR areas, which, however, displayed less variation.
4.1 Species delimitation method performance
The results obtained here were consistent with many other studies showing that different species delimitation methods can produce different delimitation scenarios when employing single-locus data (Hofmann et al., 2019). The single-locus species delimitation methods tested here, although they are extensively used throughout the literature, including for the Ophiuroidea (Khodami et al., 2014; Laakman et al., 2016; Boissin et al., 2017), are each subject to potential biases and differing conditions inherent in the empirical datasets (Hofmann et al., 2019). The five species delimitation methods used here generally recovered the same number of SSHs. despite some degree of incongruence observed in the numbers of SSHs, they were consistent in recovering more SSHs than the number of species originally recognized. Given the lack of information regarding the biodiversity and of the relationships between deep-sea ophiuroids, it was not surprising that more lineages were inferred than are currently recognized. It is likely that many of these SSHs correspond to undescribed cryptic species, but simultaneously some may be the result of genetic drift or isolated populations currently undergoing speciation. Noticeably, the BIN method in BOLD recovered a higher number of species than all other approaches. BOLD and specifically BINs can greatly improve the Linnaean taxonomic assignment in many animal groups, including echinoderms (Layton et al., 2016; Laakman et al., 2016). The low intra-cluster divergence (2.2 %) at the initial cluster step of refined single linkage (RESL) methodology (Ratnasingham and Hebert, 2013; Song et al., 2018) could be the reason why in some cases the BIN method overestimated species number, especially since there appears to be a small overlap between intraspecific and interspecific distance in our data (Fig. 4). This could be the case in the delimited Ophiocymbium spp. (Species 24, 25, 40; Fig. 2, Table S2), which were separated into numerous lineages despite the relatively low divergence between them. Generally, barcodes are well defined when the lowest interspecific distance exceeds the highest intraspecific distance, and in such cases a species delineation “threshold” will be clear. But, as the threshold can be lineage-specific, a universal threshold that fits all the branches may not exist, as coalescent depths among species will vary greatly due to differences in population size, mutation rate and speciation times (Colins and Cruickshank, 2012). Similarly, GMYC recovered a relatively high number of species (47 vs. 49 BINs). Arguably, GMYC, especially the single-threshold version of the method, is a robust species delimitation method (Fujisawa and Barraclough, 2013). GMYC performance depends on a single-locus ultrametric tree which tends to compress the coalescent events towards the tips of the tree, making it especially difficult to distinguish closely related species (Boissin et al., 2017). It has been argued that the Poisson tree process (PTP) methods generate diversity estimates that are more robust to different phylogenetic methods, while GMYC is more sensitive, but provide consistent estimates for BEAST trees (Tang et al., 2014). Nevertheless, unresolved nodes can affect both GMYC and PTP estimates, although they seem to have a greater effect on GMYC estimates (Tang et al., 2014). In contrast it seems that ABGD (initial partition) has underestimated the species number in this study, although the performance of the method improved when the recursive partition option was used. ABGD has been reported to over-lump speciose datasets with high speciation rates (Dellicour and Flot, 2018). ABGD's conservatism and GMYC's overestimation have also been shown on reef brittle stars (Boissin et al., 2017) while indicating that PTP methods show a small advantage as the most stable, suggesting the presence of additional cryptic species but without over-splitting taxa. Summarizing, despite the differences in the number of delimited species, overall the methods recovered a broadly similar number of SSHs. Congruence among different delimitation methods is a strong indication that the delimitation is correct, allowing the designation of cryptic species and rectification of taxonomic problems (Dellicour and Flot, 2018), while always when possible taking into account the morphology.
4.2 Taxonomic implications
The abyssal eastern Pacific harbours a highly diverse ecosystem. The number of ophiuroid species reported from the polymetallic nodule fields of the Pacific has now increased by 433 % from 10 (Glover et al., 2016; Amon et al., 2017) to 43 in this paper. This is the largest collection of any megafaunal taxon in the CCZ and the only one that has been studied in such detail using a comprehensive combination of morphological and genetic evidence. Remarkably, from the species reported here, 32 (75 %) are probably new to science, and some represent hitherto unknown old evolutionary lineages (see also Christodoulou et al., 2019). The discovery of new species is the direct result of increased sampling effort, in which a greater number of specimens deriving from a larger sampling surface were collected than during any previous studies in the DEA or in the CCZ, spanning five exploration contract areas and one APEI. Furthermore, the use of new sampling gears, i.e. epibenthic sledge (EBS), in relation to past historical expeditions that took place in the area, permitted the collection of fragile and minute specimens, while new DNA barcoding approaches allowed the identification of post-larvae and juveniles that lacked adult morphological characters. Overall, these data show that the brittle star biodiversity in the deep sea is still greatly underestimated, while supporting the use of DNA barcoding as an effective and time-efficient method of species delimitation to complement morphological studies. Although we do not wish to suggest that single mitochondrial locus data should be used on their own to draw taxonomic conclusions, in much the same way as using single morphological characters is discouraged (DeSalle, 2006; Hofmann et al., 2019), we do argue that single gene barcoding could be the first step in identifying previously overlooked species, while also providing a guide in cases where morphological identification is difficult. It was only recently that a transcriptome-based analysis of Ophiuroidea (O'Hara et al., 2014) instigated a major re-evaluation of morphology-based classifications (Smith et al., 1995), proving that there is still a lot to be discovered and re-evaluated within this group. Specifically in our study, DNA barcoding proved to be necessary since a significant proportion of the specimens are post-larvae juveniles, making their identification based on morphological characteristics quite challenging. DNA barcoding not only allowed species delimitation but also aided in matching post-larvae individuals with their corresponding adults, as for example in the case of Ophiosphalma glabrum where individuals ranging from 0.5 mm to 20 cm were collected (Fig. 23). Furthermore, the large-sized brittle stars collected with the remotely operated vehicle were matched with their in situ photos, allowing a more accurate estimation of morphospecies which in turn could facilitate the more accurate annotation of photos and video transects used in various biodiversity assessment studies (Tilot et al., 2018).
Mean COI genetic intraspecific distances (K2P) of brittle stars (0.00–0.057) were concordant with previous ophiuroid studies (0.00–0.042: Khodami et al., 2014 and 0.00–0.064: Boissin et al., 2017), while the mean COI interspecific genetic distances (0.052–0.512) were found to be noticeably higher. This could be attributed both to the great phylodiversity of ophiuroids collected from the polymetallic fields, spanning 11 families and five orders, and to the discovery of previously undescribed diversity up to the family level (Christodoulou et al., 2019).
4.3 Ophiuroid diversity and assemblage structure and implications for conservation in light of possible nodule mining
Mining of abyssal polymetallic nodules in the CCZ could result in severe habitat disruption and loss of benthic communities in and directly around the mined sites (Vanreusel et al., 2016). An attempt to foresee the potential impact of deep-sea mining on abyssal communities requires a profound knowledge of natural background biodiversity and ecosystem functioning, such as how many and which species are living there now, how large the species ranges are, and whether there are natural changes in diversity along environmental gradients. However, our knowledge of abyssal benthic communities is still so poor that even these simple questions remain unanswered for many groups of organisms in what is nonetheless considered to be an economically important and potentially endangered deep-sea region like the CCZ. We can now provide partial answers to these questions for the ophiuroids, one of the dominant megafaunal groups in the CCZ.
Our initial assumption was that ophiuroid diversity would be low (we expected around 10 species) based on the previous studies in the region (Glover et al., 2016; Amon et al., 2017) and on a recent review of global ophiuroid distribution, in which only 28 species were recorded for the whole tropical east Pacific at abyssal depths (Stöhr et al., 2012). Coupled with expectations of low diversity, we assumed that connectivity would be high and that most beta diversity between sites would be composed of nestedness (high) rather than species turnover (low). Under these circumstances, the APEI3 could be a good region to host most of the CCZ species and serve as a source for most of the populations. Also, we assumed that the most distant DISCOL area would display a low similarity to the CCZ.
The results of our study do not support any of these initial assumptions; in fact, they show exactly the opposite. We recorded a 4-fold higher number of species than expected, and rarefaction curves show no sign of reaching an asymptote (Fig. 18). Chao1 and ACE estimators predicted between 53 and 57 species across the region (Table 3). The ophiuroid communities were characterized by high beta diversity that is mainly composed of high turnover between areas, rather than nestedness. This means that there was a high proportion of rare species (19 species were present in only one area and 12 species were present in only two areas), which reduces nestedness and increases the potential for damage to natural populations caused by deep-sea mining at the local scale. Food availability regulated by particulate organic carbon (POC) flux seems to strongly influence diversity and abundance in abyssal ecosystems (Smith et al., 2008). The CCZ licence areas for exploration despite being all in a mesotrophic zone are not all the same. The POC flux in the CCZ shows a southeast-to-northwest gradient increasing towards its eastern edge (Vanreusel et al., 2016; Volz et al., 2018, 2020). Volz et al. (2018, 2020), after studying four of the CCZ areas studied herein and the APEI3, found that they differ in POC fluxes to the sea floor ranging from as low as 1 mg Corg m−2 d−1 in APEI3 to 2 mg Corg m−2 d−1 in the BGR area. Within this study the highest diversity values were recorded in the UKSRL, BGR and DISCOL areas. The BGR and UKSRL areas are located in the far east of the CCZ, in a region of higher POC flux (Vanreusel et al., 2016; Volz at al., 2018, 2020), which could explain higher standing stocks and higher species diversity and abundance. Food availability seems to justify why the communities of the very distant and eutrophic DISCOL area (Haeckel et al., 2001) resemble the eastern CCZ more than the geographically closer APEI3. The DISCOL area shares 11 species with the BGR and eight species with the UKSRL area, while no species are shared with APEI3. In contrast to the CCZ areas, APEI3 lies within an oligotrophic zone, exhibiting 2-fold lower POC fluxes and subsequently 2-fold lower aerobic respiration rates in comparison to the BGR area (Volz et al., 2018, 2020). APEI3 differs significantly from the other areas in additional aspects such as lower chloroplastic pigment equivalents (CPEs) and total organic carbon (TOC) values and lower sedimentation rates resulting in finer sediments with higher clay content (Hauquier et al., 2019; Volz et al., 2018, 2020). In conclusion the APEI3 biogeochemical features differ considerably from the other areas (Volz et al., 2018, 2020) and could explain the fact that APEI3 has a very different assemblage sharing only up to two species with the CCZ areas and DISCOL. Furthermore, APEI3 is mainly located outside the CCZ, north of the Clarion Fracture, a submarine mountain range characterized by a peak and trough surpassing 1800 m difference in elevation (Hall and Gurnis, 2005), which may act as a dispersal barrier for abyssal fauna.
This raised the critical question of whether the APEI3 fauna is representative for the exploration licence areas in the CCZ, especially as ISA created the APEIs based on environmental conditions but in the absence of any biological data (Wedding et al., 2013). Our results suggest that APEI3 may not be a good surrogate area for the CCZ nodule fauna. Only a small fraction of the total registered ophiuroid fauna was recorded in APEI3. This area is the most different in terms of species composition and assemblage structure (Figs. 17, 19). Removing the APEI3 from beta analysis results in a great reduction of turnover and increase in nestedness. This means that the remaining areas become more similar to each other, the total beta diversity decreases and differences between sites due to missing species out of a common species pool (nestedness) increase when APEI3 is excluded. Lower species richness and abundance in APEI3 as well as very low similarity between the APEI3 and the exploration areas, independent of distance, were also found by Vanreusel et al. (2016), Hauquier et al. (2019) and Bonifácio et al. (2020) after studying megafaunal, nematode and polychaete assemblages, respectively. Volz et al. (2018, 2020), after studying the (bio)geochemical characteristics of APEI3 as well as four eastern CCZ areas, concluded that the preservation area APEI3 does not represent the depositional conditions and biogeochemical processes that are dominating in the investigated CCZ licence areas. The results of these studies converge with ours in finding that APEI3 is ill-suited as a representative area of the recovery of the potentially mined areas. Furthermore, Taboada et al. (2018) found that APEI6 is inadequate to act as a population source for a hexactinellid sponge species and suggest the designation of a new APEI closer to the exploration areas studied. The large geographic distance between the APEIs and the explorations areas may hinder the exchange of individuals and the genetic flow among remaining populations after mining. Therefore, we strongly advocate in favour of incorporating no-mining sites within the core CCZ area with a similar nodule composition as the potential mining areas, rather than only on the periphery of the CCZ, as was already suggested by Vanreusel et al. (2016), in order to prevent the loss of biodiversity.
Biodiversity studies that focus only on known, nominal species are problematic, as they likely overlook cryptic or undiscovered lineages involved in diversification. As shown herein, most of the brittle stars recorded in the CCZ and Peru Basin lack formal Linnaean scientific names, thus widening the gap between described species and actual biodiversity, which appears to be far greater than previously estimated. Not recognizing these cryptic or undescribed taxa ensures that they remain in the shadows of research and conservation policies. These taxa could be locally endemic or rare and thus more vulnerable to human impacts such as deep-sea mining. Biodiversity studies, such as presented here, aiming to develop reference libraries while using an integrative taxonomic approach, such as presented here, will provide much-needed comprehensive and time-efficient assessments of “missed” diversity. These, in turn, may fill the gaps for adequate baseline assessment at the onset of commercial-scale mining and thus, through adequate management schemes, prevent serious species declines before they have been adequately described or even discovered.
In conclusion, it is important to note that the present study explored only a part of the polymetallic fields of the CCZ and DEA. Our dataset on ophiuroid communities in the CCZ is the largest available to date but still too small to allow for comprehensive conclusions. The conclusions for APEI3 cannot be extrapolated to other APEIs in the region. Also, our dataset is biased toward the eastern areas of the CCZ where the sampling effort was higher. Thus, although a large number of specimens were examined, it is highly likely that the true biodiversity is even much higher. Broader efforts, especially those that will include samples from the western parts of the CCZ, or from other APEIs, are likely to result in the discovery of additional diversity and will allow us to obtain a better understanding of connectivity and patterns of distribution across the CCZ. This will in turn refine our perception of the marine biodiversity of the abyssal plains and specifically of polymetallic nodule fields.
Four methods of species delineation showed concordant results and revealed 43 deep-sea ophiuroid species in the Clarion-Clipperton Zone and the DISCOL Experimental Area (Pacific Ocean), revealing an unexpectedly high diversity and showing that the most conspicuous invertebrates in the abyssal plains have been so far considerably underestimated. This study increases the number of ophiuroid species reported from polymetallic nodule fields of the Pacific by 433 %.
A comprehensive reference library including 287 novel ophiuroid sequences allocated to 43 species is produced. This reference library can facilitate the assessment of potential impacts and biodiversity loss due to deep-sea mining. It is the first time such an integrated reference library has been produced for the CCZ and the DISCOL area including both genetic and morphological information about the most emblematic mobile megafaunal inhabitants.
The biodiversity patterns observed within CCZ could be mainly attributed to differences in POC fluxes explaining the higher species numbers found in BGR and UKSRL areas. The five exploration contract areas belong to a mesotrophic province, while in contrast the APEI3 (Area of Particular Environmental Interest) is located in an oligotrophic province, which could explain the lowest diversity as well as very low similarity to the other six study areas.
Based on the results of our study, the representativeness and the appropriateness of APEI3 (Area of Particular Environmental Interest) to meet its purpose of preserving the biodiversity of the CCZ fauna are questioned, and the creation of no-mining sites within the core CCZ area is suggested.
DNA sequences, trace files, collection data and taxonomic remarks are available in the datasets CCZ_Ophiuroidea (https://doi.org/10.5883/DS-CCZ1, Christodoulou et al., 2020a) and DEA_Ophiuroidea (https://doi.org/10.5883/DS-DEA1, Christodoulou et al., 2020b) in BOLD. Furthermore, the DNA sequences and their accompanying collection data are also available in GenBank.
The supplement related to this article is available online at: https://doi.org/10.5194/bg-17-1845-2020-supplement.
PMA and AV designed the sampling. PMA, AV, SK, CFR and AH carried out the sampling and processed the specimens on board. MC performed the sequencing programme. AFH performed the phylogenomic analyses. MC, TO'H, AFH, CFR and PMA analysed and interpreted the data. MC took the lead in writing the manuscript in collaboration with TO'H and PMA, to which all other authors contributed.
The authors declare that they have no conflict of interest.
This article is part of the special issue “Assessing environmental impacts of deep-sea mining – revisiting decade-old benthic disturbances in Pacific nodule areas”. It is not associated with a conference.
The authors would like to thank Carsten Rühlemann (Federal Institute for Geosciences and Natural Resources, BGR, Hanover) for making the material from the BGR cruises BIONOD, MANGAN 2013 and MANGAN 2014 available. Special thanks are due to Nicol Mahnken for her help in photographing the specimens. This is publication number 67 from the Senckenberg am Meer Metabarcoding and Molecular Laboratory.
The cruises SO239 and SO242 were financed by the German Ministry of Education and Science (BMBF) as a contribution to the European project JPI Oceans “Ecological Aspects of Deep-Sea Mining”. This research has been supported by BMBF under contract 03F0707E. The ABYSSLINE cruises were funded by UK Seabed Resources Ltd. CESAM was financially supported by the Portuguese Foundation for Science and Technology, Ministry of Science, Technology and Higher Education (FCT/MCTES), through national funds (UID/AMB/50017/2019). Clara F. Rodrigues was funded by Portuguese national funds (OE), through FCT, I.P., in the scope of the framework contract foreseen in numbers 4, 5 and 6 of article 23 of the decree law 57/2016, on 29 August, changed by law 57/2017, on 19 July.
This paper was edited by Tina Treude and reviewed by Chris Mah and two anonymous referees.
Altschul, S. F., Gish, W., Miller, W., Myers, E. W., and Lipman D. J.: Basic local alignment search tool, J. Mol. Biol., 215, 403–410, https://doi.org/10.1016/S0022-2836(05)80360-2, 1990.
Amon, D. J., Ziegler, A. F., Dahlgren, T. G., Glover, A. G., Goineau, A., Gooday, A. J., Wiklund, H., and Smith C. R.: Insights into the abundance and diversity of abyssal megafauna in a polymetallic-nodule region in the eastern Clarion-Clipperton Zone, Sci. Rep.-UK, 6, 30492, https://doi.org/10.1038/srep30492, 2016.
Amon, D. J., Ziegler, A. F., Kremenetskaia, A., Mah, C. L, Mooi, R., O'Hara, T., Pawson, D. L., Roux, M., and Smith C. R.: Megafauna of the UKSRL exploration contract area and eastern Clarion-Clipperton Zone in the Pacific Ocean: Echinodermata, Biodivers. Data J., 5, e11794, https://doi.org/10.3897/BDJ.5.e11794, 2017.
Baselga, A.: Partitioning the turnover and nestedness components of beta diversity, Global Ecol. Biogeogr., 19, 134–143, https://doi.org/10.1111/j.1466-8238.2009.00490.x, 2010.
Baselga, A. and Orme, C. D. L.: betapart: an R package for the study of beta diversity, Methods Ecol. Evol., 3, 808–812, https://doi.org/10.1111/j.2041-210X.2012.00224.x, 2012.
Boissin, E., Hoareau, T. B., Paulay, G., and Bruggemann, J. H.: DNA barcoding of reef brittle stars (Ophiuroidea, Echinodermata) from the southwestern Indian Ocean evolutionary hot spot of biodiversity, Ecol Evol., 7, 11197–11203, https://doi.org/10.1002/ece3.3554, 2017.
Bonifácio, P., Martínez Arbizu, P., and Menot, L.: Alpha and beta diversity patterns of polychaete assemblages across the nodule province of the eastern Clarion-Clipperton Fracture Zone (equatorial Pacific), Biogeosciences, 17, 865–886, https://doi.org/10.5194/bg-17-865-2020, 2020.
Brenke, N.: An epibenthic sledge for operations on marine softbottom and bedrock, Mar. Technol. Soc. J., 39, 13–24, https://doi.org/10.4031/002533205787444015, 2005.
Calas, G.: Mineral resources and sustainable development, Elements, 13, 301–306, https://doi.org/10.2138/gselements.13.5.301, 2017.
Carney, R. S.: Zonation of deep biota on continental margins, in: Oceanography and Marine Biology: An Annual Review 43, edited by: Gibson, R. N., Atkinson, R. J. A., and Gordon, J. D. M., CRC Press, Boca Raton, USA, 211–278, https://doi.org/10.1201/9781420037449, 2005.
Carstens, B. C., Pelletier, T. A., Reid, N. M., and Satler, J. D.: How to fail at species delimitation, Mol. Ecol., 22, 4369–4383, https://doi.org/10.1111/mec.12413, 2013.
Chao, A.: Non-parametric estimation of the number of classes in a population, Scand. J. Stat., 11, 265–270, 1984.
Chazdon, R. L., Colwell, R. K., Denslow, J. S., and Guariguata, M. R.: Statistical methods for estimating species richness of woody regeneration in primary and secondary rain forests of NE Costa Rica, in: Forest biodiversity research, monitoring and modeling: Conceptual background and Old World case studies, edited by: Dallmeier, F. and Comiskey, J. A., Parthenon Publishing, Paris, France, 285–309, 1998.
Christodoulou, M., O'Hara, T. D., Hugall, A. F., and Martinez Arbizu, P.: Dark ophiuroid biodiversity in a prospective abyssal mine field, Curr. Biol., 29, 1–4, https://doi.org/10.1016/j.cub.2019.09.012, 2019.
Christodoulou, M., O'Hara, T., and Martinez Arbizu, P.: DS-CCZ1, BOLD Systems, https://doi.org/10.5883/DS-CCZ1, 2020a.
Christodoulou, M., O'Hara, T., and Martinez Arbizu, P.: DS-DEA1, BOLD Systems, https://doi.org/10.5883/DS-DEA1, 2020b.
Clark, H. L.: North Pacific Ophiurans in the collection of the United States National Museum, Bull. U.S. Natl. Mus., 75, 1–302, 1911.
Clark, A. H.: Ophiuroidea of the Hawaiian Islands, Bernice P. Bis. Mus. B., 195, 14–15, 1949.
Colins, R. A. and Cruickshank, R. H.: The seven deadly sins of DNA barcoding, Mol. Ecol. Resour., 13, 969–975, https://doi.org/10.1111/1755-0998.12046, 2012.
Colwell, R. K. and Coddington, J. A.: Estimating terrestrial biodiversity through extrapolation, Philos. T. Roy. Soc. B, 345, 101–118, https://doi.org/10.1098/rstb.1994.0091, 1994.
Danovaro, R., Aguzzi, J., Fanelli, E., Billett, D., Gjerde, K., Jamieson, A., Ramirez-Llodra, E., Smith, C. R., Snelgrove, P. V. R., Thomsen, L., and Dover, C. L.: An ecosystem-based deep-ocean strategy, Science, 355, 452–454, https://doi.org/10.1126/science.aah7178, 2017.
Dellicour, S. and Flot, J.-F.: The hitchhiker's guide to single-locus species delimitation, Mol. Ecol. Resour., 18, 1234–1246, https://doi.org/10.1111/1755-0998.12908, 2018.
DeSalle, R.: Species discovery versus species identification in DNA barcoding efforts: response to Rubinoff, Conserv. Biol., 20, 1545–1547, https://doi.org/10.1111/j.1523-1739.2006.00543.x, 2006.
Estoup, A., Largiader, C. R., Perrot, E., and Chourrout, D.: Rapid one-tube DNA extraction for reliable PCR detection of fish polymorphic markers and transgenes, Mol. Mar. Biol. Biotech., 5, 295–298, 1996.
Folmer, O. M., Black, W., and Hoen, R.: DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates, Mol. Mar. Biol. Biotech., 3, 294–299, 1994.
Fontaneto, D., Flot, J. F., and Tang, C. Q.: Guidelines for DNA taxonomy, with a focus on the meiofauna, Mar. Biodivers., 45, 433–451, https://doi.org/10.1007/s12526-015-0319-7, 2015.
Fujisawa, T. and Barraclough, T. G.: Delimiting species using single-locus data and the Generalized Mixed Yule Coalescent approach: a revised method and evaluation on simulated data sets, Syst. Biol., 62, 707–724, https://doi.org/10.1093/sysbio/syt033, 2013.
Gage, J. D. and Tyler, P. A. (Eds.): Deep-Sea Biology: A Natural History of Organisms at the Deep-Sea Floor, Cambridge University Press, Cambridge, UK, 1991.
Glover, A. G., Wiklund, H., Rabone, M., Amon, D. J., Smith, C. R., O'Hara, T. D., Mah, C. L., and Dahlgren, T. G.: Abyssal fauna of the UK-1 polymetallic nodule exploration claim, Clarion-Clipperton Zone, central Pacific Ocean: Echinodermata, Biodivers. Data J., 4, e7251, https://doi.org/10.3897/BDJ.4.e7251, 2016.
Haeckel, M., Köonig, I., Riech, V., Weber, M. E., and Suess, E.: Pore water profiles and numerical modelling of biogeochemical processes in Peru Basin deep-sea sediments, Deep-Sea Res. Pt. II, 48, 3713–3736, https://doi.org/10.1016/S0967-0645(01)00064-9, 2001.
Hall, C. E. and Gurnis, M.: Strength of fracture zones from their bathymetric and gravitational evolution, J. Geophys. Res.-Sol. Ea., 110, B01402, https://doi.org/10.1029/2004JB003312, 2005.
Hauquier, F., Macheriotou, L., Bezerra, T. N., Egho, G., Martínez Arbizu, P., and Vanreusel, A.: Distribution of free-living marine nematodes in the Clarion–Clipperton Zone: implications for future deep-sea mining scenarios, Biogeosciences, 16, 3475–3489, https://doi.org/10.5194/bg-16-3475-2019, 2019.
Hein, J., Mizell, K., Koschinsky, A., and Conrad, T.: Deep-ocean mineral deposits as a source of critical metals for high- and green-technology applications: comparison with land-based resources, Ore Geol. Rev., 51, 1–14, https://doi.org/10.1016/j.oregeorev.2012.12.001, 2013.
Hoang, D. T., Chernomor, O., von Haeseler, A., Minh, B. Q., and Vinh, L. S.: UFBoot2: Improving the ultrafast bootstrap approximation, Mol. Biol. Evol., 35, 518–522, https://doi.org/10.1093/molbev/msx281, 2018.
Hofmann, E. P., Nicholson, K. E., Luque-Montes, I. R., Köhler, G., Cerrato-Mendoza, C. A., Medina-Flores, M., Wilson, L. D., and Townsend, J. H.: Cryptic diversity, but to what extent? Discordance between single-locus species delimitation methods within mainland anoles (Squamata: Dactyloidae) of Northern Central America, Front. Genet., 10, 11, https://doi.org/10.3389/fgene.2019.00011, 2019.
Jamieson, A. (Ed.): The Hadal Zone. Life in the deepest oceans, Cambridge University Press, Cambridge, UK, 2015.
Jamieson, A. J., Fujii, T., Mayor, D. J., Solan, M., and Priede, I. G.: Hadal trenches: the ecology of the deepest places on Earth, Trends Ecol. Evol., 25, 190–197, https://doi.org/10.1016/j.tree.2009.09.009, 2009.
Kapli, P., Lutteropp, S., Zhang, J., Kobert, K., Pavlidis, P., Stamatakis, A., and Flouri, T.: Multi-rate Poisson tree processes for single-locus species delimitation under maximum likelihood and Markov chain Monte Carlo, Bioinformatics, 33, 1630–1638, https://doi.org/10.1093/bioinformatics/btx025, 2017.
Katoh, K., Misawa, K., Kuma, K. I., and Miyata, T.: MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform, Nucleic Acids Res., 30, 3059–3066, https://doi.org/10.1093/nar/gkf436, 2002.
Kearse, M., Moir, R., Wilson, A., Stones-Havas, S., Cheung, M., Sturrock, S., Buxton, S., Cooper, A., Markowitz, S., Duran, C., Thierer, T., Ashton, B., Mentjies, P., and Drummond, A.: Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data, Bioinformatics, 28, 1647–1649, https://doi.org/10.1093/bioinformatics/bts199, 2012.
Kekkonen, M. and Hebert, P. D. N.: DNA barcode-based delineation of putative species: Efficient start for taxonomic workflows, Mol. Ecol. Resour., 14, 706–715, https://doi.org/10.1111/1755-0998.12233, 2014.
Khodami, S., Martinez Arbizu, P., Stöhr, S., and Laakmann, S.: Molecular species delimitation of Icelandic brittle stars (Ophiuroidea), Pol. Polar Res., 35, 243–260, https://doi.org/10.2478/popore-2014-0011, 2014.
Laakman, S., Boos, K., Knebelsberger, T., Raupach, M. J., and Neumann, H.: Species identification of echinoderms from the North Sea by combining morphology and molecular data, Helgoland Mar. Res., 70, 18, https://doi.org/10.1186/s10152-016-0468-5, 2016.
Layton, K. K. S., Corstorphine, E. A., and Hebert, P. D. N.: Exploring Canadian echinoderm diversity through DNA barcodes, PLoS ONE, 11, e0166118, https://doi.org/10.1371/journal.pone.0166118, 2016.
Legendre, P. and Gallagher, E. D.: Ecologically meaningful transformations for ordination of species data, Oecologia, 129, 271–280, https://doi.org/10.1007/s004420100716, 2001.
Lodge, M., Johnson, D., Le Gurun, G., Wengler, M., Weaver, P., and Gunn, V.: Seabed mining: International Seabed Authority environmental management plan for the Clarion–Clipperton Zone. A partnership approach, Mar. Policy, 49, 66–72, https://doi.org/10.1016/j.marpol.2014.04.006, 2014.
Ludwig, H.: Die Ophiuren der Sammlung Plate, Zool. Jahrb., 4, 750–786, 1898.
Ludwig, H.: Ophiuroideen, in: Ergebnisse der Hamburger Magalhaensischen Sammelreise 1892/93, I Band, L. Friedericsen & Co, Hamburg, Germany, 1899.
Lusty, P. A. J. and Murton, B. J.: Deep-ocean mineral deposits: Metal resources and windows into earth processes, Elements, 14, 301–306, https://doi.org/10.2138/gselements.14.5.301, 2018.
Lyman, T.: Ophiuridae and Astrophytidae of the exploring voyage of H.M.S. Challenger, under Prof. Sir Wyville Thomson, F.R.S. Part 1, Bull. Mus. Comp. Zool., 5, 65–168, 1878.
Lyman, T.: Ophiuridae and Astrophytidae of the exploring voyage of H.M.S. Challenger under Prof. Sir Wyville Thomson, F.R.S. Part 2, Bull. Mus. Comp. Zool., 6, 17–83, 1879.
Lyman, T.: Report on the Ophiuroidea, in: Report on the Scientific Results of the Voyage of H.M.S. Challenger, Zoology, 5, edited by: Thomson, C. W. and Murray, J., H. M. Stationery Off., London, UK, 1–192, 1882.
Mero, J. L.: Chapter 11. Economic aspects of nodule mining, in: The mineral resources of the sea, edited by: Mero, J. L., Elsevier, Amsterdam, the Netherlands, 327–355, 1965.
Miller, K. A., Thompson, K. F., Johnston, P., and Santillo, D.: An overview of seabed mining including the current state of development, environmental impacts, and knowledge gaps, Front. Mar. Sci., 4, 418, https://doi.org/10.3389/fmars.2017.00418, 2018.
Mukhopadhyay, R., Ghosh, A. K., and Iyer, S. D.: The Indian Ocean nodule field: Geology and resource potential, in: Handbook of exploration and environmental geochemistry, Vol. 10, edited by: Hale, M., Elsevier, Amsterdam, the Netherlands, 1–292, 2008.
Murray, J. and Renard, A. F.: Report on deep-sea deposits based on the specimens collected during the voyage of H.M.S. Challenger in the years 1872 to 1876, in: Report on the scientific results of the voyage of H.M.S., Challenger during the years 1873–76, edited by: Thomson, C. W. and Murray, J., H. M. Stationery Off., London, UK, 1–688, 1891.
Niner, H. J., Ardron, J. A., Escobar, E. G., Gianni, M., Jaeckel, A., Jones, D. O. B., Levin, L. A., Smith, C. R., Thiele, T., Turner, P. J., Van Dover, C. L., Watling, L., and Gjerde, K. M.: Deep-sea mining with no net loss of biodiversity – An impossible aim, Front. Mar. Sci., 5, 53, https://doi.org/10.3389/fmars.2018.00053, 2018.
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A., and Minh, B. Q.: IQ-TREE: A fast and effective stochastic algorithm for estimating maximum likelihood phylogenies, Mol. Biol. Evol., 32, 268–274, https://doi.org/10.1093/molbev/msu300, 2015.
O'Hara, T. D., Hugall, A. F., Thuy, B., and Moussalli, A.: Phylogenomic resolution of the class Ophiuroidea unlocks a global microfossil record, Curr. Biol., 24, 1874–1879, https://doi.org/10.1016/j.cub.2014.06.060, 2014.
O'Hara, T. D., Hugall, A. F., Thuy, B., Stöhr, S., and Martynov, A. V.: Restructuring higher taxonomy using broad-scale phylogenomics: the living Ophiuroidea, Mol. Phylogenet. Evol., 107, 415–430, https://doi.org/10.1016/j.ympev.2016.12.006, 2017.
Oksanen, J., Kindt, R., Legendre, P., O'Hara, B., Simpson, G. L., Solymos, P., Stevens, M. H. H., and Wagner, H.: The vegan Package, Community ecology package, 10, 631–637, 2008.
Puillandre, N., Lambert, A., Brouillet, S., and Achaz, G.: ABGD, Automatic Barcode Gap Discovery for primary species Delimitation, Mol. Ecol., 21, 1864–1877, https://doi.org/10.1111/j.1365-294X.2011.05239.x, 2012.
Pons, J., Barraclough, T. G., Gomez-Zurita, J., Cardoso, A., Duran, D. P., Hazell, S., Kamoun, S., Sumlin, W. D., and Vogler, A. P.: Sequence-based species delimitation for the DNA taxonomy of undescribed insects, Syst. Biol., 55, 595–609, https://doi.org/10.1080/10635150600852011, 2006.
Ramirez-Llodra, E., Brandt, A., Danovaro, R., De Mol, B., Escobar, E., German, C. R., Levin, L. A., Martinez Arbizu, P., Menot, L., Buhl-Mortensen, P., Narayanaswamy, B. E., Smith, C. R., Tittensor, D. P., Tyler, P. A., Vanreusel, A., and Vecchione, M.: Deep, diverse and definitely different: unique attributes of the world's largest ecosystem, Biogeosciences, 7, 2851–2899, https://doi.org/10.5194/bg-7-2851-2010, 2010.
Ramirez-Llodra, E., Tyler, P. A., Baker, M. C., Bergstad, O. A., Clark, M. R., Escobar, E., Levin, L. A., Menot, L., Rowden, A. A., Smith, C. R., and Van Dover, C. L.: Man and the Last Great Wilderness: Human Impact on the Deep Sea, PLoS ONE, 6, e22588, https://doi.org/10.1371/journal.pone.0022588, 2011.
Ratnasingham, S. and Hebert, P. D. N.: A DNA-based registry for all animal species: The barcode index number (BIN) system, PLoS ONE, 8, e66213, https://doi.org/10.1371/journal.pone.0066213, 2013.
Rex, M. A. and Etter, R. J.: Deep-Sea Biodiversity: Pattern and Scale, Harvard University Press, Cambridge, USA, 2010.
Simpson, G. G.: Mammals and the nature of continents, Am. J. Sci., 241, 1–31, 1943.
Smith, A. B., Paterson, G. L. J., and Lafay, B.: Ophiuroid phylogeny and higher taxonomy: morphological, molecular and palaeontological perspectives, Zool. J. Linn. Soc.-Lond., 114, 213–243, 1995.
Smith, C. R., De Leo, F. C., Bernardino, A. F., Sweetman, A. K., and Martinez Arbizu, P.: Abyssal food limitation, ecosystem structure and climate change, Trends Ecol. Evol., 23, 518–528, https://doi.org/10.1016/j.tree.2008.05.002, 2008.
Song, C., Lin, X.-L., Wang, Q., and Wang, X.-H.: DNA barcodes successfully delimit morphospecies in a superdiverse insect genus, Zool. Scr., 47, 311–324, https://doi.org/10.1111/zsc.12284, 2018.
Sørensen, T.: A method of establishing groups of equal amplitude in plant sociology based on similarity of species content and its application to analyses of the vegetation on Danish commons, Biol. Skr. Dan. Vid. Sel., 5, 1–34, 1948.
Stöhr, S., O'Hara, T. D., and Thuy, B.: Global Diversity of Brittle Stars (Echinodermata: Ophiuroidea), PLoS ONE, 7, e31940, https://doi.org/10.1371/journal.pone.0031940, 2012.
Stratmann, T., Lins, L., Purser, A., Marcon, Y., Rodrigues, C. F., Ravara, A., Cunha, M. R., Simon-Lledó, E., Jones, D. O. B., Sweetman, A. K., Köser, K., and van Oevelen, D.: Abyssal plain faunal carbon flows remain depressed 26 years after a simulated deep-sea mining disturbance, Biogeosciences, 15, 4131–4145, https://doi.org/10.5194/bg-15-4131-2018, 2018.
Taboada, S., Riesgo, A., Wiklund, H., Paterson, G. L. J., Koutsouveli, V., Santodomingo, N., Dale, A. C., Smith, C. R., Jones, D. O. B., Dahlgren, T. G., and Glover, A. G.: Implications of population connectivity studies for the design of marine protected areas in the deep sea: An example of a demosponge from the Clarion-Clipperton Zone, Mol. Ecol., 27, 4657–4679, https://doi.org/10.1111/mec.14888, 2018.
Tang, C. Q., Humphreys, A. M., Fontaneto, D., Barraclough, T. G., and Paradis, E.: Effects of phylogenetic reconstruction method on the robustness of species delimitation using single-locus data, Methods Ecol. Evol., 5, 1086–1094, https://doi.org/10.1111/2041-210X.12246, 2014.
Thiel, H. and Schriever, G.: Deep-Sea Mining, Environmental Impact and the DISCOL Project, Ambio, 19, 245–250, 1990.
Thiel, H., Schriever, G., Ahnert, A., Bluhm, H., Borowski, C., and Vopel, K.: The large-scale environmental impact experiment DISCOL: reflection and foresight, Deep-Sea Res. Pt. II, 48, 3869–3882, https://doi.org/10.1016/S0967-0645(01)00071-6, 2001.
Tilot, V., Ormond, R., Moreno Navas, J., and Catalá, T. S.: The Benthic Megafaunal Assemblages of the CCZ (Eastern Pacific) and an Approach to their Management in the Face of Threatened Anthropogenic Impacts, Front. Mar. Sci., 5, 7, https://doi.org/10.3389/fmars.2018.00007, 2018.
Van Dover, C. L., Ardron, J. A., Escobar, E., Gianni, M., Gjerde, K. M., Jaeckel, A., Jones, D. O. B., Levin, L. A., Niner, H. J., Pendleton, L., Smith, C. R., Thiele, T., Turner, P. J., Watling, L., and Weaver, P. P. E.: Biodiversity loss from deep-sea mining, Nat. Geosci., 10, 464, https://doi.org/10.1038/ngeo2983, 2017.
Vanreusel, A., Hilario, A., Ribeiro, P. A., Menot, L., and Arbizu, P. M.: Threatened by mining, polymetallic nodules are required to preserve abyssal epifauna, Sci. Rep.-UK, 6, 26808, https://doi.org/10.1038/srep26808, 2016.
Volz, J. B., Mogollon, J. M., Geibert, W., Arbizu, P. M., Koschinsky, A., and Kasten, S.: Natural spatial variability of depositional conditions, biogeochemical processes and element fluxes in sediments of the eastern Clarion-Clipperton Zone, Pacific Ocean, Deep-Sea Res. Pt. I, 140, 159–172, https://doi.org/10.1016/j.dsr.2018.08.006, 2018.
Volz, J. B., Haffert, L., Haeckel, M., Koschinsky, A., and Kasten, S.: Impact of small-scale disturbances on geochemical conditions, biogeochemical processes and element fluxes in surface sediments of the eastern Clarion–Clipperton Zone, Pacific Ocean, Biogeosciences, 17, 1113–1131, https://doi.org/10.5194/bg-17-1113-2020, 2020.
Wedding, L. M., Friedlander, A. M., Kittinger, J. N., Watling, L., Gaines, S. D., Bennett, M., Hardy, S. M., and Smith, C. R.: From principles to practice: a spatial approach to systematic conservation planning in the deep sea, P. Roy. Soc. Lond. B Bio., 280, 1684–1684, https://doi.org/10.1098/rspb.2013.1684, 2013.