the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Soil microbial diversity and network complexity promote phosphorus transformation – a case of long-term mixed plantations of Eucalyptus and a nitrogen-fixing tree species
Jiyin Li
Yeming You
Wen Zhang
Yi Wang
Yuying Liang
Haimei Huang
Hailun Ma
Qinxia He
Angang Ming
Increased nitrogen (N) availability influences soil phosphorus (P) cycling through multiple pathways. Soil microorganisms are essential facilitating a wide range of ecosystem functions. However, the impact of how mixed plantations of Eucalyptus and N-fixing tree species affect P transformation and microbiota interactions remains unknown. Therefore, we conducted a 17 y field experiment comparing pure Eucalyptus plantations (PPs) and mixed plantations (MPs) with Eucalyptus and an N-fixing tree species to assess their effects on soil P transformation, using data collected from two soil layers (0–10 and 10–20 cm). The results showed that α-diversity indices (ACE, Chao1, and Shannon indices) were significantly higher in MPs than in PPs for both bacteria and fungi. Furthermore, MPs exhibited significantly higher relative abundances of bacterial phyla Proteobacteria (0–10 cm), Verrucomicrobia, and Rokubacteria, as well as fungal phyla Mortierllomycota, Mucoromycota, and Rozellomycota. Conversely, MPs showed lower abundances of the bacterial phyla Chloroflexi, Actinobacteria, and Planctomycetes and the fungal phylum Ascomycota. Gene copy numbers of functional genes were also elevated in MPs, including 16S rRNA, internal transcribed spacer (ITS), N functional genes [nifH (0–10 cm), AOB-amoA, narG, nirS, and nosZ (0–10 cm)], and P functional genes [phoC, phoD (0–10 cm), BPP, and pqqC]. The findings indicate that MPs can enhance soil microbial diversity, network complexity, and the relative abundance of functional genes, which involved N and P transformation, by optimizing soil nutrient levels and pH, thereby facilitating P transformation. Therefore, MPs of Eucalyptus and N-fixing tree species may represent a promising forest management strategy to improve ecosystem P benefits.
- Article
(5160 KB) - Full-text XML
-
Supplement
(780 KB) - BibTeX
- EndNote
Phosphorus (P) is a vital macronutrient for plant and microbial growth (Turner et al., 2018), while the availability of P serves as a key indicator of soil fertility and quality (Peng et al., 2021). In most ecosystems, particularly in tropical and subtropical forests, P bioavailability in soil is often limited due to intense weathering and the presence of aluminum (Al) ions and free iron (Fe) (Soltangheisi et al., 2019; Du et al., 2020). Therefore, these P reserves cannot be accessed directly by plants (Fan et al., 2019). However, plants and microorganisms have developed various strategies to access P from inorganic (Pi) and organic (Po) reservoirs and render it available for biological processes (including, e.g., assimilation by phosphate-solubilizing microorganisms and mineralization of enzymes) (Lu et al., 2022). Consequently, it is crucial to implement strategies for the sustainable management of soil P to enhance its utilization by plants, preserve soil quality, and mitigate the risk of P loss.
Soil microorganisms serve as both a reservoir and a source of phosphate ions, significantly influencing the availability of P. In addition, microorganisms play a role in maintaining soil functions such as nutrient cycling, biological activity, and plant growth, all of which are crucial for sustaining soil quality and fertility (Bünemann et al., 2008; Zhou et al., 2018; Sun et al., 2022). Microorganisms facilitate the P transformation by participating in the processes of P mineralization, solubilization, and cycling, converting P into bioavailable forms for plant uptake (Pastore et al., 2020). Specifically, the mineralization of Po is facilitated by the extracellular presence of phosphatases, which are produced principally by soil microorganisms (Nannipieri et al., 2012). Therefore, both extracellular acid (ACP) and alkaline (ALP) phosphatase activities are commonly used as indicators to assess the mineralization of Po to bioavailable Pi (Luo et al., 2019). Furthermore, P transformation is influenced by the α-diversity, structure, and composition of soil microbial communities, with pH being considered a key determinant in shaping microbial diversity and community composition (Jin et al., 2019). Microbiome co-occurrence networks are prevalently employed to scrutinize the interrelationships within microbial communities, and network attributes (e.g., the mean degree, edge quantity, and node amount) can be utilized to appraise the reciprocal ties among these communities and their reactions to modifications in cultivation paradigms (Faust, 2021; Qiu et al., 2021). Microbial network analysis can uncover the complex interactions between microorganisms, such as competition, cooperation, and antagonism, while also illuminating important ecological processes and functional relationships that are not fully captured by microbial diversity analysis alone. For instance, it can reveal processes like the transformation and cycling of key soil nutrients (e.g., P and N), which are often overlooked in traditional diversity assessments (Yao et al., 2024). Thus, gaining insight into the relationships between microbial diversity, microbial network complexity, and the transformation and cycling of P is crucial for improving soil functions and enhancing soil fertility.
The studies on genes involved in P cycling also emphasize the contribution of microbes in enhancing plant P uptake and efficiency (Dai et al., 2020). The P cycle cluster includes genes that stimulate the mineralization of Po (e.g., phoD, phoC, and BPP) (Cao et al., 2022; Khan et al., 2023) and solubilization of Pi (e.g., pqqC) (Meyer et al., 2011). The genes phoDphoC encode phosphatases, which are capable of mineralizing Po compounds into Pi (Fraser et al., 2015).
Nitrogen is a fundamental element for plant growth and development, typically coupled with P in biogeochemical cycles. The N cycle group consists of genes responsible for microbially driven nitrification (e.g., AOB-amoA), N fixation (e.g., nifH), and denitrification (e.g., nirS). Improved interaction networks among soil microbial functional groups contribute to increasing nutrient availability and enhancing the nutrient acquisition of host plants (Shi et al., 2020; Qin et al., 2024). In addition, given that both N and P are essential elements for microorganisms, an increase in N content can influence soil pH, which subsequently alters the composition of soil microbial communities and impacts the abundance of phosphatase-coding genes (phoC and phoD) (Widdig et al., 2020). Furthermore, the presence of N-fixing plants also affects P uptake by enhancing litter decomposition rates and the release of organic acids from microbial biomass, thereby accelerating nutrient cycling and improving soil fertility (Li et al., 2021). Therefore, studying the coupling of N and P cycling in soil is crucial for understanding the diversity and mechanisms of microbially driven biogeochemical cycles.
Eucalyptus is characterized by its straight trunks, well-developed horizontal root systems, and high adaptability. They are prevalent in subtropical and tropical regions, where they have significant economic and ecological value (Zhang and Wang, 2021). However, monocultures and short-term rotation management of Eucalyptus plantations have led to soil degradation, reductions in plant available soil nutrient effectiveness (e.g., the availability of nutrients such as N and P in forms that can be absorbed and utilized by plants), and soil microbial function and diversity, as well as other adverse ecological effects. Mixed species forests exert a strong positive impact on soil fertility and nutrient cycling by regulating the microbiome, including its diversity and structure (Pereira et al., 2019; Li et al., 2024). Recently, incorporating N-fixing tree species such as Acacia as a substitute for N fertilization has become widely acknowledged as one of the most effective silvicultural practices for enhancing tree N uptake and woody production in Eucalyptus plantations (Koutika and Mareschal, 2017; Zhang et al., 2023). In addition, mixing with N-fixing tree species improves N availability, P accumulation, microbial diversity, and forms a more complex and interconnected microbial network compared to pure plantations (Li et al., 2022; He et al., 2025; Yao et al., 2021). So far, the effect of N-fixing tree species on P cycling has mainly been addressed by investigating organic or inorganic P accumulation in soil from either pure or mixed stands of non-N-fixing tree species and N-fixing tree species (Yao et al., 2024).
Acacia mangium, one of the N-fixing tree species that is widely planted in many parts of the world, has clear benefits in forestry and agroforestry ecosystems (Epron et al., 2013; Koutika and Richardson, 2019). Key reasons for the widespread planting of Acacia mangium in pure or mixed plantations with other tree species with infertile soils are its capacity to change soil fauna and microbial communities (Huang et al., 2014; de Araujo Pereira et al., 2017), improve soil fertility (Tchichelle et al., 2017), and stimulate tree growth and forest productivity (Paula et al., 2015). Nevertheless, the effects of mixing N-fixing tree species on regulating the correlations between microbial diversity and the network of P transformation are still poorly understood. Phosphomonoesterase (i.e., ACP) mineralization is an essential strategy for P transformation (Luo et al., 2019; Yu et al., 2022; Wang et al., 2023), so we employed soil ACP activity to analyze the dynamics of P transformation. Here, we aimed to (1) compare variations in the structure, diversity, and stability of soil microbial communities after mixing Eucalyptus with N-fixing tree species and (2) elucidate the mechanisms through which fungal and bacterial communities, along with genes associated with N and P transformation processes, regulate P transformation. We hypothesized that (1) mixed species plantations of Eucalyptus and N-fixing tree species would alter the composition of soil microbial communities and improve microbial community diversity and network complexity in the soil; (2) introduction of N-fixing tree species may cause imbalance in soil properties (e.g., soil organic carbon, pH, and so on), microbial diversity and networks complexity, and related functional genes which co-regulate the P transformation with differential roles. Our findings will provide new insights into sustainable management practices for plantations.
2.1 Site description
The study was conducted in the Shaoping Experimental Field at the Experimental Center for Tropical Forestry, which is affiliated with the Chinese Academy of Forestry (106°56′ E, 22°03′ N). The area has a subtropical climate, with approximately 1400 mm of rainfall annually and an average yearly temperature of 21.2 °C. The landscape is characterized by low mountains and hills along with acidic red soil. Forests in this area are primarily composed of commercially managed plantations as either pure or mixed stands.
2.2 Plot design and sampling
In this study, the pure (monoculture) Eucalyptus urophylla plantations (PPs) and adjacent mixed plantations (MPs) of Eucalyptus urophylla and Acacia mangium (N-fixing tree species) were established in 2004 on the logging tracks of Pinus massoniana plantations that were established in 1977. The MPs were planted at a 1:1 mixing ratio with inter-row planting, consisting of one row of Eucalyptus urophylla and one row of Acacia mangium. In the first two consecutive years post planting, both plantations were subjected to a similar stand management regime, which included practices such as weed control and fertilization, subsequently allowing them to proceed with their natural stand development. The experimental design is described in the study conducted by Huang et al. (2017). In 2021, taking into account the differences in plantation layout and topography, five 20 m × 20 m sample plots were randomly established in each stand (PPs and MPs), ensuring that adjacent plots were maintained at a distance greater than 200 m to mitigate edge effects. The diameter at breast height, height, and stand density of every tree within each plot were assessed. Detailed information on the plantations is provided in Table S1.
Soil samples were collected in early August 2021. Soil samples were gathered from eight different points within each plot, located at 5 m intervals from the center, along angles of 0, 45, 90, 135, 180, 225, 270, and 315°. Previous studies only examine a single soil layer (usually the upper 0–10 cm) (Waithaisong et al., 2022; Chen et al., 2024). More study on the P transformation and mechanisms underlying soil microbial and biochemical interactions in different soil layers is needed to determine whether the variation of the P cycle is dependent on depth. Therefore, soil samples in our study were obtained from the depth intervals of 0–10 and 10–20 cm following the removal of extraneous materials such as small stones and dead leaves. Eight undisturbed samples from each soil layer were amalgamated into a composite sample and transported to the laboratory on ice. Each composite sample was partitioned into two aliquots: one designated for the analysis of physicochemical properties and the other reserved for genomic DNA extraction.
2.3 Soil properties and soil enzyme activity
Soil pH was measured using a 1:2.5 soil:water ratio, and soil organic carbon (SOC) was quantified using the K2Cr2O7-H2SO4 oxidation method. The total nitrogen (TN) content of soil was determined using an Auto Analyzer III in an extract obtained by digestion of the sample with H2SO4 and a catalyst (). The levels of nitrate N (NO-N) and ammonia N (NH-N) were determined by CaCl2 extraction, followed by quantitative analysis using an AutoAnalyzer III (Tsiknia et al., 2014). Total P (TP) was quantified using the molybdenum blue colorimetric method following the extraction of the samples with HClO4-H2SO4 (Murphy and Riley, 1962).
N and P metabolized by soil extracellular enzyme activity (EEA), e.g., β-1,4-N-acetylglucosaminidase (NAG) and leucine aminopeptidase (LAP) activity are involved in N acquisition, and acid phosphomonoesterase is associated with P mineralization, were quantified in a fluorescence assay conducted in a 96 well microplate (Yan et al., 2022). Soil EEA was calculated from the fluorescence readings of the enzyme after its reaction with the appropriate substrate. The assay was conducted using 200 µL of a soil suspension prepared by weighing 1.25 g of fresh soil to which sodium acetate buffer (pH 4.5) was added and stirred for 1 min to ensure consistent extraction conditions and effective solubilization of the soil constituents. Eight replicates per sample were tested. The samples were incubated in darkness at 25 °C for 3 h, after which the reaction was terminated by adding NaOH. Fluorescence was then immediately measured within the wavelength range of 365–450 nm by using a fluorescence microplate reader. Information on the substrates of the three EEA can be found in Table S2.
2.4 Soil DNA extraction and sequencing
Microbial genomic DNA was obtained from soil samples utilizing the PowerSoil DNA isolation kit (MN NucleoSpin 96 Soi, MACHEREY-NAGEL, Dueren, Germany) for subsequent analysis and measurements. The primers employed were 338F and 806R for the amplification of the V3–V4 hypervariable region of the 16S rRNA gene (Mori et al., 2014; Parada et al., 2016), while ITS1F and ITS2R were employed to amplify the ITS1 region of fungal rRNA gene loci (Adams et al., 2013; Dong et al., 2021) (Table S3). Sequencing data were processed by filtering the raw reads using Trimmomatic v0.33, removing the primers using Cutadapt v1.9.1, assembling the clean reads by overlap with Usearch v10, and removing chimeras with UCHIME v4.2 to ensure data validity. After the removal of potential chimeras, 1 600 678 and 1 550 033 high-quality bacterial and fungal reads were obtained, respectively.
The genetic potential of the soil microorganisms was assessed by real-time fluorescence quantitative PCR (qPCR) to quantitatively determine the gene copy numbers of bacteria (16S rRNA) and fungi (ITS). The genetic potential of N cycling processes was evaluated based on the abundance of functional genes involved in nitrogen fixation (nifH), nitrification (AOB-amoA), and denitrification (narG, nirS, nirK, and nosZ). Similarly, the genetic potential of P cycling processes was assessed using the abundance of functional genes involved in organic phosphorus hydrolysis (phoC, phoD, BPP) and Pi hydrolysis (pqqC). These functional genes are well established biomarkers of the biochemical pathways essential for nutrient cycling in various ecosystems. The qPCR amplification efficiencies ranged from 90 % to 110 %. The primers and references for the functional genes are reported in Table S3.
2.5 Network construction
Networks for bacteria and fungi were constructed by dividing the 20 samples into four groups consisting of two soil layers for PPs and MPs. First, sample operational taxonomic units (OTUs) were filtered, discarding those that appeared in fewer than three samples within each group (3 out of 5 replicates) (Hu et al., 2023). OTUs with a relative abundance exceeding 1 % in the bacterial and fungal communities were selected for further correlation analysis (Fan et al., 2018). The network was built according to thresholds of Pearson correlation coefficient >0.6 and P<0.05, assessed using the Hmisc package in R v4.0.5. We adjusted the P values according to the Hochberg false discovery rate test (Benjamini et al., 2006), with a cut-off of adjusted P<0.05. Network properties were computed utilizing the igraph R package and visualized using Gephi (https://gephi.org/, last access: 23 April 2024). In all figures, bacterial and fungal phyla exhibiting a relative abundance greater than 1 % within the network are represented by distinct colors.
Keystone species were identified by utilizing the connectivity within modules (Zi) and between modules (Pi). Microorganisms were classified into four categories depending on intra-module degree (Z-score) and participation coefficient (C-score) thresholds. These categories are: network hubs, module hubs, connectors, and peripherals (Poudel et al., 2016). Network hubs refer to nodes with a high degree of connectivity both globally and within individual modules; module hubs are nodes with significant connectivity restricted to a single module, connectors are nodes that facilitate strong connections between different modules, and peripheral nodes are those with few connections to other nodes (Poudel et al., 2016). Network hubs, module hubs, and connectors occupy critical positions within the network and are classified as keystone topological features. These characteristics are essential for sustaining the stability of microbial communities (Delmas et al., 2019). Consequently, OTUs associated with these nodes were designated as keystone species.
2.6 Data analyses
Microbial diversity (Shannon index) and richness (Chao1 and ACE) values were all calculated using phyloseq with default settings by Mothur (v 1.30.2) software (Schloss et al., 2009). Soil physicochemical properties, microbial community indices (such as the ACE and Shannon and Chao1 indices), as well as functional genes and enzyme activity, were analyzed in independent samples t-tests using SPSS v24.0. This statistical approach was applied to evaluate differences attributable to stand type (monoculture or mixed). Differences in soil microorganisms across stand types and soil layers were analyzed using non-metric multidimensional scaling (NMDS) with Bray–Curtis dissimilarity and analysis of similarity (ANOSIM), implemented using the vegan package in R (Oksanen et al., 2013; Knowles et al., 2019). Random forest analysis based on Pearson correlation analysis and the best multiple regression model was used to evaluate the contributions of soil properties, microbial characteristics, and functional genes involved in the N and P cycles to the variation in nitrogen and phosphorus transformation enzyme activities and to identify the major predictors based on their importance. Computation and visualization were conducted in R (Jiao et al., 2020). Correlation analysis and visualization of soil properties, microbial characteristics, and functional genes related to N and P cycling were performed in Origin 2024. A redundancy analysis (RDA) was employed to explore the multivariate associations between soil physicochemical characteristics and microorganisms. The most important soil physicochemical properties affecting bacterial and fungal phyla were identified in the RDA and visualized using CANOCO v5. A partial least squares path model (PLS-PM) was constructed using R software to assess the direct and indirect effects of mixed planting of Eucalyptus and Acacia on P transformation. A PLS-PM can reveal causal connections between observed and latent variables, and its superiority for small sample sizes has been demonstrated in simulation studies in which path modeling estimation was shown to be reliable (Monecke and Leisch, 2012; Sanchez, 2013). The goodness-of-fit statistic was used to assess the adequacy of the PLS-PM fit, with a value >0.7 indicating good model fit (Tenenhaus et al., 2004; Sanchez, 2013).
3.1 Soil properties
Significantly (P<0.05) higher values of SOC, TN, NO-N, C:P, N:P, and pH were determined in both investigated soil layers in MPs than those in PPs (Table 1); however, TP (10–20 cm) was significantly lower in MPs than in PPs (P<0.05) (Table 1).
Table 1Soil physicochemical properties in both 0–10 and 10–20 cm soil layers in PPs and MPs.
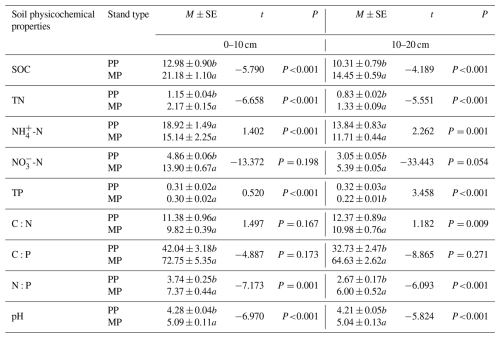
SOC: soil organic carbon; TN: total nitrogen; NH-N: ammonium nitrogen; NO-N: nitrate nitrogen; TP: total phosphorus; C:N: carbon: nitrogen ratio; C:P: carbon: phosphorus ratio; N:P: nitrogen: phosphorus ratio; pH: soil pH value; value = mean ± standard error; different lowercase letters in the table represent significant differences between PPs and MPs (P<0.05), the same below.
3.2 Bacterial and fungal community diversity and composition
In both soil layers, the bacterial ACE (0–10 cm: , P=0.001; 10–20 cm: , P<0.001), Chao1 (0–10 cm: , P=0.001; 10–20 cm: , P<0.001), and Shannon (0–10 cm: , P=0.008; 10–20 cm: , P<0.005) indices of α-diversity were significantly higher in MPs than in PPs (Fig. 1a–c). Fungal Shannon (, P=0.006) index in the 0–10 cm was also significantly higher in MPs than in PPs (Fig. 1f). The composition of bacterial and fungal community exhibited significant differences between the two plantation types and soil layers, except for the fungal communities in PPs, which did not differ between the surface and deeper soil layers (P<0.05, ANOSIM: R2=0.85, P=0.01, stress = 0.03 and R2=0.73, P=0.01, stress = 0.05, respectively, Fig. S1).
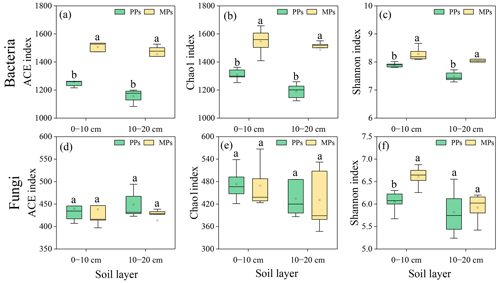
Figure 1Comparisons of (a)–(c) bacterial and (d)–(f) fungal community by α diversity index in two soil layers in PPs and MPs. Different lowercase letters in the table represent significant differences between PPs and MPs (P<0.05), the same below.
After clustering at a 97.0 % similarity level, a total of 1869 OTUs were obtained for bacteria, which revealed 21 phyla, 64 classes, 140 orders, 201 families, and 311 genera. For fungi, a total of 1128 OTUs were obtained, showing 8 phyla, 24 classes, 62 orders, 104 families, and 157 genera (Table S4). The most abundant bacterial phyla (relative abundance >1 %) in both PPs and MPs were Acidobacteria (26.83 %), Proteobacteria (22.46 %), Chloroflexi (13.95 %), Actinobacteria (13.62 %), Verrucomicrobia (11.16 %), Planctomycetes (5.6 %), and Rokubacteria (3.5 %), which represented 94.08 % of the total bacterial community in the 0–10 cm layer (Figs. 2a, b and S2a). The most abundant fungal phyla (relative abundance >1 %) in both PPs and MPs were Ascomycota (63.25 %), Basidiomycota (28.14 %), Mortierellomycota (1.77 %), Mucoromycota (1.18 %), and Rozellomycota (1.06 %), which represented 95.40 % of the total fungal community (Figs. 2c, d and S2b). The introduction of N-fixing tree species resulted in changes in the relative abundance and composition of these microbial communities, although these changes were not always statistically significant (Fig. 2).
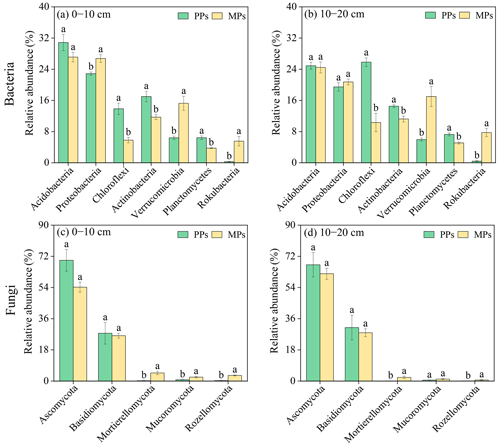
Figure 2Abundance difference of (a)–(b) bacteria and (c)–(d) fungi and based on relative abundance >1 % at phylum level.
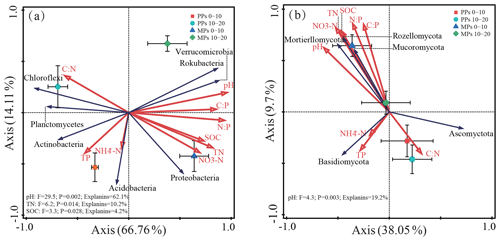
We used RDA to determine the linkage between soil microbial phyla and the specific soil physicochemical factors. The first two components of RDA axes explained 80.87 % and 47.75 % of the total variance in the relationship between soil bacterial and fungal communities and nine selective soil physicochemical factors, respectively (Fig. 3a, b). Forward selection of the nine soil physicochemical factors in the RDA ordinations showed that the bacterial communities were primarily influenced by pH, TN, and SOC (Fig. 3a), and the fungal communities were primarily influenced by pH (P<0.05) (Fig. 3b).
3.3 Microbial network complexity and stability
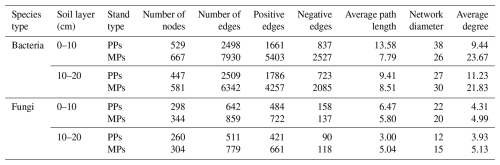
Microbial species with an average abundance of at least 1 % in the 0–10 and 10–20 cm layers of PPs and MPs were selected for network analysis. Significant differences in microbial network structure were found between PPs and MPs in both soil layers (Fig. 4a, b). In the bacterial and fungal networks, there were significantly more nodes in MPs than in PPs (Table 2). Therefore, compared to PPs, MPs significantly stimulated the complexity of the co-occurrence network, particularly in the 0–10 cm. Positive correlations (bacterial networks: ranging = 0.665–0.712, fungal networks: ranging = 0.754–0.849) were determined for both PPs and MPs (Table 2). Compared with PPs, the average path lengths in MPs were shorter (except for the fungal network in the 10–20 cm) and the network diameter was smaller (except for the bacterial network in the 10–20 cm) and had a higher average degree for both the bacterial and the fungal networks in both soil layers (Table 2).
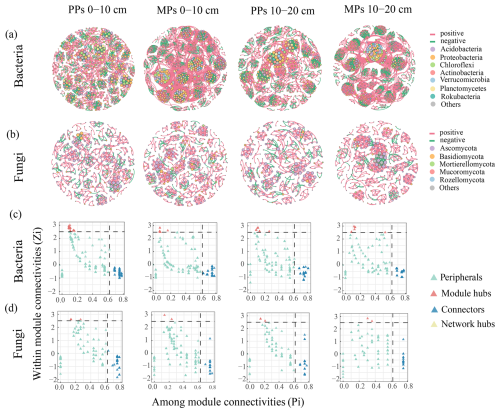
Figure 4Co-occurrence network characteristics of (a) bacterial and (b) fungal communities. The node color represents the phyla with relative abundance greater than 1 %, and the node size represents the degree. The Zi-Pi plot (c–d) predicts keystone OTUs in (c) bacterial and (d) fungal networks.
The Zi–Pi plot showed that network hubs were absent from the bacterial and fungal networks, with keystone species instead concentrated in connectors and module hubs (Fig. 4c, d). Bacterial keystone OTUs were primarily found in the top three phyla, Proteobacteria, Acidobacteriota, and Actinobacteriota (Fig. 4c). Fungal keystone OTUs were likewise concentrated in the top three phyla, Ascomycota, Basidiomycota, and Mucoromycota (Fig. 4d).
3.4 Microbial functional genes involved in N and P transformation and enzyme activity
Introducing Acacia mangium into the Eucalyptus urophylla plantation increased the abundances of functional genes involved in N and P transformation (Figs. 5 and 6). Specifically, the abundances of the N-related functional genes nifH (, P=0.003), AOB-amoA (, P=0.003), narG (, P=0.036), nirS (, P=0.005), and nosZ (, P=0.031) in the 0–10 cm and of AOB-amoA (, P=0.039), narG (, P=0.038), and nirS (, P=0.002) in the 10–20 cm were significantly higher in MPs than in PPs (Fig. 5a–f).
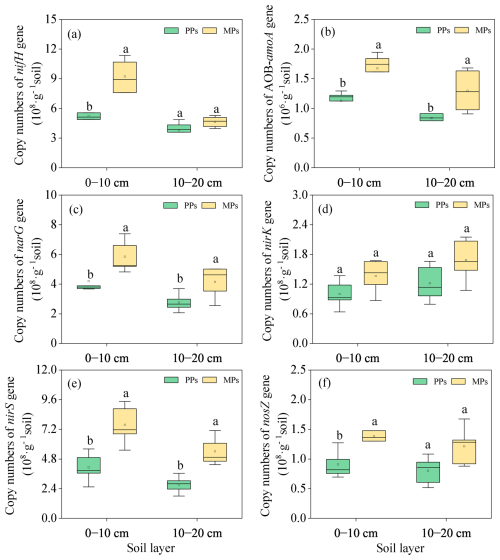
Figure 5Comparison of the abundance of functional genes involved in nitrogen fixation (nifH) (a), nitrification (AOB-amoA) (b), and denitrification [narG (c), nirK (d), nirS (e), and nosZ (f)] in two soil layers in PPs and MPs.
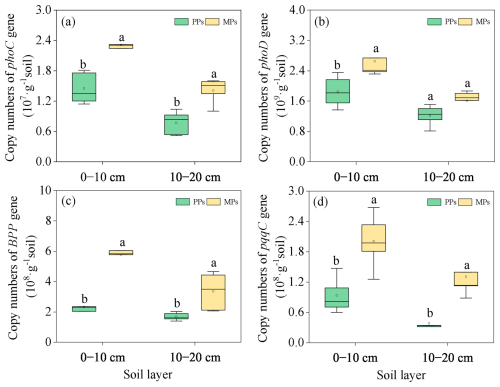
Figure 6Comparison of the abundance of functional genes involved in Po hydrolysis [phoC (a), phoD (b), BPP (c)] and Pi hydrolysis (pqqC) (e) in two layers in PPs and MPs.
The abundances of the P functional genes phoC (0–10 cm: , P=0.003; 10–20 cm: , P=0.003), phoD (0–10 cm: , P=0.020), BPP (0–10 cm: , P<0.001; 10–20 cm: , P=0.018), and pqqC (0–10 cm: , P=0.006; 10–20 cm: , P=0.002) in both soil layers were significantly higher in MPs than in PPs, with the exception of phoD in the 10–20 (Fig. 6).
The EEA analysis results showed that NAG (, P<0.001), LAP (, P=0.035), and ACP (, P=0.001) in the 0–10 cm were significantly higher in MPs than in PPs, by 97.31 %, 31.72 %, and 64.35 % respectively (Fig. 7). In the 10–20 cm, NAG (, P<0.001), LAP (, P=0.012), and ACP (, P=0.003) were also significantly higher in MPs than in PPs, by 24.02 %, 88.54 %, 39.83 %, and 47.72 %, respectively (Fig. 7). The qPCR results showed significantly higher levels of 16S rRNA (0–10 cm: , P<0.001; 10–20 cm: , P=0.002) and ITS (0–10 cm: , P<0.001; 10–20 cm: , P=0.001) in MPs than in PPs (Fig. S3).
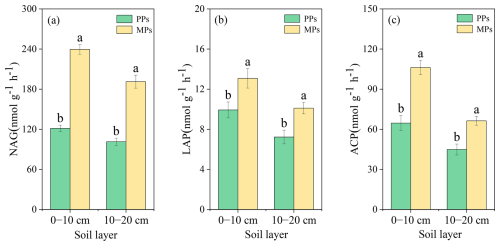
Figure 7Comparisons of extracellular soil enzyme activity of (a) β-1,4-N-acetylglucosaminidase for chitin degradation (NAG); (b) leucine aminopeptidase for protein degradation, (LAP); and (c) acid phosphatase for catalyzing the hydrolysis of phosphate monoesters (ACP) in two soil layers in PPs and MPs.
3.5 Integrating variation in microbial diversity and network complexity with P transformation
The random forest analysis results showed that NAG, LAP, and ACP activities were explained by soil properties, microbial characteristics, and functional genes involved in the N and P cycles to 84.09 %, 58.95 %, and 75.51 %, respectively (Fig. 8). The results showed significant positive correlations for NAG, LAP, and ACP with SOC, TN, NO-N, C:P, N:P, and pH; for the three enzymes with 16S rRNA, ACEbacteria, Chao1bacteria, Shannonbacteria, nodesbacteria, edgesbacteria, and average degreebacteria (P<0.05); for NAG, LAP, and ACP with ITS, Shannonfungi, edgesfungi, and average degreefungi; for LAP and ACP with nodesfungi; for NAG, LAP, and ACP with nifH, AOB-amoA, narG, and nirS; for NAG and LAP with nosZ; and for NAG, LAP, and ACP with phoC, phoD, BPP, and pqqC (all P<0.05). In addition, NAG was significantly negatively correlated with the average path lengthbacteria (P<0.05). Soil physicochemical properties (SOC, TN, NO-N), bacterial community diversity and network complexity, as well as functional genes involved in the N (nifH) and P (phoC) cycles, are strong positive predictors of the variation in EEA.
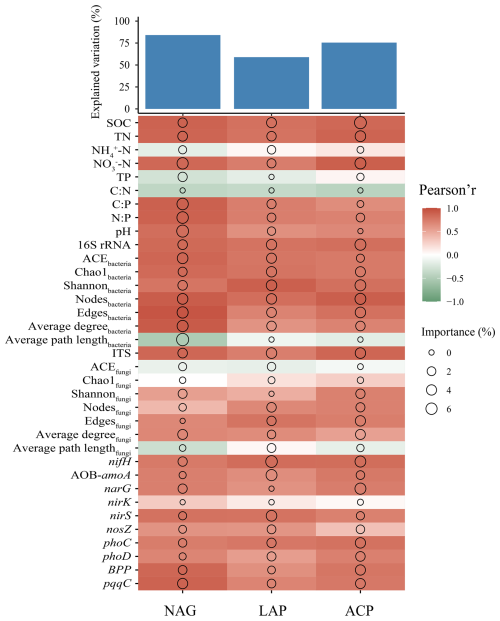
Figure 8The potential biological contributions of soil properties, microbial influences, and functional genes related to N and P cycling to the activity of N and P transformation enzymes. The size of the circles represents the importance of the variables and the color indicates the Pearson correlation.
In the model of P transformation, the variance of 75.7 %, 71.5 %, 96.1 %, 83.9 %, 76.2 %, and 69.5 % could be explained by soil properties, fungal properties, bacterial properties, N functional genes, P functional genes, and N transformation, respectively, within a goodness-of-fit index of 0.782 (Fig. 9a). N transformation and P functional genes (phoC, phoD, and BPP) had a strong direct influence on P transformation, with path coefficients of 0.283 and 0.605, respectively (P<0.01). The diversity and complexity of the network also had favorable effects on N and P functional genes, exerting a substantial influence on P transformation. The overall influence of each factor on P transformation in soil followed the order: soil properties > P functional genes > bacterial properties > N functional genes > fungal properties > N transformation (Fig. 9b). Overall, the mixture of Eucalyptus with N-fixing tree species directly induces alterations in soil properties, which subsequently influence soil microbial characteristics, functional genes involved in N and P cycling, as well as P transformation, ultimately regulating P transformation.
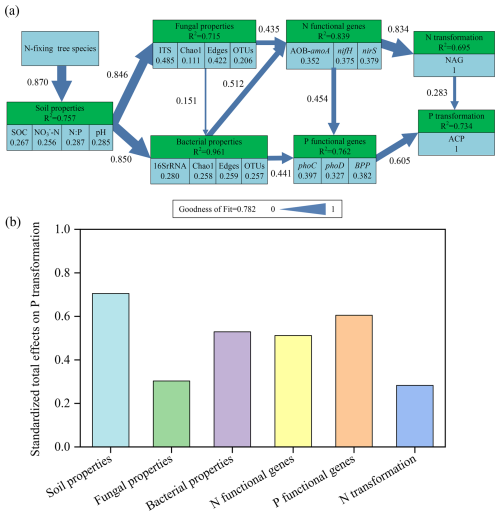
Figure 9(a) Path model describing the control pathways of P transformation (ACP activity) and (b) standardized total effects (including both direct and indirect effects) on P transformation derived from PLS-PM. The light blue in (a) represents the observation variable, the light green represents the latent variable, the number under the observation variable represents the contribution weight of the observation variable to the latent variable, the number and the width of the arrow on the arrow represent the standardized path coefficient between the latent variables, and R2 represents the explanation rate of the model to the latent variable.
4.1 Soil microbial diversity and network response in a mixed plantation of Eucalyptus and N-fixing tree species
The mixed planting of Eucalyptus with N-fixing species significantly impacted the soil microbial community structure, increasing microbial diversity and network complexity. With methodological advances that enable a more comprehensive understanding of soil microbial diversity and networks, we know that soil microorganisms are not only involved in nutrient (e.g., N and P) transformations but also shape the soil habitat by multiple biophysical and biogeochemical processes (Philippot et al., 2024). In our study, the combination of Eucalyptus and N-fixing Acacia mangium enhanced soil nutrient content and altered the stoichiometric ratios of C, N, and P (Table 1). Mixed plantations with N-fixing tree species have higher litter quantity and quality, which enhances nutrient retention and acquisition capacity (Huang et al., 2014), stimulates microbial growth, and promotes microbial aggregation and metabolism, thereby increasing microbial diversity (Guo et al., 2019) (Figs. 1 and S1). These findings align with those of a previous study, which demonstrated that the incorporation of Eucalyptus with N-fixing tree species increased the abundance and diversity of microorganisms, while also revealing variability in community structure across different stands (Li et al., 2023). The composition and diversity of soil microbial communities are primarily driven by ratios (Delgado-Baquerizo et al., 2017). The availability of essential nutrients such as N, P, and Fe are controlled by the soil C supply, while the lower C:N ratio in mixed plantations promotes the formation of various C components, thereby increasing SOC input, which subsequently influences the structure of the microbial communities and their co-occurrence patterns (Yuste et al., 2011; Qiu et al., 2021). Interestingly, in this study, the TP content in MPs was significantly lower than that in PPs (Table 1), which may be a result of increased plant uptake due to higher biomass. Additionally, the high soil N content in MPs with N-fixing tree species may positively influence plant growth, potentially stimulating P uptake (Li et al., 2016). In subtropical regions, characterized by high temperatures and heavy rainfall, P leaching is substantial; however, the introduction of N-fixing tree species increases N content, which may shift the limitation from N to P in MPs. In this context, plants are likely to recycle P more efficiently (See et al., 2015; Lang et al., 2016). Therefore, P returned to the soil through decomposition would be reduced.
In natural habitats, soil microbial communities form intricate arrays and robustly structured networks that allow adaptation to shifting environments (de Vries et al., 2018). The complexity and diversity of microbial communities in soil are fundamental to ecosystem persistence and resilience, as they both reinforce ecological functions and offer a robust defense against external disruptions (Guo et al., 2021). The complexity of the topological structure and connectivity between nodes influences the overall stability of microbial networks and their resilience to environmental disturbances (Yuan et al., 2021). The overwhelming predominance of positive over negative correlations indicated microbial adaptation to similar ecological niches through co-operation (Gao et al., 2022). Networks characterized by higher connectivity and larger numbers of interrelationships are better equipped to withstand environmental changes, thereby preserving the functional stability of the ecosystem (Cornell et al., 2023). Our study showed that N-fixing tree species mixed plantations increased the complexity of bacterial and fungal networks (Fig. 4), as demonstrated by a higher number of nodes and edges, with positive associations predominating over negative ones, indicating stronger interactions between microorganisms (Ma et al., 2020; Niraula, 2021). Random forest analysis also revealed a robust positive association between the number of nodes and the diversity of fungal and bacterial species expressing enzymes responsible for N and P transformation (Fig. 8). These results align with our hypothesis, suggesting that Eucalyptus mixed with N-fixing tree species increases the complexity of microbial networks (Guo and Gong, 2024). The relative abundances of Proteobacteria, Rokubacteria, and Verrucomicrobia in the bacterial community were also higher in MPs than in PPs (particularly in the 0–10 cm), as were the relative abundances of Mortierllomycota, Mucoromycota, and Rozellomycota in the fungal community. Several edaphic factors collectively influenced the structure of both communities, among which pH was the most important (Fig. 3a, b). These findings are in line with earlier research, which demonstrated that soil pH was a key determinant in shaping the structure and composition of microbial communities (Siciliano et al., 2014; Cheng et al., 2020). According to our Zi–Pi plots, the keystone species of the bacterial community were members of phyla Proteobacteria, Acidobacteriota, and Actinobacteria, and those of the fungal community belonged to Ascomycota, Basidiomycota, and Mucoromycota. The ability of leguminous plant species to establish symbiotic associations with root nodule bacteria, commonly referred to as rhizobia, is well established (e.g., Stougaard, 2000; Yang et al., 2022). The phylum Proteobacteria is one of the largest and phenotypically most diverse divisions, which includes gram-negative bacteria such as rhizobia. Furthermore, the N-fixing ability of rhizobia in the phylum Proteobacteria is a key contributor to maintaining the complexity and stability of microbial networks (Sprent and Platzmann, 2001; Fu et al., 2022). Among fungi, Ascomycota is the dominant phylum in soil worldwide (Egidi et al., 2019). In the present study, the relative abundance of Ascomycetes showed dominance in both PPs and MPs, but the relative abundance diminished in MPs. Although keystone taxa may not always be abundant, they play a vital role in shaping microbial communities and maintaining their ecological functions through specific regulatory pathways that affect community structure and function (Banerjee et al., 2018; Liu et al., 2022). For example, a prior study demonstrated that keystone taxa played a critical role in increasing the complexity of microbial networks, enhancing plant health and biomass, and promoting the hydrolysis of organophosphorus compounds through enzymatic activity (Qiao et al., 2024; Zeng et al., 2024).
4.2 Association of microbial diversity and networks with P transformation and key environmental drivers
Our study showed that the abundance of functional genes related to N and P cycles significantly increases after intercropping with N-fixing tree species, which supports our second hypothesis (Figs. 5 and 6). In contrast to this finding, Qin et al. (2024) reported that although planting N-fixing tree species with Eucalyptus enhanced the complexity and stability of N and P functional gene networks, it reduced the abundances of these genes. This discrepancy can be explained by shifts in soil microbial communities related to N and P cycles, which consequently affect the microbial functions that respond to environmental changes (Graham et al., 2016; Zhang et al., 2021). A previous study also found that the microbial community associated with a mixed plantation of Eurograndis and Amangium differed from that associated with monocultures of either species, attributable to positive effects of the mixture on soil P and nitrate levels, which enhanced the abundances of N and P functional genes (Rachid et al., 2013).
Biological N fixation is a fundamental ecosystem process that involves the conversion of atmospheric N into a form usable by plants, which, facilitated by a highly diverse group of microorganisms, significantly enhances soil fertility and promotes plant growth (Burns and Hardy, 2012; Soumare et al., 2020). All N-fixing microorganisms carry functional nifH genes that encode a component of nitrogenase and act as markers of the abundance and diversity of N-fixing microorganisms across various environmental contexts (Wang et al., 2018). Our results indicate that the relative abundance of P functional genes was significantly higher after the introduction of N-fixing tree species compared to pure Eucalyptus plantations (Fig. 6). Both phoC and phoD are functional genes that encode phosphatase activity needed for P solubilization and mineralization and are thus critically involved in promoting soil P availability (Tian et al., 2021; Cao et al., 2022). The P cycling gene pqqC, which encodes the P-mobilizing enzyme pyrroloquinoline quinone synthase, is a marker of phosphate-mobilizing bacteria (Meyer et al., 2011). The predominant bacteria containing phoD and pqqC are primarily members of the Actinobacteria and Proteobacteria (Tan et al., 2013; Hu et al., 2018), whose community structure remained unchanged with an increase in soil P pools (Ragot et al., 2015). We observed a higher abundance and diversity of phoD-, phoC-, and pqqC-bearing soil microorganisms; higher abundances of these genes in soil were correlated with higher soil SOC and TN contents (Luo et al., 2019; Cao et al., 2022). Our study also identified significantly positive correlations between most N and P functional genes and 16S rRNA, as well as the ACE, Chao1, and Shannon indexes in bacterial communities, whereas a significant positive correlation was determined only between the ITS region and the Shannon index in fungal communities (Fig. S4). This variation can be attributed to the significant positive impact that high levels of available nutrients have on the development of bacterial communities in the soil (Ming et al., 2016).
The significant positive correlations detected for the N enzymes NAG and LAP with AOB-amoA, nifH, and the denitrification genes nirS, nosZ, and narG determined in our study suggest that, after the introduction of N-fixing tree species, the microbial community facilitated soil N transformation by increasing the abundance of N cycling genes. Both random forest analysis and PLS-PM analyses indicated that P transformation reflected the interaction of biological and non-biological factors in ecological processes influenced by the introduction of N-fixing tree species (Figs. 8 and 9). Complex interactions between bacteria, fungi, and P cycle genes have been shown to promote microbial community stability while facilitating P transformation processes (Liu et al., 2024). Eucalyptus mixed with N-fixing tree species also increased soil TN and the NH-N content, which increased ACP activity and thus soil Po mineralization. The higher soil pH in MPs than in PPs was likely driven by exchange interactions involving hydroxide minerals and functional groups (Table 1), which enhanced the conversion of potentially labile Pi into plant available P via competitive adsorption (Hinsinger, 2001; Kang et al., 2021).
Together, these results indicate that forest management practices that have Eucalyptus mixed with N-fixing tree species will improve soil physicochemical properties, microbial community diversity, and correlations between microbial N and P cycling genes, thereby promoting soil P transformation.
This study outlines the benefits of incorporating mixed N-fixing tree species with Eucalyptus, specifically highlighting their positive effects on P transformation. The presence of Acacia was shown to alter soil physicochemical properties, improving soil bacterial and fungal community diversity, network complexity, and the abundance of N and P cycling functional genes, ultimately driving P transformation. Increases in soil nutrient content, particularly SOC, TN, and NO-N, as well as the increase in pH that occurred in MPs influenced soil microbial diversity. PLS-PM analysis revealed that mixed plantations have significantly enhanced correlations between P transformation and microbial functional genes that mediate N and P cycling. Our findings offer fresh insights into the predictive capacity of potential shifts in the belowground microbial communities for soil functionality within mixed plantation ecosystems involving N-fixing tree species and Eucalyptus.
The data related to this paper can be downloaded from Zenodo (https://doi.org/10.5281/zenodo.16901157, Li and Huang, 2025).
The supplement related to this article is available online at https://doi.org/10.5194/bg-22-4221-2025-supplement.
JL, XH, and YY conceived and designed the study. JL, XH, YY, and WZ processed and analyzed data from the field experiments. JL, WZ, YL, HH, HM, and QH conducted the fieldwork. JL and WZ performed laboratory analysis. JL completed the analysis of the data and prepared the original draft of the manuscript. XH, YY, YW, and AM helped to review and edit the manuscript. All of the authors gave approval for the final manuscript.
The contact author has declared that none of the authors has any competing interests.
Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims made in the text, published maps, institutional affiliations, or any other geographical representation in this paper. While Copernicus Publications makes every effort to include appropriate place names, the final responsibility lies with the authors.
This research was funded by grants from the National Natural Science Foundation of China (nos. 32171755 and 32471727), the Guangxi Natural Science Foundation (no. 2025GXNSFAA069288), and the scientific research capacity building project for Youyiguan Forest Ecosystem Observation and Research Station of Guangxi (no. AD25069098).
This research was funded by grants from the National Natural Science Foundation of China (nos. 32171755 and 32471727), the Guangxi Natural Science Foundation (no. 2025GXNSFAA069288), and the scientific research capacity building project for Youyiguan Forest Ecosystem Observation and Research Station of Guangxi (no. AD25069098).
This paper was edited by Daniel S. Goll and reviewed by Dejun Li and two anonymous referees.
Adams, R. I., Miletto, M., Taylor, J. W., and Bruns, T. D.: Dispersal in microbes: fungi in indoor air are dominated by outdoor air and show dispersal limitation at short distances, ISME J., 7, 1262–1273, https://doi.org/10.1038/ismej.2013.28, 2013.
Banerjee, S., Schlaeppi, K., and van der Heijden, M. G.: Keystone taxa as drivers of microbiome structure and functioning, Nat. Rev. Microbiol, 16, 567–576, https://doi.org/10.1038/s41579-018-0024-1, 2018.
Benjamini, Y., Krieger, A. M., and Yekutieli, D.: Adaptive linear step-up procedures that control the false discovery rate, Biometrika, 93, 491–507, https://doi.org/10.1093/biomet/93.3.491, 2006.
Bünemann, E. K., Smernik, R. J., Marschner, P., McNeill, A. M.: Microbial synthesis of organic and condensed forms of phosphorus in acid and calcareous soils, Soil Biol. Biochem., 40, 932–946, https://doi.org/10.1016/j.soilbio.2007.11.012, 2008.
Burns, R. C. and Hardy, R. W.: Nitrogen fixation in bacteria and higher plants, Springer Science & Business Media 21:192, 2012.
Cao, N., Zhi, M., Zhao, W., Pang, J., Hu, W., Zhou, Z., and Meng, Y.: Straw retention combined with phosphorus fertilizer promotes soil phosphorus availability by enhancing soil P-related enzymes and the abundance of phoC and phoD genes, Soil Till. Res., 220, 105390, https://doi.org/10.1016/j.still.2022.105390, 2022.
Cheng, J., Zhao, M., Cong, J., Qi, Q., Xiao, Y., Cong, W., Deng, Y., Zhou, J., and Zhang, Y.: Soil pH exerts stronger impacts than vegetation type and plant diversity on soil bacterial community composition in subtropical broad-leaved forests, Plant Soil, 450, 273–286, https://doi.org/10.1007/s11104-020-04507-2, 2020.
Chen, S., Lin, L., Deng, Y., Yuan, S., and Zhang, N.: Tree species richness and mycorrhizal types drive soil nitrogen cycling by regulating soil microbial community composition and diversity in tropical forests, Forest Ecol. Manag., 569, 122187, https://doi.org/10.1016/j.foreco.2024.122187, 2024.
Cornell, C. R., Zhang, Y., Ning, D., Xiao, N., Wagle, P., Xiao, X., and Zhou, J.: Land use conversion increases network complexity and stability of soil microbial communities in a temperate grassland, ISME J., 17, 2210–2220, https://doi.org/10.1038/s41396-023-01521-x, 2023.
Dai, Z., Liu, G., Chen, H., Chen, C., Wang, J., Ai, S., Wei, D., Li, D. Ma, B., Tang, C., Brookes, P. C., and Xu, J.: Long-term nutrient inputs shift soil microbial functional profiles of phosphorus cycling in diverse agroecosystems, ISME J., 14, 757–770, https://doi.org/10.1038/s41396-019-0567-9, 2020.
Pereira, A. P. A., Andrade, P. A. M., de Andrade, P. A. M., Bini, D., Durrer, A., Robin, A., Bouillet, J. P., Andreote, F. D., and Cardoso, E. J. B. N.: Shifts in the bacterial community composition along deep soil profiles in monospecific and mixed stands of Eucalyptus grandis and Acacia mangium, PloS one, 12, e0180371, https://doi.org/10.1371/journal.pone.0180371, 2017.
Delgado-Baquerizo, M., Reich, P. B., Khachane, A. N., Campbell, C. D., Thomas, N., Freitag, T. E., Al-Soud, W. A., Sørensen, S., Bardgett, R. D., and Singh, B. K.: It is elemental: soil nutrient stoichiometry drives bacterial diversity, Environ. Microbiol., 19, 1176–1188, https://doi.org/10.1111/1462-2920.13642, 2017.
Delmas, E., Besson, M., Brice, M. H., Burkle, L. A., Dalla Riva, G. V., Fortin, M. J., Gravel, D., Guimarães, P. R., Guimarães Jr., P. R., Hembry, D. H., Newman, E. A., Olesen, J. M., Pires, M. M., Yeakel, J. D., and Poisot, T.: Analysing ecological networks of species interactions, Biol. Rev., 94, 16–36, https://doi.org/10.1111/brv.12433, 2019.
de Vries, F. T., Griffiths, R. I., Bailey, M., Craig, H., Girlanda, M., Gweon, H. S., Hallin, S., Kaisermann, A., Keith, A. M., Kretzschmar, M., Lemanceau, P., Lumini, E., Mason, K. E., Oliver, A., Ostle, N., Prosser, J. I., Thion, C., Thomson, B., and Bardgett, R. D.: Soil bacterial networks are less stable under drought than fungal networks, Nat. Commun., 9, 3033, https://doi.org/10.1038/s41467-018-05516-7, 2018.
Dong, H., Ge, J., Sun, K., Wang, B., Xue, J., Wakelin, S. A., Wu, J., Sheng, W., Xu, Q., Jiang, P., Chen, J., and Qin, H.: Change in root-associated fungal communities affects soil enzymatic activities during Pinus massoniana forest development in subtropical China, Forest Ecol. Manag., 482, 118817, https://doi.org/10.1016/j.foreco.2020.118817, 2021.
Du, E., Terrer, C., Pellegrini, A. F., Ahlström, A., van Lissa, C. J., Zhao, X., X, N., Wu, X., and Jackson, R. B.: Global patterns of terrestrial nitrogen and phosphorus limitation, Nat. Geosci., 13, 221–226, https://doi.org/10.1038/s41561-019-0530-4, 2020.
Egidi, E., Delgado-Baquerizo, M., Plett, J. M., Wang, J., Eldridge, D. J., Bardgett, R. D., Maestre, F. T., and Singh, B. K.: A few Ascomycota taxa dominate soil fungal communities worldwide, Nat. Commun., 10, 2369, https://doi.org/10.1038/s41467-019-10373-z, 2019.
Epron, D., Nouvellon, Y., Mareschal, L., e Moreira, R. M., Koutika, L. S., Geneste, B., Delgado-Rojas, J, S., Laclau J, P., Sola, G., Gonçalves, J, L, M., and Bouillet, J. P.: Partitioning of net primary production in Eucalyptus and Acacia stands and in mixed-species plantations: two case-studies in contrasting tropical environments, Forest Ecol. Manag., 301, 102–111, https://doi.org/10.1016/j.foreco.2012.10.034, 2013.
Fan, K., Weisenhorn, P., Gilbert, J. A., and Chu, H.: Wheat rhizosphere harbors a less complex and more stable microbial co-occurrence pattern than bulk soil, Soil Biol. and Biochem., 125, 251–260, https://doi.org/10.1016/j.soilbio.2018.07.022, 2018.
Fan, Y., Zhong, X., Lin, F., Liu, C., Yang, L., Wang, M., Chen, G., Chen, Y., and Yang, Y.: Responses of soil phosphorus fractions after nitrogen addition in a subtropical forest ecosystem: Insights from decreased Fe and Al oxides and increased plant roots, Geoderma, 337, 246–255, https://doi.org/10.1016/j.geoderma.2018.09.028, 2019.
Faust, K.: Open challenges for microbial network construction and analysis, ISME J., 15, 3111–3118, https://doi.org/10.1038/s41396-021-01027-4, 2021.
Fraser, T., Lynch, D. H., Entz, M. H., and Dunfield, K. E.: Linking alkaline phosphatase activity with bacterial phoD gene abundance in soil from a long-term management trial, Geoderma, 257, 115–122, https://doi.org/10.1016/j.geoderma.2014.10.016, 2015.
Fu, L., Yan, Y., Li, X., Liu, Y., and Lu, X.: Rhizosphere soil microbial community and its response to different utilization patterns in the semi-arid alpine grassland of northern Tibet, Front. Microbiol, 13, 931795, https://doi.org/10.3389/fmicb.2022.931795, 2022.
Gao, C., Xu, L., Montoya, L., Madera, M., Hollingsworth, J., Chen, L., Purdom, E., Singan, V., Vogel, J., Hutmacher, R. B., Dahlberg, J. A., Coleman-Derr, D., Lemaux, P. G., and Taylor, J. W.: Co-occurrence networks reveal more complexity than community composition in resistance and resilience of microbial communities, Nat. Commun., 13, 3867, https://doi.org/10.1038/s41467-022-31343-y, 2022.
Graham, E. B., Knelman, J. E., Schindlbacher, A., Siciliano, S., Breulmann, M., Yannarell, A., Beman, J. M., Abell, G., Philippot, L., Prosser, J., Foulquier1, A., Yuste, J. C., Glanville, H. C., Jones, D. L., Angel, R., Salminen, J., Newton, R. J., Bürgmann, H., Ingram, L. J., Hamer, U., Siljanen, H. M. P., Peltoniemi, K., Potthast, K., Bañeras, L., Hartmann, M., Banerjee, S., Yu, R. Q., Nogaro, G., Richter, A., Koranda, M., Castle, S. C., Goberna, M., Song, B., Chatterjee, A., Nunes, O. C., Lopes, A. R., Cao, Y., Kaisermann, A., Hallin, S., Strickland, M. S., Garcia-Pausas, J., Barba, J., Kang, H., Isobe, K., Papaspyrou, S., Pastorelli, R., Lagomarsino, A., Lindström, E. S., Basiliko, N., and Nemergut1, D. R.: Microbes as engines of ecosystem function: when does community structure enhance predictions of ecosystem processes?, Front. Microbial, 7, 214, https://doi.org/10.3389/fmicb.2016.00214, 2016.
Guo, Q. and Gong, L.: Compared with pure forest, mixed forest alters microbial diversity and increases the complexity of interdomain networks in arid areas, Microbiology Spectrum, 12, e02642-23, https://doi.org/10.1128/spectrum.02642-23, 2024.
Guo, Y., Hou, L., Zhang, Z., Zhang, J., Cheng, J., Wei, G., and Lin, Y.: Soil microbial diversity during 30 years of grassland restoration on the Loess Plateau, China: Tight linkages with plant diversity, Land Degrad. Dev., 30, 1172–1182, https://doi.org/10.1002/ldr.3300, 2019.
Guo, Y., Xu, T., Cheng, J., Wei, G., and Lin, Y.: Above-and belowground biodiversity drives soil multifunctionality along a long-term grassland restoration chronosequence, Sci. Total Environ., 772, 145010, https://doi.org/10.1016/j.scitotenv.2021.145010, 2021.
He, Y., Wen Y., Li, K., Ye, S., Zhang, H., He, F., Fan, R., and Wu, H.: Responses of soil multifunctionality, microbial diversity, and network complexity to tree species mixing in Eucalyptus plantations, Ind. Crop. Prod., 225, 120575, https://doi.org/10.1016/j.indcrop.2025.120575, 2025.
Hinsinger, P.: Bioavailability of soil inorganic P in the rhizosphere as affected by root-induced chemical changes: a review, Plant Soil, 237, 173–195, https://doi.org/10.1023/A:1013351617532, 2001.
Hu, J. P., Zhang, M. X., Lü, Z. L., He, Y. Y., Yang, X. X., Khan, A., Xiong, Y. C., Fang, X. L., Dong, Q. M., and Zhang, J. L.: Grazing practices affect phyllosphere and rhizosphere bacterial communities of Kobresia humilis by altering their network stability, Sci. Total Environ., 900, 165814, https://doi.org/10.1016/j.scitotenv.2023.165814, 2023.
Hu, Y., Xia, Y., Sun, Q. I., Liu, K., Chen, X., Ge, T., Zhu, B., Zhu, Z., Zhang, Z., Su, Y.: Effects of long-term fertilization on phoD-harboring bacterial community in Karst soils, Sci. Total Environ., 628, 53–63, https://doi.org/10.1016/j.scitotenv.2018.01.314, 2018.
Huang, X., Liu, S., Wang, H., Hu, Z., Li, Z., and You, Y.: Changes of soil microbial biomass carbon and community composition through mixing nitrogen-fixing species with Eucalyptus urophylla in subtropical China, Soil Biol. Biochem., 73, 42–48, https://doi.org/10.1016/j.soilbio.2014.01.021, 2014.
Huang, X., Liu, S., You, Y., Wen, Y., Wang, H., and Wang, J.: Microbial community and associated enzymes activity influence soil carbon chemical composition in Eucalyptus urophylla plantation with mixing N2-fixing species in subtropical China, Plant Soil, 414, 199–212, https://doi.org/10.1007/s11104-016-3117-5, 2017.
Jiao, S., Yang, Y., Xu, Y., Zhang, J., and Lu, Y.: Balance between community assembly processes mediates species coexistence in agricultural soil microbiomes across eastern China, ISME J., 14, 202–216, https://doi.org/10.1038/s41396-019-0522-9, 2020.
Jin, X., Liu, Y., Hu, W., Wang, G., Kong, Z., Wu, L., and Ge, G.: Soil bacterial and fungal communities and the associated nutrient cycling responses to forest conversion after selective logging in a subtropical forest of China, Forest Ecol. Manag., 444, 308–317, https://doi.org/10.1016/j.foreco.2019.04.032, 2019.
Kang, L., Zhang, G., and Chu, G.: Split delivering phosphorus via fertigation to a calcareous soil increased P availability and maize yield (Zea mays L.) by reducing P fixation, J. Soil. Sediment., 21, 2287–2300, https://doi.org/10.1007/s11368-021-02914-1, 2021.
Khan, A., Zhang, G., Li, T., and He, B.: Fertilization and cultivation management promotes soil phosphorus availability by enhancing soil P-cycling enzymes and the phosphatase encoding genes in bulk and rhizosphere soil of a maize crop in sloping cropland, Ecotox. Environ., Safe, 264, 115441, https://doi.org/10.1016/j.ecoenv.2023.115441, 2023.
Knowles, S. C. L., Eccles, R. M., and Baltrūnaitė, L.: Species identity dominates over environment in shaping the microbiota of small mammals, Ecol. Lett., 22, 826–837, https://doi.org/10.1111/ele.13240, 2019.
Koutika, L. S. and Mareschal, L.: Acacia and eucalypt change P, N and C concentrations in POM of Arenosols in the Congolese coastal plains, Geoderma Reg., 11, 37–43, https://doi.org/10.1016/j.geodrs.2017.07.009, 2017.
Koutika, L. S. and Richardson, D. M.: Acacia mangium Willd: benefits and threats associated with its increasing use around the world, For. Ecosyst., 6, 2, https://doi.org/10.1186/s40663-019-0159-1, 2019.
Lang, F., Bauhus, J., Frossard, E., George, E., Kaiser, K., Kaupenjohann, M., Krüger, J., Matzner, E., Polle, A., Prietzel, J., Rennenberg, H., and Wellbrock, N.: Phosphorus in forest ecosystems: new insights from an ecosystem nutrition perspective, J. Plant Nutr. Soil Sc., 179, 129–135, https://doi.org/10.1002/jpln.201500541, 2016.
Li, J. and Huang, X.: Data set for “Soil microbial diversity and network complexity promote phosphorus transformation – a case of long-term mixed plantations of Eucalyptus and a nitrogen-fixing tree species”, in: Biogeosciences, Guangxi University, Zenodo [data set], https://doi.org/10.5281/zenodo.16901157, 2025.
Li, C., Xu, Y., Wang, Z., Zhu, W., and Du, A.: Mixing planting with native tree species reshapes soil fungal community diversity and structure in multi-generational eucalypt plantations in southern China, Front. Microbiol. Fro., 14, 1132875, https://doi.org/10.3389/fmicb.2023.1132875, 2023.
Li, M., You, Y., Tan, X., Wen, Y., Yu, S., Xiao, N., Shen, W., and Huang, X.: Mixture of N2-fixing tree species promotes Po accumulation and transformation in topsoil aggregates in a degraded karst region of subtropical China, Geoderma, 413, 115752, https://doi.org/10.1016/j.geoderma.2022.115752, 2022.
Li, N., Zhang, Y., Qu, Z., Liu, B., Huang, L., Ming, A., and Sun, H.: Mixed and continuous cropping eucalyptus plantation facilitated soil carbon cycling and fungal community diversity after a 14-year field trail, Ind. Crop. Prod., 210, 118157, https://doi.org/10.1016/j.indcrop.2024.118157, 2024.
Li, Q., Lv, J., Peng, C., Xiang, W., Xiao, W., and Song, X.: Nitrogen-addition accelerates phosphorus cycling and changes phosphorus use strategy in a subtropical Moso bamboo forest, Environ. Res. Lett., 16, 024023, https://doi.org/10.1088/1748-9326/abd5e1, 2021.
Li, Y., Niu, S., and Yu, G.: Aggravated phosphorus limitation on biomass production under increasing nitrogen loading: A meta-analysis, Glob. Change Biol., 22, 934-943, https://doi.org/10.1111/gcb.13125, 2016.
Liu, S., Yu, H., Yu, Y., Huang, J., Zhou, Z., Zeng, J., Chen, P., Xiao, F., He, Z., and Yan, Q.: Ecological stability of microbial communities in Lake Donghu regulated by keystone taxa, Ecol. Indic., 136, 108695, https://doi.org/10.1016/j.ecolind.2022.108695, 2022.
Liu, S., Li, H., Xie, X., Chen, Y., Lang, M., and Chen, X.: Long-term moderate fertilization increases the complexity of soil microbial community and promotes regulation of phosphorus cycling genes to improve the availability of phosphorus in acid soil, Appl. Soil Ecol., 194, 105178, https://doi.org/10.1016/j.apsoil.2023.105178, 2024.
Lu, X., Wen, L., Sun, H., Fei, T., Liu, H., Ha, S., Tang, S., and Wang, L.: Responses of soil respiration to phosphorus addition in global grasslands: A meta-analysis, J. Clean. Prod., 349, 131413, https://doi.org/10.1016/j.jclepro.2022.131413, 2022.
Luo, G., Sun, B., Li, L., Li, M., Liu, M., Zhu, Y., Guo, S., Ling, N., and Shen, Q.: Understanding how long-term organic amendments increase soil phosphatase activities: insight into phoD-and phoC-harboring functional microbial populations, Soil Biol. Biochem., 139, 107632, https://doi.org/10.1016/j.soilbio.2019.107632, 2019.
Ma, B., Wang, Y., Ye, S., Liu, S., Stirling, E., Gilbert, J. A., Faust, K., Knight, R., Jansson, J. K., Cardona, C., Röttjers, L., and Xu, J.: Earth microbial co-occurrence network reveals interconnection pattern across microbiomes, Microbiome, 8, 82, https://doi.org/10.1186/s40168-020-00857-2, 2020.
Meyer, J. B., Frapolli, M., Keel, C., and Maurhofer, M.: Pyrroloquinoline quinone biosynthesis gene pqqC, a novel molecular marker for studying the phylogeny and diversity of phosphate-solubilizing pseudomonads, Appl. Environ. Microb., 77, 7345–7354, https://doi.org/10.1128/AEM.05434-11, 2011.
Ming, L. I., Ming, L. I. U., Li, Z. P., Jiang, C. Y., and Meng, W. U.: Soil N transformation and microbial community structure as affected by adding biochar to a paddy soil of subtropical China, J. Integr. Agr., 15, 209–219, https://doi.org/10.1016/S2095-3119(15)61136-4, 2016.
Monecke, A. and Leisch, F.: semPLS: structural equation modeling using partial least squares, J. Stat. Softw., 48, 1–32, https://doi.org/10.18637/jss.v048.i03, 2012.
Mori, H., Maruyama, F., Kato, H., Toyoda, A., Dozono, A., Ohtsubo, Y., Nagata, Y., Fujiyama, A., Tsuda, M., and Kurokawa, K.: Design and experimental application of a novel non-degenerate universal primer set that amplifies prokaryotic 16S rRNA genes with a low possibility to amplify eukaryotic rRNA genes, DNA Res., 21, 217–227, https://doi.org/10.1093/dnares/dst052, 2014.
Murphy, J. and Riley, J. P.: A modified single solution method for the determination of phosphate in natural waters, Anal. Chim. Acta, 27, 31–36, https://doi.org/10.1016/S0003-2670(00)88444-5, 1962.
Nannipieri, P., Giagnoni, L., Renella, G., Puglisi, E., Ceccanti, B., Masciandaro, G., Fornasier, F., Moscatelli, M. C., and Marinari, S.: Soil enzymology: classical and molecular approaches, Biol. Fert. Soil., 48, 743–762, https://doi.org/10.1007/s00374-012-0723-0, 2012.
Niraula, S.: Effects of a N2-Fixing Biofertilizer on the Rhizosphere Microbiome and the Influence of Biochar on Soil Fertility and Microbial Communities, The University of Texas at Arlington, 2021.
Oksanen, J., Blanchet, F. G., Kindt, R., Legendre, P., Minchin, P. R., O'hara, R. B., Solymos, P., Stevens, M. H. H., Szoecs, E., Wagner, H., Barbour, M., Bedward, M., Bolker, B., Borcard, D., Carvalho, G., Chirico, M., Caceres, M. D., Durand, S., Evangelista, H. B. A., FitzJohn, R., Friendly, M., Furneaux, B., Hannigan, G., Hill, M. O., Lahti, L., McGlinn, D., Ouellette, M. H., Cunha, E. R., Smith, T., Stier, A., Braak, C. J. K. T., and Weedon, J.: Package “vegan”, Community ecology package, version, 2, 1–295, https://github.com/vegandevs/vegan (last access: 29 April 2025), 2013.
Parada, A. E., Needham, D. M., and Fuhrman, J. A.: Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples, Environ. Microbiol., 18, 1403–1414, https://doi.org/10.1111/1462-2920.13023, 2016.
Pastore, G., Kaiser, K., Kernchen, S., and Spohn, M.: Microbial release of apatite-and goethite-bound phosphate in acidic forest soils, Geoderma, 370, 114360, https://doi.org/10.1016/j.geoderma.2020.114360, 2020.
Paula, R. R., Bouillet, J. P., Trivelin, P. C. O., Zeller, B., de Moraes Gonçalves, J. L., Nouvellon, Y., Bouvet, J, M., Plassard, C., and Laclau, J. P.: Evidence of short-term belowground transfer of nitrogen from Acacia mangium to Eucalyptus grandis trees in a tropical planted forest, Soil Biol. Biochem., 91, 99–108, https://doi.org/10.1016/j.soilbio.2015.08.017, 2015.
Peng, Y., Duan, Y., Huo, W., Xu, M., Yang, X., Wang, X., Wang, B., Blackwell, M. S. A., and Feng, G.: Soil microbial biomass phosphorus can serve as an index to reflect soil phosphorus fertility, Biol. Fert. Soils, 57, 657–669, https://doi.org/10.1007/s00374-021-01559-z, 2021.
Pereira, A. P. A., Durrer, A., Gumiere, T., Gonçalves, J. L. M., Robin, A., Bouillet, J. P., Wang, J., Verma, J. P., Singh, B. K., and Cardoso, E. J. B. N.: Mixed Eucalyptus plantations induce changes in microbial communities and increase biological functions in the soil and litter layers, Forest Ecol. Manag., 433, 332–342, https://doi.org/10.1016/j.foreco.2018.11.018, 2019.
Philippot, L., Chenu, C., Kappler, A., Rillig, M. C., and Fierer, N.: The interplay between microbial communities and soil properties, Nat. Rev. Microbiol., 22, 226–239, https://doi.org/10.1038/s41579-023-00980-5, 2024.
Poudel, R., Jumpponen, A., Schlatter, D. C., Paulitz, T. C., Gardener, B. M., Kinkel, L. L., and Garrett, K. A.: Microbiome networks: a systems framework for identifying candidate microbial assemblages for disease management, Phytopathology, 106, 1083–1096, https://doi.org/10.1094/PHYTO-02-16-0058-FI, 2016.
Qiao, Y., Wang, T., Huang, Q., Guo, H., Zhang, H., Xu, Q., Shen, Q., and Ling, N.: Core species impact plant health by enhancing soil microbial cooperation and network complexity during community coalescence, Soil Biol. Biochem., 188, 109231, https://doi.org/10.1016/j.soilbio.2023.109231, 2024.
Qin, F., Yang, F., Ming, A., Jia, H., Zhou, B., Xiong, J., and Lu, J.: Mixture enhances microbial network complexity of soil carbon, nitrogen and phosphorus cycling in Eucalyptus plantations, Forest Ecol. Manag., 553, 121632, https://doi.org/10.1016/j.foreco.2023.121632, 2024.
Qiu, L., Zhang, Q., Zhu, H., Reich, P. B., Banerjee, S., van der Heijden, M. G., Sadowsky, M. J., Ishii, S., Jia, X., Shao, M., Liu, B., Jiao, H., Li, H., and Wei, X.: Erosion reduces soil microbial diversity, network complexity and multifunctionality, ISME J., 15, 2474–2489, https://doi.org/10.1038/s41396-021-00913-1, 2021.
Rachid, C. T., Balieiro, F. D. C., Peixoto, R. S., Pinheiro, Y. A. S., Piccolo, M. D. C., Chaer, G. M., and Rosado, A. S.: Mixed plantations can promote microbial integration and soil nitrate increases with changes in the N cycling genes, Soil Biol. Biochem., 66, 146–153, https://doi.org/10.1016/j.soilbio.2013.07.005, 2013.
Ragot, S. A., Kertesz, M. A., and Bünemann, E. K.: phoD alkaline phosphatase gene diversity in soil, Appl. Environ. Microb., 81, 7281–7289, https://doi.org/10.1128/AEM.01823-15, 2015.
Sanchez, G.: PLS path modeling with R. Berkeley: Trowchez Editions, http://www.gastonsanchez.com/PLS (last access: 7 October 2024) Path Modeling with R.pdf, 2013.
Schloss, P. D., Westcott, S. L., Ryabin, T., Hall, J. R., Hartmann, M., Hollister, E. B., Lesniewski, R. A., Oakley, B. B., Parks, D. H., Robinson, C. J., Sahl, J. W., Stres, B., Thallinger, G. G., Horn, D. J. V., and Weber, C. F.: Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities, Appl. Environ. Microb., 75, 7537–7541, https://doi.org/10.1128/AEM.01541-09, 2009.
See, C. R., Yanai, R. D., Fisk, M. C., Vadeboncoeur, M. A., Quintero, B. A., and Fahey, T. J.: Soil nitrogen affects phosphorus recycling: foliar resorption and plant–soil feedbacks in a northern hardwood forest, Ecology, 96, 2488–2498, https://doi.org/10.1890/15-0188.1, 2015.
Shi, Y., Delgado Baquerizo, M., Li, Y., Yang, Y., Zhu, Y. G., Peñuelas, J., and Chu, H.: Abundance of kinless hubs within soil microbial networks are associated with high functional potential in agricultural ecosystems, Environ. Int., 142, 105869, https://doi.org/10.1016/j.envint.2020.105869, 2020.
Siciliano, S. D., Palmer, A. S., Winsley, T., Lamb, E., Bissett, A., Brown, M. V., van Dorst, J., Ji, M., Ferrari, B. C., Grogan, P., Chu, H., and Snape, I.: Soil fertility is associated with fungal and bacterial richness, whereas pH is associated with community composition in polar soil microbial communities, Soil Biol. Biochem., 78, 10–20, https://doi.org/10.1016/j.soilbio.2014.07.005, 2014.
Soltangheisi, A., Withers, P. J., Pavinato, P. S., Cherubin, M. R., Rossetto, R., Do Carmo, J. B., da Rocha, G., C., and Martinelli, L. A.: Improving phosphorus sustainability of sugarcane production in Brazil, GCB Bioenergy, 11, 1444–1455, https://doi.org/10.1111/gcbb.12650, 2019.
Soumare, A., Diedhiou, A. G., Thuita, M., Hafidi, M., Ouhdouch, Y., Gopalakrishnan, S., and Kouisni, L.: Exploiting biological nitrogen fixation: a route towards a sustainable agriculture, Plants, 9, 1011, https://doi.org/10.3390/plants9081011, 2020.
Sprent, J. I. and Platzmann, J.: Nodulation in legumes, Kew: Royal Botanic Gardens, 2001.
Stougaard, J.: Regulators and regulation of legume root nodule development, Plant Physiol., 124, 531–540, https://doi.org/10.1104/pp.124.2.531, 2000.
Sun, H., Wu, Y., Zhou, J., Yu, D., and Chen, Y.: Microorganisms drive stabilization and accumulation of organic phosphorus: An incubation experiment, Soil Biol. Biochem., 172, 108750, https://doi.org/10.1016/j.soilbio.2022.108750, 2022.
Tan, H., Barret, M., Mooij, M. J., Rice, O., Morrissey, J. P., Dobson, A., Griffiths, B., and O'Gara, F.: Long-term phosphorus fertilisation increased the diversity of the total bacterial community and the phoD phosphorus mineraliser group in pasture soils, Biol. Fert. Soils, 49, 661–672, https://doi.org/10.1007/s00374-012-0755-5, 2013.
Tchichelle, S. V., Epron, D., Mialoundama, F., Koutika, L. S., Harmand, J. M., Bouillet, J. P., and Mareschal, L.: Differences in nitrogen cycling and soil mineralisation between a eucalypt plantation and a mixed eucalypt and Acacia mangium plantation on a sandy tropical soil, South, Forests, 79, 1–8, https://doi.org/10.2989/20702620.2016.1221702, 2017.
Tenenhaus, M., Amato, S., and Esposito Vinzi, V.: A global goodness-of-fit index for PLS structural equation modeling. In Proceedings of the XLII SIS Scientific Meeting, 739–742, Padova, CLEUP, https://www.researchgate.net/publication/284462849 (last accesds: 7 October 2024), 2004.
Tian, J., Ge, F., Zhang, D., Deng, S., and Liu, X.: Roles of phosphate solubilizing microorganisms from managing soil phosphorus deficiency to mediating biogeochemical P cycle, Biology, 10, 158, https://doi.org/10.3390/biology10020158, 2021.
Tsiknia, M., Tzanakakis, V. A., Oikonomidis, D., Paranychianakis, N. V., and Nikolaidis, N. P.: Effects of olive mill wastewater on soil carbon and nitrogen cycling, Appl. Microbiol. Biot., 98, 2739–2749, https://doi.org/10.1007/s00253-013-5272-4, 2014.
Turner, B., Brenes Arguedas, T., and Condit, R.: Pervasive phosphorus limitation of tree species but not communities in tropical forests, Nature, 555, 367–370, https://doi.org/10.1038/nature25789, 2018.
Waithaisong, K., Robin, A., l'Huillery, V., Abadie, J., Sauvage, F. X., Chemardin, P., Mareschal, L., Bouillet, J.-P., Gonçalves, J. L. M., and Plassard, C.: Organic phosphorus immobilization in microbial biomass controls how N2-fixing trees affect phosphorus bioavailability in two tropical soils, Environmental Advances, 8, 100247, https://doi.org/10.1016/j.envadv.2022.100247, 2022.
Wang, Q., Wang, J., Li, Y., Chen, D., Ao, J., Zhou, W., Shen, D., Li, Q., Huang, Z., and Jiang, Y.: Influence of nitrogen and phosphorus additions on N2-fixation activity, abundance, and composition of diazotrophic communities in a Chinese fir plantation, Sci. Total Environ., 619, 1530–1537, https://doi.org/10.1016/j.scitotenv.2017.10.064, 2018.
Wang, Y., Luo, D., Xiong, Z., Wang, Z., and Gao, M.: Changes in rhizosphere phosphorus fractions and phosphate-mineralizing microbial populations in acid soil as influenced by organic acid exudation, Soil Till. Res., 225, 105543, https://doi.org/10.1016/j.still.2022.105543, 2023.
Widdig, M., Heintz-Buschart, A., Schleuss, P. M., Guhr, A., Borer, E. T., Seabloom, E. W., and Spohn, M.: Effects of nitrogen and phosphorus addition on microbial community composition and element cycling in a grassland soil, Soil Biol. Biochem, 151, 108041, https://doi.org/10.1016/j.soilbio.2020.108041, 2020.
Yan, J., Huang, X., Su, X., Zhang, W., Gao, G., and You, Y.: Introducing N2-Fixing Tree Species into Eucalyptus Plantation in Subtropical China Alleviated Carbon and Nitrogen Constraints within Soil Aggregates, Forests, 13, 2102, https://doi.org/10.3390/f13122102, 2022.
Yang, J., Lan, L., Jin, Y., Yu, N., Wang, D., and Wang, E.: Mechanisms underlying legume–rhizobium symbioses, J. Integr. Plant Biol., 64, 244–267, https://doi.org/10.1111/jipb.13207, 2022.
Yao, X., Zhang, Q., Zhou, H., Zhu, H., Nong, Z., Ye, S., and Deng, Q.: Introduction of Dalbergia odorifera enhances nitrogen absorption on Eucalyptus through stimulating microbially mediated soil nitrogen-cycling, For. Ecosyst., 8, 59, https://doi.org/10.1186/s40663-021-00339-3, 2021.
Yao, X., Hui, D., Xing, S., Zhang, Q., Chen, J., Li, Z., Xu, Y., and Deng, Y.: Mixed plantations with N-fixing tree species maintain ecosystem stoichiometry: Implication for sustainable production, Soil Biol. Biochem., 191, 109356, https://doi.org/10.1016/j.soilbio.2024.109356, 2024.
Yu, Q., Ma, S., Ni, X., Ni, X., Guo, Z., Tan, X., Zhong, M., Hanif, MA., Zhu, J., Ji, C., and Zhu, B.: Long-term phosphorus addition inhibits phosphorus transformations involved in soil arbuscular mycorrhizal fungi and acid phosphatase in two tropical rainforests, Geoderma, 425, 116076, https://doi.org/10.1016/j.geoderma.2022.116076, 2022.
Yuan, M. M., Guo, X., Wu, L., Zhang, Y. A., Xiao, N., Ning, D., Shi, Z., Zhou, X., Wu, L., Yang, Y., Tiedje, J. M., and Zhou, J.: Climate warming enhances microbial network complexity and stability, Nat. Clim. Change, 11, 343–348, https://doi.org/10.1038/s41558-021-00989-9, 2021.
Yuste, J. C., Penuelas, J., Estiarte, M., Garcia-Mas, J., Mattana, S., Ogaya, R., Pujol, M., and Sardans, J.: Drought-resistant fungi control soil organic matter decomposition and its response to temperature, Glob. Change Biol., 17, 1475–1486, https://doi.org/10.1111/j.1365-2486.2010.02300.x, 2011.
Zeng, Q., Peñuelas, J., Sardans, J., Zhang, Q., Zhou, J., Yue, K., Chen, C., Yang, Y., and Fan, Y.: Keystone bacterial functional module activates P-mineralizing genes to enhance enzymatic hydrolysis of organic P in a subtropical forest soil with 5-year N addition, Soil Biol. Biochem., 192, 109383, https://doi.org/10.1016/j.soilbio.2024.109383, 2024.
Zhang, M., O'Connor, P. J., Zhang, J., and Ye, X.: Linking soil nutrient cycling and microbial community with vegetation cover in riparian zone, Geoderma, 384, 114801, https://doi.org/10.1016/j.geoderma.2020.114801, 2021.
Zhang, W., You, Y., Su, X., Yan, J., Gao, G., Ming, A., Shen, W., and Huang, X.: Introducing N2-fixing tree species into Eucalyptus plantations promotes soil organic carbon sequestration in aggregates by increasing microbial carbon use efficiency, Catena, 231, 107321, https://doi.org/10.1016/j.catena.2023.107321, 2023.
Zhang, Y. and Wang, X.: Geographical spatial distribution and productivity dynamic change of eucalyptus plantations in China, Sci. Rep.-UK, 11, 19764, https://doi.org/10.1038/s41598-021-97089-7, 2021.
Zhou, Y., Boutton, T. W., and Wu, X. B.: Soil phosphorus does not keep pace with soil carbon and nitrogen accumulation following woody encroachment, Glob. Change Biol., 24, 1992–2007, https://doi.org/10.1111/gcb.14048, 2018.