the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Marine eukaryote community responses to the climate and oceanographic changes in Storfjordrenna (southern Svalbard) over the past ∼ 13.3 kyr BP: insights from sedimentary ancient DNA analysis
Magdalena Łącka
Marek Zajączkowski
Dhanushka Devendra
Ngoc-Loi Nguyen
Jan Pawłowski
Joanna Pawłowska
Sedimentary ancient DNA (sedaDNA) metabarcoding is an emerging method for reconstructing the responses of marine organisms to past climate and oceanographic changes, including rare and non-fossilized taxa. Marine sedaDNA records from the Arctic are scarce, especially those focusing on the impact of environmental shifts on the biodiversity and functional composition of marine eukaryote communities. Here, we present a sedaDNA eukaryotic record from a sediment core retrieved in Storfjordrenna, southern Svalbard, spanning the termination of the Bølling–Allerød, the Younger Dryas, and the Holocene (13.3–1.3 kyr BP). We successfully recovered the eukaryotic communities and identified them by their ecological roles. Our study showed that the eukaryotic biodiversity in Storfjordrenna remained relatively stable, except during transitions between major climatic intervals. These shifts were characterized by changes in richness and relative abundance, driven by factors such as perennial ice cover, surface water cooling, and subsurface Atlantic water influx. Cercozoans and Marine Stramenopiles (MAST) emerged as dominant heterotrophs, characterized by high ecological flexibility and broad tolerance. Primary productivity was primarily driven by Arctic water (ArW) associated phytoplankton, including diatoms (Thalassiosira and Chaetoceros), green algae (Micromonas), and autotrophic dinoflagellates (Polarella glacialis) as well as the mixoplanktonic silicoflagellate Pseudopedinella elastica. Amplicon sequence variant (ASV)-based indicator analysis revealed that uncultured cercozoan lineages and MAST taxa were primarily associated with Atlantic water (AW) proxies, whereas parasitic dinoflagellates (Dino-group I) and choanoflagellates were more closely aligned with ArW proxies. Analysis of indicator responses shows the complex interactions within eukaryotic communities, and reveals a strong association among functional ecological groups that impact ecosystem productivity and regulation. This complexity highlights the limitations of traditional single-proxy approaches to accurately reconstructing paleoenvironmental conditions. Our study demonstrates the potential of high-resolution marine sedaDNA metabarcoding in elucidating responses to past climate changes, improving our understanding of the intricate interactions within eukaryotic communities, and enhancing our knowledge of marine ecosystems.
- Article
(4926 KB) - Full-text XML
-
Supplement
(2819 KB) - BibTeX
- EndNote
The Arctic marine ecosystem is undergoing rapid and profound changes, primarily driven by climate warming (IPCC, 2023; Polyakov et al., 2017, 2020). A prominent feature of these changes is the increased influx of AW into the region, a phenomenon known as Atlantification. This process is associated with warming and rise of sea surface temperatures (SST), reduced sea-ice cover, altered salinity patterns, and changes in nutrient dynamics (Årthun et al., 2012; Polyakov et al., 2017).
These transformations in the marine environment are altering the biodiversity of the Arctic region and impacting the function and resilience of its ecosystems (Benner et al., 2019; Hallegraeff, 2010; Ribeiro et al., 2024). The Storfjordrenna region in southern Svalbard is an ideal location to study these shifts, having experienced significant climate-driven changes over the last ∼ 14 000 years, driven by meltwater discharge and the interaction between cold ArW and warmer AW inflows (Łącka et al., 2019, 2015; Pawłowska et al., 2020; Telesiński et al., 2024). The region's biodiversity has been shaped by, and remains sensitive to, these fluctuating and dynamic environmental conditions (Bensi et al., 2024; Deb and Bailey, 2023; Górska et al., 2022; Hop et al., 2019). Understanding how biodiversity adapts to such changes is essential for reconstructing past ecological responses to climate change and for predicting future trends. Although the impact of climate change on Arctic marine ecosystems is well documented (Deb and Bailey, 2023; Wassmann et al., 2010), relatively few studies have examined marine ecosystems using sedaDNA to analyze long-term biodiversity patterns (Grant et al., 2024; Pawłowska et al., 2020; Zimmermann et al., 2023). Recent developments in sedaDNA techniques have increased our ability to extract and analyze DNA from marine environments, providing valuable insights into eukaryotic communities and their responses to environmental changes over geological time scales (Grant et al., 2024; Harðardóttir et al., 2024; Zimmermann et al., 2021). Moreover, several studies have suggested that the sedaDNA or environmental DNA (eDNA) approach has the potential to use specific ASVs as proxies/bioindicators, even when their taxonomy is unknown thereby strengthening the connection between ASVs and ecological functions and environmental changes (Grant et al., 2024; Harðardóttir et al., 2024; Li et al., 2023; Lin et al., 2022; Pawłowska et al., 2020; Perret-Gentil et al., 2021, 2017; Zimmermann et al., 2021). Studies of marine eukaryotic sedaDNA have demonstrated that even low-resolution records can provide significant data on the shifts in marine communities over time, offering insight into past ecosystem dynamics (Grant et al., 2024). For example, recent studies have demonstrated that the sedaDNA approach can be used to reconstruct interactions between sea ice cover, ocean temperatures, and eukaryotic community composition (Armbrecht, 2020; Grant et al., 2024; Harðardóttir et al., 2024; Zimmermann et al., 2023, 2021). However, there is still a significant lack of suitable-resolution marine eukaryotic sedaDNA records from the Arctic, especially those focusing on the, ecosystem-oriented approaches that integrate ecological roles and biotic interactions to better link past biodiversity changes with ecosystem functioning and environmental changes.
This study addresses this issue by reconstructing the long-term history of marine eukaryotic communities from Storfjordrenna, southern Svalbard, using sedaDNA metabarcoding using V1V2 primers that capture both planktonic and benthic taxa with adequate taxonomic resolution for ecological interpretation (Fonseca et al., 2010; Lindeque et al., 2013; Sinniger et al., 2016). The sedaDNA record is supported by previously published sedimentological, micropaleontological, and geochemical records (Łącka et al., 2019, 2020, 2015; Telesiński et al., 2024). By focusing on eukaryotic communities associated with ArW and AW masses, we aim to assess their structure, ecological roles, and potential as indicators of past environmental conditions. Our approach seeks to identify how marine ecosystem have responded to significant climate-driven changes in this region and how these responses can improve our understanding of the future trajectories of Arctic marine biodiversity in the context of ongoing climate warming.
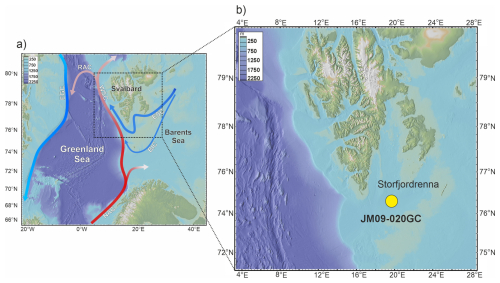
Figure 1(a) Map of the study area and (b) location of core JM09-020GC (yellow dot). Red arrows indicate warm currents, and blue arrows indicate cold currents. Abbreviations: NAC: North Atlantic Current, WSC: West Spitsbergen Current, RAC: Return Atlantic Current, ESC: East Spitsbergen Current, BIC: Bear Island Current, EGC: East Greenland Current.
Storfjorden is an open fjord system located in the Svalbard archipelago, between the islands of Spitsbergen, Barentsøya, and Edgeøya (Fig. 1a). The cross-shelf through Storfjordrenna is located south of Storfjorden's mouth. The hydrography of Storfjorden and Storfjordrenna is primarily governed by the interplay of two major water masses: AW and ArW. AW is characterized by relatively high temperatures (> 3 °C) and high salinity (> 34.95 PSU), whereas ArW exhibits lower temperatures (< 0.5 °C) and salinity ranging from 34.3 to 34.8 PSU (Bensi et al., 2024; Skogseth et al., 2020; Sundfjord et al., 2017). AW is transported northwards by the Norwegian Atlantic Current, which bifurcates upon entering the Barents Sea into the West Spitsbergen Current and the North Cape Current (Blindheim and Østerhus, 2005). In contrast, ArW enters the region via the East Spitsbergen Current and the Bear Island Current, bringing cold, less saline waters into the Barents Sea (Hopkins, 1991). AW enters Storfjordrenna in a cyclonic manner, following the bathymetry. The Polar Front, which separates AW and ArW water masses, is located along the slope of Storfjordrenna (Bensi et al., 2024). The biological and geochemical signals preserved in the sediments represent an integrated response to both gradual and abrupt climatic and oceanographic changes (Łącka et al., 2019, 2015). Therefore, the site's proximity to the Arctic Front facilitates the detection of subtle ecosystem responses to long-term warming and changing oceanographic regimes.
3.1 Sediment core and age model
Gravity core JM09-020-GC was collected in 2009 at a depth of 253 m in Storfjordrenna, northwestern Barents Sea during the expedition of R/V Jan Mayen (Fig. 1b). The core was stored and processed according to the methods described by (Łącka et al., 2019, 2020, 2015). The chronology of the core was established based on AMS14C radiocarbon dating and one additional tie point defined by the appearance of vivianite micro-concretions in a sediment layer set to 12.8 kyr BP, coinciding with the onset of the Younger Dryas (Table S1 in the Supplement) (Łącka et al., 2020). The dates published first by Łącka et al. (2015), and Łącka et al. (2020) were recalibrated using the Marine20 calibration curve (Fig. S1 in the Supplement, Table S1) (Heaton et al., 2020). The palaeoceanographic history of Storfjordrenna over the past ∼ 14 000 years is well documented through detailed, multi-proxy reconstructions. These include analyses of fossil foraminifera assemblages, isotopic composition of foraminiferal tests, grainsize and elemental composition of sediments, alkenones (Łącka et al., 2019, 2015), and dinoflagellate cysts (Telesiński et al., 2024).
3.2 sedaDNA workflow
3.2.1 DNA extraction, amplification, and sequencing
Approximately 10 g of sediment was collected from 55 sediment layers using sterile spoons and transferred to sterile containers. DNA extractions were performed using the DNeasy PowerMax Soil Kit (Qiagen), following the manufacturer's instructions. All DNA extracts were stored at −20 °C until PCR amplification.
The V1V2 region of the 18S rDNA (with a length of ∼ 340 bp) was amplified by PCR using the forward primer SSU_FO4mod (5′-GCT TGW CTC AAA GAT TAA GCC-3′) and the reverse primer SSU_R22 (3′-CCT GCT GCC TTC CTT RGA-5′) (Fonseca et al., 2010; Lindeque et al., 2013), which were tagged with a unique 8-nucleotide sequence at their 5′ ends (Esling et al., 2015). We followed the protocols established and positively tested by Pawłowska et al. (2020, 2014) and Lejzerowicz et al. (2013), with minor adjustments to the number of PCR cycles based on sample-specific amplification performance. Each sample was amplified in triplicate and each PCR reaction was performed in a total volume of 25 µL, which included 1.5 µL of 1.5 mM MgCl2 (Applied Biosystems, USA), 2.5 µL of 10× PCR buffer II (Applied Biosystems), 0.5 µL of 0.2 mM deoxynucleotide triphosphates (Promega, USA), 0.5 µL of 20 mg mL−1 bovine serum albumin (Invitrogen Ultrapure, USA), 1 µL of 10 µM of each primer, 0.2 µL of AmpliTaq Gold DNA polymerase (Applied Biosystems) and 2 µL of template DNA. The amplification conditions consisted of a pre-denaturation step at 95 °C for 5 min, followed by 50 cycles of denaturation at 95 °C for 30 s, annealing at 57 °C for 30 s and extension at 72 °C for 1 min, followed by a final extension step at 72 °C for 5 min. PCR products, including negative control for each unique combination of tag-encoded primers, were verified by agarose gel electrophoresis. PCR products were purified using the High Pure PCR Cleanup Micro Kit (Roche) and quantified using a Qubit 2.0 fluorometer. Libraries were pooled in equimolar quantities and the sequence library was prepared using a TruSeq library-preparation kit (Illumina). Samples were then loaded into a MiSeq instrument for a paired-end run of 2 × 250 cycles. The sequencing was performed at the University of Geneva.
3.2.2 Data quality control and processing
The raw sequencing reads for each sample were processed using the SLIM web application (Dufresne et al., 2019). In brief, the module demultiplexer was used to demultiplex the raw reads according to their unique tags in the forward and reverse reads. Quality filtering, chimera removal and ASVs table generation were performed using DADA2 v.1.16 with pseudo-pool parameters (Callahan et al., 2015).
The ASVs were then curated using the LULU package v.0.1.0 (Froslev et al., 2017) with the default parameters. The taxa assignment of the ASVs was performed using VSEARCH against the taxonomically curated PR2 database v.4.14.1 (Guillou et al., 2012), which contains functional annotations. We used a Last Common Ancestor approach, assigning to the consensual taxonomic rank to up to reference sequences with at least 80 % similarity as a threshold for the dataset. The ASVs were also assigned to functional groups with at least 95 % similarity; with the functional attributes of Ibarbalz et al. (2019). The ASVs assigned to prokaryotes (bacteria and archaea) were removed in order to analyze only eukaryotic ASVs. Additionally, fungi and gymnamoebae were removed due to the high risk of contamination (Armbrecht, 2020). Unique ASVs (occurring in only one sample), short sequences (< 200 bp), rare ASVs (having < 100 reads), and low read count samples (< 1000 reads) were removed from the dataset. Additionally, the unassigned sequences were blasted with NCBI to further clarify the taxonomic composition. The Cumulative Sum Scaling (CSS) technique (scale factor – 0.9) was used to transform the read counts in the dataset and using `cssNorm' function in the genomeSeq package (Paulson et al., 2013), and used for the downstream statistical analysis in the study.
3.3 Statistical analysis
Data analysis was performed in R v.4.5.2 (Team, 2025) using several R packages. The relative abundances of reads and ASVs for each eukaryote group were calculated and plotted using ggplot2 (Wickham, 2016) and Grapher 24.1.213.
Three alpha diversity indices were calculated for all samples (ASV richness q=0, Shannon index q=1, and Simpson index q=2) based on the functions in the vegan package (Oksanen et al., 2025). Shannon diversity was compared between the main groups using the Kruskal–Walli's rank test in the stats package (Kruskal and Wallis, 1952). Significance between the groups was determined using a pairwise Wilcoxon rank sum test with an adjusted p value (Benjamini–Hochberg) in the ggpubr package (Kassambara, 2026). A Principal Coordinate Analysis (PCoA) ordination was generated using the Bray–Curtis dissimilarity matrix calculated using the ape package (Paradis and Schliep, 2019) and stringer package (Wickham, 2025) to assess beta diversity and visualize dissimilarities in eukaryotic community composition among the samples. Permutational Multivariate Analysis of Variance (PERMANOVA) with 999 permutations via the adonis2 function in the vegan package (Oksanen et al., 2025).
A co-occurrence heatmap representing most of the families in the study was generated. The corrplot and pheatmap packages (Kolde, 2025; Wei and Simko, 2024) were used to analyze the correlation between eukaryote families based on spearman method (a cutoff mark in correlation as > 0.5, and p value Benjamini–Hochberg (BH) adjusted < 0.05). Environmental parameters represented by palaeoceanographic proxies were used to identify the response of eukaryote species using three analytical methods. Seven proxies were used, including indicators of: (i) sea surface temperature (SST ) Łącka et al. (2019), (ii) AW (foraminifera Nonionellina labradorica and Buccella frigida Łącka et al. (2015), and dinocyst Operculodinium centrocarpum (Telesiński et al., 2024), (iii) ArW/meltwater (% C37:4 Łącka et al. (2019), (iv) glaciomarine condition (foraminifera Elphidium excavatum and Cassidulina reniforme Łącka et al. (2015), (v) sea ice (dinocyst Echinidinium karaense Telesiński et al. (2024), and (vi) bottom current dynamics (mean grain size 0–63 µm) Łącka et al. (2015). Multidimensional Fuzzy Set Ordination (MFSO) correlation plot, and Fuzzy set ordination (FSO) plots were generated for each environmental variable to assess their influence on eukaryote communities and identify key proxies for downstream analysis (Roberts, 2008). Firstly, a heatmap of sparse partial least squares (sPLS) regression between ASVs, and proxies was generated using the spls and cim function in mixOmics package (Froslev et al., 2017; Rohart et al., 2017). The potential ASV based indicators were selected based on a correlation coefficient threshold of > 0.3 and BH adjusted p value of < 0.05. Secondly, a Spearman correlation heatmap of the top 100 most significant ASVs (rho > 0.3 and p-adjust < 0.05) as generated using the pheatmap package. Finally, the dataset was analyzed using DEseq2 analysis, and the dataset was curated based on log2FoldC ≥ 1 (BH-adjusted p value < 0.05), and lower base mean (Love et al., 2014). Potential indicator ASVs were categorized based on their correlation strength and consistent detection across at least two methods or strong association with multiple paleo-proxies.
4.1 Metabarcoding data
A total of 2 620 808 raw sequence reads were generated from 55 samples. After initial quality filtering and denoising with DADA2 in SLIM, eight samples were excluded due to low number of reads (< 300). An additional five samples were removed during the downstream eukaryote-specific quality control process described in the Methods. This reduced our dataset to 1 609 500 sequence reads and 273 ASVs in 42 samples (Tables S2 and S3 in the Supplement).
4.2 Alpha diversity
Alpha diversity indices varied across time interval; however, Shannon diversity showed no significant differences among intervals (Kruskal–Wallis, p = 0.48; Fig. S2 in the Supplement, Table S2). This apparent discrepancy reflects that species richness is sensitive to rare taxa, whereas Shannon diversity accounts for both richness and evenness, making it less responsive to occasional low-abundance ASVs. Overall, the number of observed ASVs ranged from 4 to 144 (Fig. S2, Table S2). The highest values were observed in the Younger Dryas, the Early and Late Holocene, particularly at 12.3, 11.3, 9.5, 4.0, and 2.8–2.3 kyr BP. In contrast, a significant decrease in richness was observed around 13.3, 11.8, 2.17, and 1.8 kyr BP (Fig. S2, Table S2). Similar trends were revealed by both the Shannon and Simpson indices, with minimal diversity observed around 12.8, 11.7, 9.2 kyr, and 3.4 kyr BP. Due to limited data resolution, no clear trends in alpha diversity could be discerned between 9.2 kyr BP and 3.4 kyr BP. However, a decline was evident after 3.4 kyr BP, continuing towards 1.3 kyr BP (Fig. S2).
4.3 Beta Diversity
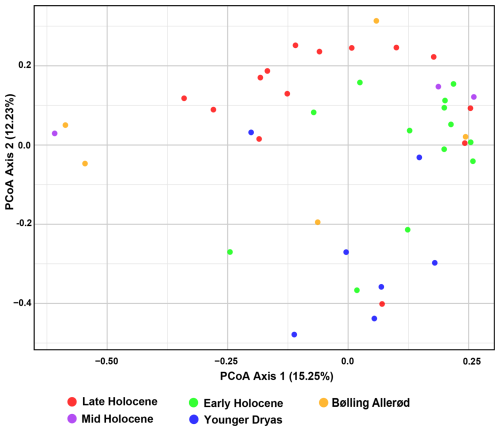
Beta diversity analyses revealed minor changes in community composition during the transitions from the Bølling–Allerød to the Younger Dryas, and from the Younger Dryas to the Holocene (Fig. 2). The PCoA plot revealed partial overlap across different time intervals, with Early Holocene and Younger Dryas samples were widely dispersed, while Late Holocene samples clustered separately along Axis 2, respectively. In contrast, Bølling–Allerød and Mid Holocene samples were largely scattered along the Axis 2 (Fig. 2). PERMANOVA results confirmed a significant effect of the time intervals on eukaryote community (R2 = 0.16, p < 0.05), with significant pairwise differences detected between the Late and Early Holocene, Late Holocene and Younger Dryas, Early Holocene and Younger Dryas, and Younger Dryas and Bølling–Allerød (Table S4 in the Supplement). FSO plots revealed a significant relationship between the samples and several paleo-environmental proxies, including the dinocyst Operculodinium centrocarpum from (Telesiński et al., 2024), ArW/meltwater indicator (% C37:4) (Łącka et al., 2019), and sea surface temperature (SST ) (Łącka et al., 2019) (Fig. S3 in the Supplement). In the MFSO framework, these proxies together explained a moderate proportion of the variation in community composition (cumulative fuzzy set correlation r = 0.454) (Fig. S3).
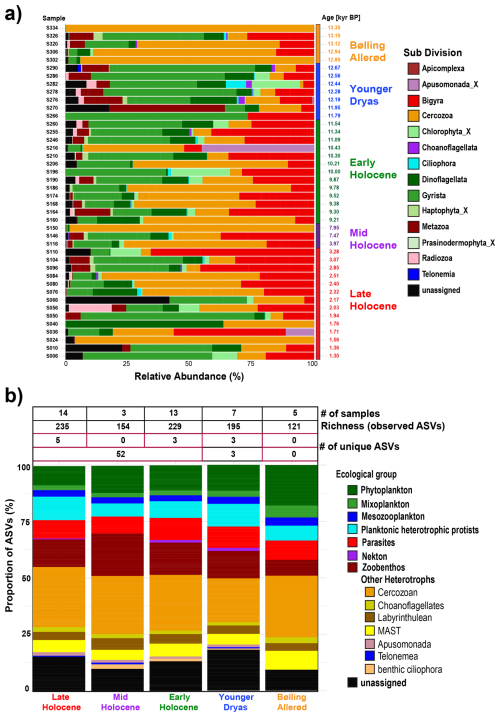
Figure 3(a) Bar plot showing the downcore distribution of eukaryotic sub-divisions based on their relative abundance. (b) Proportional richness of distinct ecological groups across selected time periods (Bølling–Allerød, Younger Dryas, Early Holocene, Mid Holocene, and Late Holocene), expressed as the percentage of ASVs. The accompanying table provides the number of samples, the total number of observed ASVs and unique ASVs within each climate time interval.
4.4 Community composition
Within the dataset, a total of 236 ASVs were assigned to 14 Sub-Divisions, while 37 ASVs remained unassigned (Table S3 in the Supplement). The Cercozoa was the most abundant sub-division, comprising 67 ASVs, accounting for 24.54 % of the total ASVs (Fig. 3, Table S3). Overall, the taxonomic structure of eukaryotes based on read abundance fluctuated significantly between samples (Fig. 3a). In contrast, ASV richness remained stable across different time periods except during the Bølling–Allerød period (Fig. 3b). The number of unique ASVs was highest during the Late Holocene, with five unique ASVs identified. The Younger Dryas and Early Holocene each exhibited three unique ASVs. Conversely, no distinctive ASVs were identified during the Mid Holocene and Bølling–Allerød periods. Across the entire Holocene, a total of 52 unique ASVs were recorded (Fig. 3b).
The ASVs were categorized based on their ecological roles, such as phytoplankton (31 ASVs), mixoplankton (7 ASVs), mesozooplankton (8 ASVs), planktonic heterotrophic protists (23 ASVs), parasites (24 ASVs), zoobenthos (36 ASVs), nekton (2 ASVs), and other heterotrophs (104 ASVs) (Fig. 3b, Table S3). The latter category comprises multiple taxonomic groups characterized by complex habitats and feeding behaviors, many of which have poorly understood ecological roles. This group includes Cercozoa (67 ASVs), Labyrinthulea (11 ASVs), Choanoflagellatea (5 ASVs), MAST (15 ASVs), benthic ciliophora (2 ASVs), Apusomonada (3 ASVs), and one ASV from the Telonemea flagellate group. The unassigned taxa (37 ASVs) also remained ecologically uncategorized (Fig. 3b).
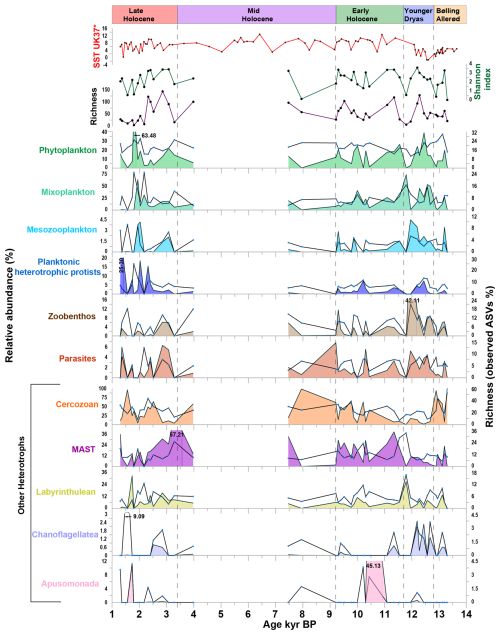
Figure 4Relative abundance and richness (expressed as observed ASVs percentage) of major ecological groups, along with Shannon index, richness, and sea surface temperature (SST UK'37) from Łącka et al. (2019). Lines represented ASVs abundance (%), and area represented the read abundance (%).
The phytoplankton community consisted of diatoms, green algae, haptophytes, and autotrophic dinoflagellates, most of which were associated with ArW (Fig. 4, Table S3). In terms of read abundance, Thalassiosira spp. and Chaetoceros sp. dominated among diatoms, while Micromonas polaris was the dominant species within the green algae. The haptophytes group was primarily represented by Phaeocytis sp., whereas the Gymnodinium spp. and sea-ice-associated species Polarella glacialis were dominant within the autotrophic dinoflagellate group (Fig. S4 in the Supplement). The mixoplankton community was primarily composed of mixotrophic dinoflagellates and silicoflagellates. In terms of read abundance, mixotrophic dinoflagellates were mainly present from the Younger Dryas to the beginning of the Early Holocene, whereas mixotrophic silicoflagellates, represented by Pseudopedinella sp., was present throughout the entire core (Fig. S4) (Sects. S1 and S2 in the Supplement).
The planktonic heterotrophic protists group included radiolarians, pelagic ciliates, dinoflagellates, and silicoflagellates. These groups were identified as being present at specific time periods, e.g. ∼ 12.4 to ∼ 10.2 kyr, and ∼ 2.3 to 1.3 kyr BP (Fig. S5 in the Supplement, Table S3). The mesozooplankton group comprises small metazoans and was dominated by arthropods (Copepoda and Malacostraca) and larvaceans (Appendicularia). The copepod Calanus spp. represented the majority of the mesozooplankton around the study area (Fig. S5) (Sects. S3 and S4 in the Supplement).
The zoobenthos was recorded as the most diverse group, primarily representing macrobenthic species (Fig. 4). This group included annelids, ascidiacean, molluscs, cnidarians, and echinoderms (Fig. S6 in the Supplement, Table S3). Zoobenthos taxa were most abundant around ∼ 9.3 kyr BP, as well as between ∼ 12.3 kyr BP (Fig. 4) (Sect. S5 in the Supplement).
The parasites were represented by 6 classes: Dinophyceae (Syndiniales), Gregarinomorphea, Paragregarea, Peronosporea, Hyphochytrea, and Enoplea (Fig. 4, Table S3). Among them, Syndiniales have the highest abundance and diversity, with 18 ASVs (mainly uncultured) detected throughout the studied time interval (Fig. S7 in the Supplement) (Sect. S6 in the Supplement). The nekton group included two ASVs assigned to Arctic cod (order Gnathostomata), which were detected only during the Younger Dryas and Early Holocene.
Among other heterotrophs, Cercozoa were dominant, accounting for a significant proportion of reads throughout the study period (Fig. 4, Table S3). Five classes of Cercozoa were identified: Ascetosporea, Phytomyxea, Granofilosea, Thecofilosea, and Imbricatea. Thecofilosea exhibited the highest richness with 51 ASVs (Fig. S8 in the Supplement, Table S3). Ecologically, cercozoans can be classified as parasitic, predatory, and bacterivorous. Cryothecomonas spp. were identified as predatory, while those in the classes Ascetosporea and Phytomyxea were classified as parasites. Other cercozoan ASVs were identified only to the family level, limiting precise ecological interpretations.
The other heterotrophs also include the MAST (Marine Stramenopiles), Labyrinthulea, Choanoflagellata and Apusomonada. The MAST group included 16 ASVs, representing four main sub-clades: MAST-1, MAST-3, MAST-9, and MAST-12. MAST-9 and MAST-12 dominated throughout the studied time period and exhibited high richness (Fig. S7, Table S3).
The Labyrinthulea included saprotrophic Thraustochytriaceae, and Aplanochytriidae, revealing a dominant presence and high richness around ∼ 11.8 to ∼ 2.2 kyr BP, and ∼ 1.7 kyr BP (Fig. 4). Most choanoflagellate ASVs belonged to environmental clades, except for Calliacantha sp., which was dominated around ∼ 12.7 kyr BP to ∼ 12.2 kyr BP (Fig. 4). Apusomonada, represented by the class Apusomonadea, appeared during certain time intervals, especially the Early Holocene (∼ 10.4 kyr BP), and Late Holocene (∼ 1.7 kyr BP) (Fig. 4, Table S3) (Sect. S7 in the Supplement).
4.5 Indicator taxa for Arctic and Atlantic water conditions
A total of 44 ASVs were identified as potential indicator taxa using three analytical approaches (sPLS, Spearman correlation, and DESeq) (Figs. S9 and S10 in the Supplement, Table S5 in the Supplement). Of these, 16 ASVs were identified as potential AW indicators, and belonged to the following groups: phytoplankton (1), planktonic heterotrophic protists (1), cercozoans (6), MAST (3), zoobenthos (1), labyrinthulean (1), and unassigned ASVs (1). The AW plankton indicators included a green algae Pyramimonas sp. (ASV55), heterotrophs-Silicoflagellate Pteridomonas sp. (ASV105), and a pelagic ciliate Cyclotrichium sp. (ASV265), while the AW benthic indicators comprised a polychaete Tharyx sp. (ASV278) (Fig. S11 in the Supplement, Table S5).
In contrast, 28 ASVs were associated with ArW (Table S5), primarily parasites (6), zoobenthos (3), phytoplankton (4), choanoflagellates (1), planktonic heterotrophic protists (3), mesozooplankton (1), mixoplankton (1), benthic ciliate (1), nekton (1), cercozoans (1), and unassigned ASVs (6) (Fig. S12 in the Supplement, Table S5). Potential ArW ASV-based indicators were identified among both planktonic and benthic taxa. Planktonic ASVs comprised autotrophic dinoflagellates (Prorocentrum sp., ASV64 and ASV153; Gymnodinium sp., ASV29), diatoms (Chaetoceros gelidus, ASV18), mixotrophic dinoflagellate (Heterocapsa rotundata, ASV204), pelagic ciliate (Strombidium sp., ASV123), radiolarian (Heteracon sp., ASV54; Acanthoplegma sp., ASV677), and mesozooplankton (Oikopleura sp., ASV500). Parasites included three Syndiniales dinoflagellates (ASV213, ASV341, and ASV391), an apicomplexan (Paralecudina sp., ASV958), and a parasitic nematode (Mermithidae sp., ASV101). Benthic indicators included benthic ciliate (Holosticha sp., ASV87), echinoderm (Ctenodiscus sp., ASV104), and bivalve (Tridonta sp., ASV137). Additionally, Calliacantha natans (ASV92) was identified as an ArW indicator in our study (Table S5).
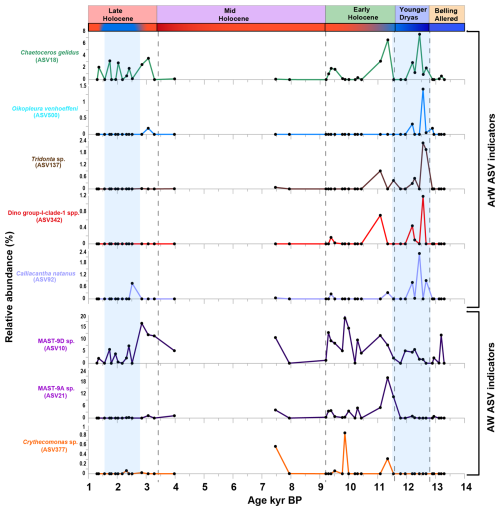
Figure 5Potential ASV-based indicators of AW and ArW conditions from the study (correlation > 0.4, p < 0.05, positive results from at least two method (displayed data as relative abundance %, and included the ASVs recorded in more than four samples).
Eight ASVs were identified as the most robust ASV-based indicators of AW and ArW conditions, based on their consistent occurrence through time (detected in more than four samples) and strong statistical support (correlation coefficient > 0.4; p < 0.05) (Fig. 5).
4.6 Ecological interactions among eukaryote families
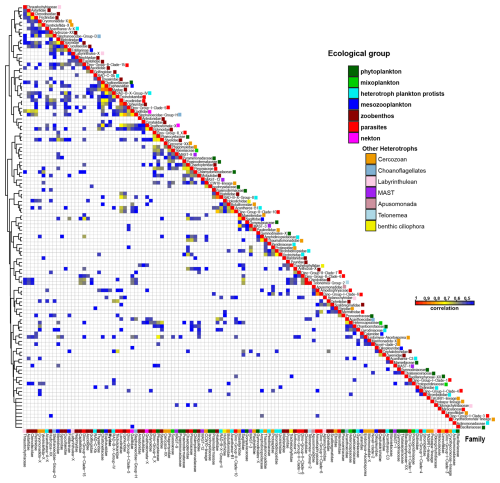
Spearman correlation analysis (r > 0.5, adjusted p < 0.001) was used to explore potential ecological interactions among eukaryotic families inhabiting similar environmental niches (Fig. 6, Table S6 in the Supplement). Parasitic cercozoans (Ascetosporea and Phagomyxidae) strongly correlated with algae families (Phaeocystaceae, Thalassiosiraceae, Pyramimonadaceae, and Prasinodermataceae), and dinoflagellates (Suessiaceae, and Gymnodiniaceae). Other cercozoans (CCW10-lineage, Novel-Clade-2, Cryothecomonas-lineage, and Ventricleftida) revealed significant correlations with algal and dinoflagellate groups (Fig. 6, Table S6). Among MAST groups, MAST-12 showed positive associations with algae families (Prasinodermataceae, Thalassiosiraceae, and Chaetocerotaceae) and parasites (Pirsoniaceae), while MAST-9 correlated with multiple phytoplankton families (Fig. 6, Table S6). Parasitic dinoflagellates (dino-group-II) showed strong correlation with haptophyte algae (Phaeocystaceae), diatom (Thalassiosiraceae), and dinoflagellates (Suessiaceae, and Gymnodiniaceae). Parasitic alveolates of the family Lecudinidae showed a strong positive correlation with the mesozooplankton (Malacostraca), radiolarians and ophiuroids (Fig. 6, Table S6). Another parasitic superfamily of Stramenopiles, the Pirsoniaceae displayed strong associations with various taxa, including MAST-12, cercozoans (Thaumatomonadidae), green algae (Prasinodermataceae, and Chlamydomonadales), haptophytes (Phaeocystaceae), diatoms (Thalassiosiraceae), silicoflagellates (Actinomonadaceae), dinoflagellates (Amphidiniopsidaceae), and polychaetae (Chaetopteridae) (Fig. 6, Table S6).
This study expands our knowledge of eukaryotic communities' patterns in the Storfjordrenna over the past 13 300 years by providing high-resolution sedaDNA records of both fossilized and non-fossilized groups. We demonstrate how these communities responded to major climatic shifts since the Bølling–Allerød and highlight key ecological interactions among major taxonomic groups. These findings enhance our understanding of how environmental changes have shaped eukaryotic biodiversity in southern Svalbard.
5.1 Impacts of oceanographic changes on the eukaryotic community in Storfjordrenna
5.1.1 Bølling–Allerød (13.30 to 12.80 kyr BP)
The eukaryotic community in our record during the Bølling–Allerød reflected oceanographic conditions resembling those observed today in glacier-proximal areas of Arctic fjords, characterized by high turbidity due to meltwater discharge and the presence of colder, fresher waters (Łącka et al., 2019; Zajączkowski, 2008). Previous studies support this interpretation: the grounding line of the Svalbard Barents Ice Sheet (SBIS) retreated from Storfjordrenna before 13.95 kyr BP (Łącka et al., 2015), coinciding with SST reaching modern-like values (Łącka et al., 2019). However, despite elevated SST, primary productivity remained low likely due to the suppressive effect of turbid meltwater input from the retreating ice sheet (Łącka et al., 2015). Furthermore, biomarker data indicated a dominance of fresher ArW over AW, which has been linked to reduced primary productivity (Łącka et al., 2019). These conditions favored the development of a eukaryotic community dominated by heterotrophs, capable of thriving in such extreme environments. The most abundant taxa, in terms of both sequence reads and ASV richness, were bacterivorous cercozoans (Fig. 4). The dominant cercozoan was Limnofila sp., a genus primarily found in fresh and brackish waters (Mylnikov et al., 2015; Nikolaev et al., 2003), the presence of Limnofila sp. may reflect either local ecological conditions or allochthonous input via riverine transport or ice-rafted debris (Andruszkiewicz et al., 2019; Jo et al., 2025; Nguyen et al., 2026). Other important bacterivorous heterotrophs were MAST, particularly MAST-9D and MAST-12A, which are known to be adapted to extreme environmental conditions (Labarre et al., 2021; Lin et al., 2022; Obiol et al., 2024).
Despite lower read abundance, phytoplankton, mixoplankton, and planktonic heterotrophic protists showed high ASV richness during this period (Fig. 4). The phytoplankton community was dominated by autotrophic, sea-ice associated taxa, such as dinoflagellates Polarella glacialis (Harðardóttir et al., 2024) and Gymnodinium spp. (Kubiszyn and Wiktor, 2016), and diatom Thalassiosira spp. (Luddington et al., 2016). The mixoplankton was represented by the silicoflagellate, Pseudopedinella elastica (Fig. S4), which has been described as bacterivorous under conditions of limited light and nutrients (Gerea et al., 2016). The mesozooplankton community was primarily composed of the herbivorous Calanus spp. and the omnivorous Metridia longa. These species have been previously observed in the Svalbard region, with Calanus spp. dominating in terms of biomass (Daase et al., 2008). Despite unfavorable conditions caused by meltwater influx and low nutrient availability, both primary and secondary productivity persisted, likely concentrated in ice-proximal and frontal zones where the stratification enhanced nutrient retention and water column stability (Łącka et al., 2019, 2015).This suggests that environments near retreating ice sheets can act as biological hotspots, supporting productivity through ice-associated blooms and complex interactions within the microbial food web.
5.1.2 Younger Dryas (12.80 to 11.70 kyr BP)
The eukaryotic community during the Younger Dryas reflected the dramatic environmental changes that occurred at that time. The most notable change was the rapid decrease in biodiversity during the Bølling–Allerød and Younger Dryas transition, when alpha diversity indices reached near-zero values (Fig. 4). The reorganization of oceanographic conditions most likely caused a temporary slowdown of Atlantic meridional overturning circulation (AMOC) and a reduction in AW inflow (Łącka et al., 2020). This led to strong stratification, formation of perennial ice cover, and anoxic conditions at the bottom (Łącka et al., 2020). The presence of perennial ice cover led to a significant reduction in primary productivity in Storfjordrenna (Łącka et al., 2019). However, the sedaDNA record also revealed the presence of phyto- and mixoplankton during this period, especially presence of phytoplankton Thalassiosira spp. and Gymnodinium sp., and silicoflagellate P. elastica. This may suggest that, although limited, primary productivity still occurred under the ice. The detection of herbivorous mesozooplankton Calanus spp., and predatory Cryothecomonas spp., also coincided with the presence of phytoplankton. Notably, the early Younger Dryas also revealed a short-term increase in the relative abundance and diversity of zoobenthos, primarily polychaetae (Barantolla sp.) and molluscs (Tridonta sp. and Talochlamys sp.) (Fig. S6).
During the latter part of the Younger Dryas (after ∼ 12.4 kyr BP), increasing advection of AW and SST warming led to the replacement of perennial sea ice by seasonal ice cover (Łącka et al., 2019, 2020). This shift was followed by the development of a more diverse benthic foraminifera community (Łącka et al., 2015). Similarly, the sedaDNA record displayed the increase in the richness and abundance of zoobenthic taxa, mainly annelids, molluscs, and echinoderms (Fig. S6). However, the alkenone record suggested that the warming was may associated with low primary productivity, probably due to the continuous input of turbid meltwater from the decaying SBIS (Łącka et al., 2015).
In contrast, the sedaDNA record indicated a sudden phytoplankton bloom in the late Younger Dryas, dominated by M. polaris, Thalassiosira spp., Chaetoceros gelidus, and Gymnodinium spp. Micromonas polaris is typically associated with Arctic sea-ice environments (Bachy et al., 2022), and was recorded during the periods of low SST. Chaetoceros gelidus is known for its high tolerance under variable light and ocean acidification conditions (Biswas, 2022; Ribeiro et al., 2024) and may play a key role in plankton blooms and primary productivity, particularly during the Younger Dryas (Fig. S4). Phytoplankton blooms stimulated the development of secondary producers, mainly pelagic ciliates, and radiolarians, as well as mesozooplankton copepods (Calanus spp.) (Fig. S5). Altogether, these findings indicate that the latter part of the Younger Dryas (after ∼ 12.4 kyr BP) was characterized by periods of accelerated AW inflow and higher SST (Risebrobakken et al., 2010; Wollenburg et al., 2004), which promoted phytoplankton and zooplankton growth, and enhanced development of the benthic community.
5.1.3 Early Holocene (11.70 to 9.20 kyr BP)
The transition from the Younger Dryas to the Early Holocene was characterized by a significant decrease in biodiversity and the dominance of mixoplankton, primarily silicoflagellate P. elastica, and dinoflagellates such as Biecheleria sp. and Gotoius sp. According to Łącka et al. (2020), the onset of the Early Holocene was associated with a short-term decrease in SST and a decrease in foraminiferal fauna abundance. The low biodiversity and dominance of mixotrophic plankton observed in the sedaDNA record might be a consequence of this short-term deterioration in environmental conditions.
However, the further development of the Early Holocene was driven by an increasing influence of AW in the area, which was followed by an increase in SST and productivity (Devendra et al., 2023; Telesiński et al., 2018). Moreover, Arctic Front was located close to the Spitsbergen coast, leading to the formation of a highly productive frontal zone (Łącka et al., 2019). The amelioration of environmental conditions during the Early Holocene (Łącka et al., 2015) was reflected in a sudden peak of alpha diversity of overall eukaryotic community, accompanied by a notable increase in the richness and abundance of key ecological groups, including phytoplankton, zoobenthos, parasites, and other heterotrophs such as cercozoans and MAST (Fig. 4). However, taxa associated with sea-ice were an important component of the assemblage, suggesting that sea-ice formation still occurred in Storfjordrenna. Despite AW dominance, the presence of a brackish water cercozoan Limnofila sp., green algae M. polaris and Pyramimonas sp., as well as sea-ice-indicator P. glacialis may suggest episodic presence of sea-ice, and transition of Arctic Front (Figs. S4 and S8). The overall high eukaryotic biodiversity in the Early Holocene, particularly the diversity of phytoplankton, mixoplankton, mesozooplankton, and the gradual increase in MAST-9 species related to warm water further support the establishment of warm-water conditions with high nutrient availability (Łącka et al., 2019) (Figs. S4, S5, S7, and Table S5).
5.1.4 Mid Holocene (9.20 to 3.40 kyr BP)
Samples from the Mid Holocene, spanning the period between 4.0 and 7.5 kyr BP, show low eukaryote DNA recovery, likely due to extensive degradation, and are dominated by fungi and amoebozoan DNA, particularly Acanthamoeba, which constitutes a large proportion of the recovered sequences. The dominance of amoebozoan DNA may reflect elevated microbial activity, potentially accelerating post-depositional DNA degradation (Anderson, 2017; Boere et al., 2011; Butler and Rogerson, 1995). As a result, lack of eukaryotic sedaDNA data from the period between ∼ 7.5 and 4.0 kyr BP (see Results), restricting interpretation to the early Mid Holocene and the transition toward the Late Holocene. Therefore, this interpretation should be treated with caution due to the limited number of samples analyzed in this interval. The beginning of the Mid Holocene (9.2 kyr BP) in Storfjordrenna was marked by a significant drop in biodiversity, followed by an increase after 8.0 kyr BP. The species composition was predominantly composed of cercozoans, mainly Limnofila sp. and Cryothecomonas spp. (Fig. S8). Another important component of the eukaryotic assemblage was MAST species, including MAST-9D and MAST-12B, which had previously been recorded in the north Atlantic region (Lopez-Garcia et al., 2007; Newbold et al., 2012) (Fig. S7). The community composition resembled the one from Bølling–Allerød, dominated by heterotrophic taxa adapted to unfavorable environmental conditions. This aligns with evidence of a minor cooling event between 9.0 kyr BP and 8.0 kyr BP, as proposed in previous studies (Łącka et al., 2015).
In contrast, despite their relatively low abundance, the communities of phytoplankton, mesozooplankton, and planktonic heterotrophic protists displayed relatively high diversity in the early Mid-Holocene. This period in Storfjordrenna was characterized by limited ice rafting, variable SST, and interplay between the AW and ArW water masses rather than a continuous impact of AW (Łącka et al., 2019, 2015). Furthermore, the low alkenone flux indicated low primary productivity throughout the mid-Holocene (Łącka et al., 2019), which is consistent with the reduced representation of both phyto- and zooplankton taxa in the sedaDNA record and supports an interpretation of sustained low productivity–related community signals during this interval. Low productivity was also observed at that time in the Norwegian and Svalbard shelves, potentially due to the limited nutrient availability. According to Łącka et al. (2019), the reduction in primary productivity resulted from enhanced vertical stratification, which reduced vertical mixing in the water column, and thus, limited the nutrient resuspension to the surface. An alternative explanation is the early spring bloom, that occurs in the ice-free waters, and the subsequent development of mesozooplankton that graze on phytoplankton, thereby reducing the flux of organic matter to the bottom. However, the presence of low abundance sequences assigned to both phytoplankton and planktonic heterotrophic protists, as well as the increase in bacterivorous taxa, likely supports the first scenario. Overall, the lack of sea ice, and the variability in water masses and SST observed at the beginning of the Mid Holocene, created an unstable environment, which favored tolerant heterotrophic eukaryotes such as cercozoans or MAST.
5.1.5 Late Holocene (3.40 to 1.30 kyr BP)
The onset of the Late Holocene was marked by an increase in eukaryotic biodiversity, followed by a sharp decrease around 2.0 kyr BP. During this period, eukaryotic communities were predominantly composed of cercozoan and MAST (Fig. 4). Cercozoan abundance and richness exhibited an increasing, yet variable trend throughout the Late Holocene, whereas MAST decreased progressively over time.
Both phytoplankton and planktonic heterotrophic protists exhibited high richness, but variable abundance throughout the Late Holocene (Fig. 4). Furthermore, the presence of parasitic species, including the Syndiniales dinoflagellate (dino-group-I and dino-group-II) as well as the diatom-associated parasitic Pirsonia sp. (Schweikert and Schnepf, 1997), co-occurred with the phytoplankton suggesting that parasitic interactions may have influenced phytoplankton dynamics during the Late Holocene. (Fig. S7, Table S6). The Late Holocene coincided with the so-called Neoglacial cooling, which spanned the last 4.0 kyr BP. This period was characterized by a decline in SST (Risebrobakken et al., 2010), limited AW inflow and strengthening of ArW flow, which led to the formation of extensive ice cover (Berben et al., 2014; Devendra et al., 2023; Martrat et al., 2003). Records from Storfjordrenna also showed a cooling in the area, associated with enhanced ice rafting (Łącka et al., 2019, 2015). Thus, the increased abundance of phytoplankton in general, and ice-associated species P. glacialis in particular, is probably an effect of the cooling of surface waters and the formation of sea-ice, which launched convective water mixing and nutrient resuspension to the surface. In consequence, primary productivity increased, stimulating the development of the planktonic heterotrophic protists community (Fig. S5).
5.2 sedaDNA environmental indicators
This study identified 46 potential eukaryotic indicator taxa associated with AW and ArW conditions. Several of these taxa exhibited consistent temporal patterns that aligned with paleoenvironmental proxies, supporting their potential for long-term reconstructions. In contrast, others appeared only sporadically, which reduces their reliability as potential indicators. AW-associated taxa were primarily represented by cercozoans and MAST, while ArW-associated taxa included diatoms, dinoflagellates, choanoflagellates, Arctic zoobenthos, and zooplankton.
Bacterivorous cercozoans, including the Ventricleftida (ASV46), the Protaspa-lineage (ASV83, and ASV257), Ventricleftida spp. (ASV46) and the Cryothecomonas spp. (ASV377), were identified as potential AW indicators (Fig. S11, Table S5). However, their identification is currently based exclusively on molecular data, limiting ecological and biogeographical context and weakening their use in environmental reconstructions (Labarre et al., 2021; Obiol et al., 2024). Similarly, members of the bacterivorous MAST-9 group, notably MAST-9A and MAST-9D, were exclusively detected in AW conditions (Fig. 5, Table S5), which is consistent with their known tropical-to-temperate distribution. Within the phytoplankton communities, Pyramimonas parkeae (a green alga that prefers higher temperature regions; Bock et al., 2021), and the planktonic heterotrophic pelagic ciliate Cyclotrichium sp. (commonly found in warmer waters; Dirmenci et al., 2010; Xu et al., 2005), were also identified as potential AW indicators. However, they only occurred in a brief temporal window near the end of the Early Holocene, so their reliability as indicators needs to be verified by further studies (Fig. S11).
The relatively high number of cold-water species recorded was probably due to favorable overall conditions in the study area. The autotrophic Prorocentrum spp. are known to be toxin-producing, bloom-forming species with broad global distributions, including polar regions (Cen et al., 2020; Goncharenko et al., 2021; Stoecker and Lavrentyev, 2018; Tillmann et al., 2022). In the present study, both taxa were primarily detected at the onset of the Younger Dryas (Table S6). Their limited distribution suggests that they may be unreliable as long-term environmental indicators. Similarly, Heterocapsa rotundata a mixotrophic dinoflagellate commonly associated with harmful algal blooms in Arctic and North Atlantic waters (Rintala et al., 2010; Wu et al., 2022), was identified as a potential ArW indicator (Fig. S12). The genus Holosticha is a widespread benthic ciliate, associated with sea ice (Berger, 2003; Petz et al., 1995; Wilbert and Song, 2008), and known to feed on diatoms and flagellates (Lei et al., 2005), which was also identified as a potential ArW indicator (Fig. S12). However, their presence was confined to the Bølling–Allerød and Younger Dryas or the Younger Dryas and Early Holocene intervals, respectively, limiting their reliability as a long-term proxy for ArW conditions (Fig. S12).
In contrast, the ArW-associated diatom species Chaetoceros gelidus was consistently abundant (Fig. 5), contributing to bloom formation and demonstrating adaptability to low light conditions (Biswas, 2022; Hoppe et al., 2018). Among the zooplankton, two taxa were identified as potential indicators: the planktonic heterotrophic radiolarian species Heteracon sp. and the mesozooplankton filter feeder appendicularian Oikopleura vanhoeffeni (Deibel, 1986, 1988) (Fig. 5). Likewise, the cold-water bivalve species Tridonta sp. which is commonly found in the North Atlantic and Arctic region (Marincovich Jr et al., 2002; Petersen, 2001), demonstrated strong potential as an indicator species. Within parasitic dinoflagellates, three potential indicators belonging to dino-group-I, mostly associated with sea-ice conditions (Clarke et al., 2019), were identified (Fig. S12). Choanoflagellate recorded in the study can be identified as a sea-ice associated group due to the presence of potential ArW indicator taxa (Buck and Garrison, 1988; Thomsen and Østergaard, 2017) (Fig. 5). These taxa were all consistently present throughout the study period, suggesting a stable association with cold marine conditions.
C. gelidus, O. vanhoeffeni, C. natans, Dino-group-I-clade-I spp., and Tridonta sp. exhibited the strongest ArW indicator potential, reflected by their consistent detection across samples and their concordant associations with multiple independent paleoenvironmental proxies. In contrast, MAST-9D sp., MAST-9A sp., and Cryothecomonas spp. showed the strongest indicators of AW influence (Fig. 5). However, their Spearman correlation coefficients between environmental variables ranged from 0.3 to 0.6 (p < 0.05), indicating a weak to moderate association. This may be due to a combination of interspecific competition and the influence of multiple external environmental variables in the study area. These interacting factors contribute to the complexity of the ecosystem and limit the effectiveness of using single-proxy approaches when interpreting the responses of indicator species in paleoenvironmental reconstructions. Further studies of eukaryotic communities in other Arctic regions are therefore needed to validate these taxa as robust indicator species.
5.3 Interactions within eukaryotic community structure in Storfjordrenna
The biodiversity of eukaryotic communities in Storfjordrenna was previously influenced by the interplay between ArW and AW masses, as well as sea-ice coverage over the past 13.30 kyr BP. Throughout the study period, eukaryotic alpha diversity remained relatively stable, with a notable exception during the transitions between major climatic intervals suggesting that these changes were driven primarily by species replacement, rather than by loss of richness or evenness (Figs. 4 and S2). This may imply that key trophic interactions and ecosystem functions persisted, reflecting functional resilience of Storfjordrenna eukaryotic communities during periods of environmental change. Notably, biodiversity peaks coincided with the presence of sea-ice margins and frontal zones, environments known to promote phytoplankton growth and primary productivity (Fig. 4).
Analysis of phytoplankton diversity revealed a consistent presence of green algae throughout the study period, except during the Bølling–Allerød interstadial, when diatoms dominated. Taxonomic abundance showed dynamic fluctuations, with gradual declines observed during the transitions between major climatic intervals. This suggests that environmental instability may have influenced the structure of the phytoplankton community.
Key contributors to primary productivity in the Storfjordrenna were diatoms (Thalassiosira spp., and Chaetoceros spp.), green algae (M. polaris), and autotrophic dinoflagellates (P. glacialis, and Gymnodinium spp.). Additionally, Spearman rank correlation analysis showed that the family Actinomonadaceae, mainly represented by Pseudopedinella sp., was positively associated with diatoms. Spearman rank correlation analysis also revealed a positive association between parasitic cercozoans and phytoplankton communities (Fig. 6, Table S6), indicating that the presence of parasitic cercozoans may play a significant role in shaping ecological interactions within phytoplankton assemblages (Bass et al., 2009; Cavalier-Smith and Chao, 2003; Hartikainen et al., 2014). Conversely, the parasitic apicomplexan family Lecudinidae was associated with zoobenthos (e.g., Heteroconchia and Ophiurida) and mesozooplankton (e.g., Malacostraca), may highlight their parasitic relationships with marine invertebrates (Rueckert et al., 2015) (Fig. 6, Table S6). Parasitic dinoflagellates (dino-group-II) were positively associated with haptophytes and diatoms. The parasitic nanoflagellate Pirsoniaceae: Pirsonia sp., demonstrated a positive correlation with autotrophic microbes, including, dinoflagellates, diatoms, and silicoflagellates (Kühn et al., 2004; Schweikert and Schnepf, 1997). This raises questions about the nature of their ecological interactions, and whether they are strictly parasitic or co-occur under similar environmental conditions. These findings emphasize the need for further investigation to understand the mechanisms driving these interactions and their broader implications for microbial community dynamics.
Cercozoans have emerged as the most dominant group within the eukaryotic community, in terms of both abundance and species richness. Cercozoan community included taxa previously recorded in various habitats including fresh and marine environments (Chantangsi and Leander, 2010; Irwin et al., 2019). Their occurrence across a wide range of environmental conditions highlights their ecological flexibility and broad tolerance. Although cercozoans as a group exhibit high richness, only a few lineages, such as the Imbricata-novel clade 2, Protaspa spp., Cryothecomonas spp., and Ascetosporea, persisted consistently throughout the study period, while most of the others were restricted to specific time intervals (Fig. S8). This suggests that, although cercozoans as a whole group may not be sensitive to environmental shifts, individual lineages are likely to be more responsive.
MAST species constituted a major microbial group within the eukaryotic community, with MAST-9 dominant overall and MAST-12 particularly prevalent during the late Holocene. These two MAST subgroups are commonly associated with temperate regions or extreme environments such as cold seeps (Lin et al., 2022; Obiol et al., 2024). The statistical analysis identified potential warm-water indicator species within these groups (Fig. 5, Table S5). Based on the co-occurrence relation, the MAST-12, and MAST-9 subgroup revealed a positive correlation (> 0.6) with the parasitic family of Pirsoniaceae and the phytoplankton families (Fig. 6, Table S6), providing insight into their ecological activities within the eukaryotic community. Further analysis of the ecological traits and distribution of micro eukaryotic taxa, mainly cercozoans and MAST, could provide deeper insights into their ecological responses within the Storfjordrenna ecosystem.
Using sedaDNA metabarcoding, we reconstructed the paleoecology of eukaryotic communities in Storfjordrenna over the last 13.30 kyr BP, elucidating their sensitivity and adaptability to environmental variables. Most eukaryotic ASVs were assigned ecological functional roles via higher taxonomic classification, considered reliable due to extensive ribosomal reference databases. However, taxa with complex life cycles (e.g., dinoflagellates, ciliates) required genus/species-level identification for more accurate functional assignment, necessitating a cautious approach. While some classification errors are inevitable, their proportion in the dataset is expected to be minimal. Overall, the eukaryotic biodiversity in Storfjordrenna remained relatively stable, except during transitions between major climatic intervals, indicating that community changes were primarily driven by species replacement rather than loss of richness or evenness, which maintained ecosystem function. Peaks of biodiversity coincided with the presence of sea-ice margins and frontal zones; environments known to foster favorable conditions for phytoplankton development. Cercozoans and MAST emerged as dominant groups, demonstrating their ecological flexibility and broad tolerance. This study revealed that primary productivity in the Storfjordrenna region was mainly driven by phytoplankton, including diatoms (Thalassiosira spp., Chaetoceros spp.), green algae (Micromonas spp.), and autotrophs dinoflagellates (P. glacialis) as well as mixoplankton species such as Pseudopedinella elastica. Our approach revealed that, despite significant species turnover, functional diversity and ecosystem functions remained largely stable, highlighting the resilience of Arctic planktonic communities. Several potential ASV-indicators were identified through multi-method analyses, including taxa associated with specific water masses. Our findings also underscore the complex interplay of environmental drivers shaping community composition, revealing both positive and negative associations among key microbial taxa. Our findings highlight the potential of sedaDNA for reconstructing past eukaryotic communities and detecting environmental change. However, to fully unlock the potential of sedaDNA approach in palaeoecological studies, it is essential to develop reference databases for accurate and precise taxonomic identification of sequences, and to provide modern reference data on the ecology and distribution of Arctic microbial eukaryotes. Therefore, improving taxonomic resolution and validating indicator taxa remain essential for establishing robust palaeoecological indicators in Storfjordrenna and the broader Svalbard region.
Raw sequencing reads generated in this study were deposited in the NCBI Short Read Archive (SRA) database under Bioproject accession no. PRJNA1299363, and the remaining data used for this study can be found in the Supplement.
The supplement related to this article is available online at https://doi.org/10.5194/bg-23-2525-2026-supplement.
HN and JoP, designed the study. JoP extract the DNA. HN analyzed the DNA data, and performed bioinformatic, statistical analyses, and interpret the results. JoP, JaP and N-LN helped with the bioinformatic analysis, and interpret the results. MŁ and DD help to clarify the age depth model, and the paleoenvironmental data. HN drafted the paper, and prepared the figures, and tables. All authors contributed to data interpretation and writing of the manuscript.
The contact author has declared that none of the authors has any competing interests.
Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims made in the text, published maps, institutional affiliations, or any other geographical representation in this paper. The authors bear the ultimate responsibility for providing appropriate place names. Views expressed in the text are those of the authors and do not necessarily reflect the views of the publisher.
We thank the captain and crew of R/V Jan Mayen, as well as the cruise participants, in particular Steinar Iversen, for their help at sea.
This research has been supported by the National Science Centre, Poland (grant no. 2022/47/B/ST10/03050).
This paper was edited by Mark Lever and reviewed by Connie Lovejoy and one anonymous referee.
Anderson, O. R.: Amoebozoan Lobose Amoebae (Tubulinea, Flabellinea, and Others), in: Handbook of the Protists, edited by: Archibald, J. M., Simpson, A. G. B., and Slamovits, C. H., Springer International Publishing, Cham, 1279–1309, https://doi.org/10.1007/978-3-319-28149-0_2, 2017.
Andruszkiewicz, E. A., Koseff, J. R., Fringer, O. B., Ouellette, N. T., Lowe, A. B., Edwards, C. A., and Boehm, A. B.: Modeling Environmental DNA Transport in the Coastal Ocean Using Lagrangian Particle Tracking, Front. Mar. Sci., 6, https://doi.org/10.3389/fmars.2019.00477, 2019.
Armbrecht, L.: The Potential of Sedimentary Ancient DNA to Reconstruct Past Ocean Ecosystems, Oceanography, 33, https://doi.org/10.5670/oceanog.2020.211, 2020.
Årthun, M., Eldevik, T., Smedsrud, L. H., Skagseth, Ø., and Ingvaldsen, R. B.: Quantifying the Influence of Atlantic Heat on Barents Sea Ice Variability and Retreat, J. Climate, 25, 4736–4743, https://doi.org/10.1175/JCLI-D-11-00466.1, 2012.
Bachy, C., Sudek, L., Choi, C. J., Eckmann, C. A., Nöthig, E. M., Metfies, K., and Worden, A. Z.: Phytoplankton Surveys in the Arctic Fram Strait Demonstrate the Tiny Eukaryotic Alga Micromonas and Other Picoprasinophytes Contribute to Deep Sea Export, Microorganisms, 10, 961, https://doi.org/10.3390/microorganisms10050961, 2022.
Bass, D., Chao, E. E. Y., Nikolaev, S. I., Yabuki, A., Ishida, K., Berney, C., Pakzad, U., Wylezich, C., and Cavalier-Smith, T.: Phylogeny of novel naked filose and reticulose Cercozoa: Granofilosea cl. n. and Proteomyxidea revised, Protist, 160, 75–109, https://doi.org/10.1016/j.protis.2008.07.002, 2009.
Benner, I., Irwin, A. J., and Finkel, Z. V.: Capacity of the common Arctic picoeukaryote Micromonas to adapt to a warming ocean, Limnol. Oceanogr. Lett., 5, 221–227, https://doi.org/10.1002/lol2.10133, 2019.
Bensi, M., Nilsen, F., Ferre, B., Skogseth, R., Moskalik, M., Korhonen, M., Vogedes, D., Kovacevic, V., de Mendoza, F. P., and Ingrosso, G.: The Atlantification process in Svalbard: a broad view from the SIOS Marine Infrastructure network (ARiS)8293871148, Svalbard Integrated Arctic Earth Observing System, 138–151, Zenodo, https://doi.org/10.5281/zenodo.14425672, 2024.
Berben, S. M. P., Husum, K., Cabedo-Sanz, P., and Belt, S. T.: Holocene sub-centennial evolution of Atlantic water inflow and sea ice distribution in the western Barents Sea, Clim. Past, 10, 181–198, https://doi.org/10.5194/cp-10-181-2014, 2014.
Berger, H.: Redefinition of Holosticha Wrzesniowski, 1877 (Ciliophora, Hypotricha), Eur. J. Protistol., 39, 373–379, https://doi.org/10.1078/0932-4739-00006, 2003.
Biswas, H.: A story of resilience: Arctic diatom Chaetoceros gelidus exhibited high physiological plasticity to changing CO2 and light levels, Front. Plant Sci., 13, 1028544, https://doi.org/10.3389/fpls.2022.1028544, 2022.
Blindheim, J. and Østerhus, S.: The Nordic Seas, Main Oceanographic Features, in: The Nordic Seas: An Integrated Perspective, edited by: Drange, H., Dokken, T., Furevik, T., Gerdes, R., and Berger, W., Geophysical Monograph Series, American Geophysical Union, 11–37, https://doi.org/10.1029/158GM03, 2005.
Bock, N. A., Charvet, S., Burns, J., Gyaltshen, Y., Rozenberg, A., Duhamel, S., and Kim, E.: Experimental identification and in silico prediction of bacterivory in green algae, ISME J., 15, 1987–2000, https://doi.org/10.1038/s41396-021-00899-w, 2021.
Boere, A. C., Sinninghe Damsté, J. S., Rijpstra, W. I. C., Volkman, J. K., and Coolen, M. J. L.: Source-specific variability in post-depositional DNA preservation with potential implications for DNA based paleoecological records, Org. Geochem., 42, 1216–1225, https://doi.org/10.1016/j.orggeochem.2011.08.005, 2011.
Buck, K. R. and Garrison, D. L.: Distribution and abundance of choanoflagellates (Acanthoecidae) across the ice-edge zone in the Weddell Sea, Antarctica, Mar. Biol., 98, 263–269, https://doi.org/10.1007/Bf00391204, 1988.
Butler, H. and Rogerson, A.: Temporal and Spatial Abundance of Naked Amoebae (Gymnamoebae) in Marine Benthic Sediments of the Clyde Sea Area, Scotland, J. Eukaryot. Microbiol., 42, 724–730, https://doi.org/10.1111/j.1550-7408.1995.tb01624.x, 1995.
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J., and Holmes, S. P.: DADA2: High resolution sample inference from amplicon data, Nat. Methods, 13, 581–583, https://doi.org/10.1038/nmeth.3869, 2015.
Cavalier-Smith, T. and Chao, E. E. Y.: Phylogeny and classification of phylum Cercozoa (Protozoa), Protist, 154, 341–358, https://doi.org/10.1078/143446103322454112, 2003.
Cen, J., Wang, J., Huang, L., Ding, G., Qi, Y., Cao, R., Cui, L., and Lü, S.: Who is the “murderer” of the bloom in coastal waters of Fujian, China, in 2019?, J. Oceanol. Limnol., 38, 722–732, https://doi.org/10.1007/s00343-019-9178-6, 2020.
Chantangsi, C. and Leander, B. S.: An SSU rDNA barcoding approach to the diversity of marine interstitial cercozoans, including descriptions of four novel genera and nine novel species, Int. J. Syst. Evol. Microbiol., 60, 1962–1977, https://doi.org/10.1099/ijs.0.013888-0, 2010.
Clarke, L. J., Bestley, S., Bissett, A., and Deagle, B. E.: A globally distributed Syndiniales parasite dominates the Southern Ocean micro-eukaryote community near the sea-ice edge, ISME J., 13, 734–737, https://doi.org/10.1038/s41396-018-0306-7, 2019.
Daase, M., Eiane, K., Aksnes, D. L., and Vogedes, D.: Vertical distribution of Calanus spp. and Metridia longa at four Arctic locations, Mar. Biol. Res., 4, 193–207, https://doi.org/10.1080/17451000801907948, 2008.
Deb, J. C. and Bailey, S. A.: Arctic marine ecosystems face increasing climate stress, Environ Rev., 31, 403–451, https://doi.org/10.1139/er-2022-0101, 2023.
Deibel, D.: Feeding mechanism and house of the appendicularian Oikopleura vanhoeffeni, Mar. Biol., 93, 429–436, https://doi.org/10.1007/Bf00401110, 1986.
Deibel, D.: Filter feeding by Oikopleura vanhoeffeni: grazing impact on suspended particles in cold ocean waters, Mar. Biol., 99, 177–186, https://doi.org/10.1007/Bf00391979, 1988.
Devendra, D., Łącka, M., Szymańska, N., Szymczak-Żyła, M., Krajewska, M., Weiner, A. K. M., De Schepper, S., Simon, M. H., and Zajączkowski, M.: The development of ocean currents and the response of the cryosphere on the Southwest Svalbard shelf over the Holocene, Global Planet. Change, 228, 104213, https://doi.org/10.1016/j.gloplacha.2023.104213, 2023.
Dirmenci, T., Dündar, E., Deniz, G., Arabaci, T., Martin, E., and Jamzad, Z.: Morphological, karyological and phylogenetic evaluation of Cyclotrichium: a piece in the tribe Mentheae puzzle, Turk. J. Bot., 34, 159–170, https://doi.org/10.3906/bot-0912-3, 2010.
Dufresne, Y., Lejzerowicz, F., Perret-Gentil, L. A., Pawlowski, J., and Cordier, T.: SLIM: a flexible web application for the reproducible processing of environmental DNA metabarcoding data, BMC Bioinform., 20, https://doi.org/10.1186/s12859-019-2663-2, 2019.
Esling, P., Lejzerowicz, F., and Pawlowski, J.: Accurate multiplexing and filtering for high-throughput amplicon-sequencing, Nucleic Acids Res., 43, 2513–2524, https://doi.org/10.1093/nar/gkv107, 2015.
Fonseca, V. G., Carvalho, G. R., Sung, W., Johnson, H. F., Power, D. M., Neill, S. P., Packer, M., Blaxter, M. L., Lambshead, P. J. D., Thomas, W. K., and Creer, S.: Second-generation environmental sequencing unmasks marine metazoan biodiversity, Nat. Commun., 1, 98, https://doi.org/10.1038/ncomms1095, 2010.
Froslev, T. G., Kjoller, R., Bruun, H. H., Ejrnaes, R., Brunbjerg, A. K., Pietroni, C., and Hansen, A. J.: Algorithm for post-clustering curation of DNA amplicon data yields reliable biodiversity estimates, Nat. Commun., 8, 1188, https://doi.org/10.1038/s41467-017-01312-x, 2017.
Gerea, M., Saad, J. F., Izaguirre, I., Queimaliños, C., Gasol, J. M., and Unrein, F.: Presence, abundance and bacterivory of the mixotrophic algae Pseudopedinella (Dictyochophyceae) in freshwater environments, Aquat. Microbe. Ecol., 76, 219–232, https://doi.org/10.3354/ame01780, 2016.
Goncharenko, I., Krakhmalnyi, M., Velikova, V., Ascencio, E., and Krakhmalnyi, A.: Ecological niche modeling of toxic dinoflagellate Prorocentrum cordatum in the Black Sea, Ecohydrol. Hydrobiol., 21, 747–759, https://doi.org/10.1016/j.ecohyd.2021.05.002, 2021.
Górska, B., Gromisz, S., Legeżyńska, J., Soltwedel, T., and Włodarska-Kowalczuk, M.: Macrobenthic diversity response to the atlantification of the Arctic Ocean (Fram Strait, 79° N) – A taxonomic and functional trait approach, Ecol. Indic., 144, 109464, https://doi.org/10.1016/j.ecolind.2022.109464, 2022.
Grant, D. M., Steinsland, K., Cordier, T., Ninnemann, U. S., Ijaz, U. Z., Dahle, H., De Schepper, S., and Ray, J. L.: Sedimentary ancient DNA sequences reveal marine ecosystem shifts and indicator taxa for glacial-interglacial sea ice conditions, Quaternary Sci. Rev., 339, 108619, https://doi.org/10.1016/j.quascirev.2024.108619, 2024.
Guillou, L., Bachar, D., Audic, S., Bass, D., Berney, C., Bittner, L., Boutte, C., Burgaud, G., de Vargas, C., Decelle, J., del Campo, J., Dolan, J. R., Dunthorn, M., Edvardsen, B., Holzmann, M., Kooistra, W. H. C. F., Lara, E., Le Bescot, N., Logares, R., Mahé, F., Massana, R., Montresor, M., Morard, R., Not, F., Pawlowski, J., Probert, I., Sauvadet, A. L., Siano, R., Stoeck, T., Vaulot, D., Zimmermann, P., and Christen, R.: The Protist Ribosomal Reference database (PR2): a catalog of unicellular eukaryote Small Sub-Unit rRNA sequences with curated taxonomy, Nucleic Acids Res., 41, D597-D604, https://doi.org/10.1093/nar/gks1160, 2012.
Hallegraeff, G. M.: Ocean Climate Change, Phytoplankton Community Responses, and Harmful Algal Blooms: A Formidable Predictive Challenge, J. Phycol., 46, 220–235, https://doi.org/10.1111/j.1529-8817.2010.00815.x, 2010.
Harðardóttir, S., Haile, J. S., Ray, J. L., Limoges, A., Van Nieuwenhove, N., Lalande, C., Grondin, P., Jackson, R., Skaar, K. S., Heikkilä, M., Berge, J., Lundholm, N., Massé, G., Rysgaard, S., Seidenkrantz, M., De Schepper, S., Lorenzen, E. D., Lovejoy, C., and Ribeiro, S.: Millennial-scale variations in Arctic sea ice are recorded in sedimentary ancient DNA of the microalga Polarella glacialis, Commun. Earth Environ., 5, https://doi.org/10.1038/s43247-023-01179-5, 2024.
Hartikainen, H., Ashford, O. S., Berney, C., Okamura, B., Feist, S. W., Baker-Austin, C., Stentiford, G. D., and Bass, D.: Lineage-specific molecular probing reveals novel diversity and ecological partitioning of haplosporidians, ISME J., 8, 177–186, https://doi.org/10.1038/ismej.2013.136, 2014.
Heaton, T. J., Köhler, P., Butzin, M., Bard, E., Reimer, R. W., Austin, W. E. N., Bronk Ramsey, C., Grootes, P. M., Hughen, K. A., Kromer, B., Reimer, P. J., Adkins, J., Burke, A., Cook, M. S., Olsen, J., and Skinner, L. C.: Marine20 – The Marine Radiocarbon Age Calibration Curve (0–55,000 cal BP), Radiocarbon, 62, 779–820, https://doi.org/10.1017/rdc.2020.68, 2020.
Hop, H., Wold, A., Vihtakari, M., Daase, M., Kwasniewski, S., Gluchowska, M., Lischka, S., Buchholz, F., and Falk-Petersen, S.: Zooplankton in Kongsfjorden (1996–2016) in Relation to Climate Change, in: The Ecosystem of Kongsfjorden, Svalbard, edited by: Hop, H. and Wiencke, C., Advances in Polar Ecology, vol. 2, Springer, Cham, 229–300, https://doi.org/10.1007/978-3-319-46425-1_7, 2019.
Hopkins, T. S.: The GIN Sea: A synthesis of its physical oceanography and literature review 1972–1985, Earth-Sci. Rev. 30, 175–318, https://doi.org/10.1016/0012-8252(91)90001-V, 1991.
Hoppe, C. J. M., Wolf, K. K. E., Schuback, N., Tortell, P. D., and Rost, B.: Compensation of ocean acidification effects in Arctic phytoplankton assemblages, Nat. Clim. Change, 8, 529–533, https://doi.org/10.1038/s41558-018-0142-9, 2018.
Ibarbalz, F. M., Henry, N., Brandao, M. C., Martini, S., Busseni, G., Byrne, H., Coelho, L. P., Endo, H., Gasol, J. M., Gregory, A. C., Mahe, F., Rigonato, J., Royo-Llonch, M., Salazar, G., Sanz-Saez, I., Scalco, E., Soviadan, D., Zayed, A. A., Zingone, A., Labadie, K., Ferland, J., Marec, C., Kandels, S., Picheral, M., Dimier, C., Poulain, J., Pisarev, S., Carmichael, M., Pesant, S., Tara Oceans, C., Babin, M., Boss, E., Iudicone, D., Jaillon, O., Acinas, S. G., Ogata, H., Pelletier, E., Stemmann, L., Sullivan, M. B., Sunagawa, S., Bopp, L., de Vargas, C., Karp-Boss, L., Wincker, P., Lombard, F., Bowler, C., and Zinger, L.: Global Trends in Marine Plankton Diversity across Kingdoms of Life, Cell, 179, 1084–1097.e1021, https://doi.org/10.1016/j.cell.2019.10.008, 2019.
IPCC: Climate Change 2022 – Impacts, Adaptation and Vulnerability: Working Group II Contribution to the Sixth Assessment Report of the Intergovernmental Panel on Climate Change, Cambridge University Press, Cambridge, https://doi.org/10.1017/9781009325844, 2023.
Irwin, N. A. T., Tikhonenkov, D. V., Hehenberger, E., Mylnikov, A. P., Burki, F., and Keeling, P. J.: Phylogenomics supports the monophyly of the Cercozoa, Mol. Phylogenet. Evol., 130, 416–423, https://doi.org/10.1016/j.ympev.2018.09.004, 2019.
Jo, T. S., Murakami, H., and Nakadai, R.: Spatial dispersal of environmental DNA particles in lentic and marine ecosystems: An overview and synthesis, Ecol. Indic., 174, 113469, https://doi.org/10.1016/j.ecolind.2025.113469, 2025.
Kassambara, A.: ggpubr: 'ggplot2' Based Publication Ready Plots, R package version 0.6.3, https://doi.org/10.32614/CRAN.package.ggpubr (last access: 10 March 2026), 2026.
Kolde, R.: pheatmap: Pretty Heatmaps, R package version 1.0.13, https://doi.org/10.32614/CRAN.package.pheatmap (last access: 10 March 2026), 2025.
Kruskal, W. H. and Wallis, W. A.: Use of ranks in one-criterion variance analysis, J. Am. Stat. Assoc., 47, 583–621, https://doi.org/10.1080/01621459.1952.10483441, 1952.
Kubiszyn, A. M. and Wiktor, J. M.: The Gymnodinium and Gyrodinium (Dinoflagellata: Gymnodiniaceae) of the West Spitsbergen waters (1999–2010): biodiversity and morphological description of unidentified species, Polar Biol., 39, 1739–1747, https://doi.org/10.1007/s00300-015-1764-2, 2016.
Kühn, S., Medlin, L., and Eller, G.: Phylogenetic position of the parasitoid nanoflagellate Pirsonia inferred from nuclear-encoded small subunit ribosomal DNA and a description of Pseudopirsonia n. gen. and Pseudopirsonia mucosa (Drebes) comb. nov, Protist, 155, 143–156, https://doi.org/10.1078/143446104774199556, 2004.
Labarre, A., Lopez-Escardo, D., Latorre, F., Leonard, G., Bucchini, F., Obiol, A., Cruaud, C., Sieracki, M. E., Jaillon, O., Wincker, P., Vandepoele, K., Logares, R., and Massana, R.: Comparative genomics reveals new functional insights in uncultured MAST species, ISME J., 15, 1767–1781, https://doi.org/10.1038/s41396-020-00885-8, 2021.
Łącka, M., Zajączkowski, M., Forwick, M., and Szczuciński, W.: Late Weichselian and Holocene palaeoceanography of Storfjordrenna, southern Svalbard, Clim. Past, 11, 587–603, https://doi.org/10.5194/cp-11-587-2015, 2015.
Łącka, M., Cao, M., Rosell-Melé, A., Pawłowska, J., Kucharska, M., Forwick, M., and Zajączkowski, M.: Postglacial paleoceanography of the western Barents Sea: Implications for alkenone-based sea surface temperatures and primary productivity, Quaternary Sci. Rev., 224, 105973, https://doi.org/10.1016/j.quascirev.2019.105973, 2019.
Łącka, M., Michalska, D., Pawlowska, J., Szymanska, N., Szczucinski, W., Forwick, M., and Zajaczkowski, M.: Multiproxy paleoceanographic study from the western Barents Sea reveals dramatic Younger Dryas onset followed by oscillatory warming trend, Sci. Rep., 10, 15667, https://doi.org/10.1038/s41598-020-72747-4, 2020.
Lei, Y., Xu, K., and Choi, J. K.: Holosticha hamulata n. sp. and Holosticha heterofoissneri Hu and Song, 2001, two urostylid ciliates (protozoa, ciliophora) from intertidal sediments of the yellow sea, J. Eukaryote. Microbiol., 52, 310–318, https://doi.org/10.1111/j.1550-7408.2005.00039.x, 2005.
Lejzerowicz, F., Esling, P., Majewski, W., Szczuciński, W., Decelle, J., Obadia, C., Arbizu, P. M., and Pawlowski, J.: Ancient DNA complements microfossil record in deep-sea subsurface sediments, Biol. Lett., 9, 20130283, https://doi.org/10.1098/rsbl.2013.0283, 2013.
Li, X., Li, F., Min, X., Xie, Y., and Zhang, Y.: Embracing eDNA and machine learning for taxonomy-free microorganisms biomonitoring to assess the river ecological status, Ecol. Indic., 155, 110948, https://doi.org/10.1016/j.ecolind.2023.110948, 2023.
Lin, Y. C., Chin, C. P., Yang, J. W., Chiang, K. P., Hsieh, C. H., Gong, G. C., Shih, C. Y., and Chen, S. Y.: How communities of marine stramenopiles varied with environmental and biological variables in the Subtropical Northwestern Pacific Ocean, Microbe. Ecol., 83, 916–928, https://doi.org/10.1007/s00248-021-01788-7, 2022.
Lindeque, P. K., Parry, H. E., Harmer, R. A., Somerfield, P. J., and Atkinson, A.: Next generation sequencing reveals the hidden diversity of zooplankton assemblages, PLOS One, 8, e81327, https://doi.org/10.1371/journal.pone.0081327, 2013.
Lopez-Garcia, P., Vereshchaka, A., and Moreira, D.: Eukaryotic diversity associated with carbonates and fluid-seawater interface in Lost City hydrothermal field, Environ. Microbiol., 9, 546–554, https://doi.org/10.1111/j.1462-2920.2006.01158.x, 2007.
Love, M. I., Huber, W., and Anders, S.: Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2, Genome Biol., 15, 550, https://doi.org/10.1186/s13059-014-0550-8, 2014.
Luddington, I. A., Lovejoy, C., and Kaczmarska, I.: Species-rich meta-communities of the diatom order Thalassiosirales in the Arctic and northern Atlantic Ocean, J. Plankton Res., 38, 781–797, https://doi.org/10.1093/plankt/fbw030, 2016.
Marincovich Jr, L., Barinov, K. B., and Oleinik, A. E.: The Astarte (Bivalvia: Astartidae) that document the earliest opening of Bering Strait, J. Paleontol., 76, 239–245, https://doi.org/10.1666/0022-3360(2002)076<0239:TABATD>.0.CO;2, 2002.
Martrat, B., Grimalt, J. O., Villanueva, J., van Kreveld, S., and Sarnthein, M.: Climatic dependence of the organic matter contributions in the north eastern Norwegian Sea over the last 15,000 years, Org. Geochem., 34, 1057–1070, https://doi.org/10.1016/s0146-6380(03)00084-6, 2003.
Mylnikov, A. P., Weber, F., Jurgens, K., and Wylezich, C.: Massisteria marina has a sister: Massisteria voersi sp. nov., a rare species isolated from coastal waters of the Baltic Sea, Eur. J. Protistol., 51, 299–310, https://doi.org/10.1016/j.ejop.2015.05.002, 2015.
Newbold, L. K., Oliver, A. E., Booth, T., Tiwari, B., DeSantis, T., Maguire, M., Andersen, G., van der Gast, C. J., and Whiteley, A. S.: The response of marine picoplankton to ocean acidification, Environ. Microbiol., 14, 2293–2307, https://doi.org/10.1111/j.1462-2920.2012.02762.x, 2012.
Nguyen, N. L., Pawłowska, J., Szymańska, N., Zajaczkowski, M., Weiner, A. K. M., De Schepper, S., and Pawłowski, J.: Assessing the passive dispersal of benthic foraminifera through environmental DNA, Limnol. Oceanogr., 71, e70294, https://doi.org/10.1002/lno.70294, 2026.
Nikolaev, S. I., Berney, C., Fahrni, J., Mylnikov, A. P., Aleshin, V. V., Petrov, N. B., and Pawlowski, J.: Gymnophrys cometa and Lecythium sp. are core Cercozoa: evolutionary implications, Acta Protozool., 42, 183–190, 2003.
Obiol, A., Del Campo, J., de Vargas, C., Mahe, F., and Massana, R.: How marine are Marine Stramenopiles (MAST)? A cross-system evaluation, FEMS Microbiol. Ecol., 100, https://doi.org/10.1093/femsec/fiae130, 2024.
Oksanen, J., Simpson, G., Blanchet, F., Kindt, R., Legendre, P., Minchin, P., O'Hara, R., Solymos, P., Stevens, M., Szoecs, E., Wagner, H., Barbour, M., Bedward, M., Bolker, B., Borcard, D., Borman, T., Carvalho, G., Chirico, M., De Caceres, M., Durand, S., Evangelista, H., FitzJohn, R., Friendly, M., Furneaux, B., Hannigan, G., Hill, M., Lahti, L., Martino, C., McGlinn, D., Ouellette, M., Ribeiro Cunha, E., Smith, T., Stier, A., Ter Braak, C., and Weedon, J.: vegan: Community Ecology Package, R package version 2.7-2, https://doi.org/10.32614/CRAN.package.vegan, 2025.
Paradis, E. and Schliep, K.: ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R, Bioinformatics, 35, 526–528, https://doi.org/10.1093/bioinformatics/bty633, 2019.
Paulson, J. N., Stine, O. C., Bravo, H. C., and Pop, M.: Differential abundance analysis for microbial marker-gene surveys, Nat. Methods, 10, 1200–1202, https://doi.org/10.1038/nmeth.2658, 2013.
Pawłowska, J., Lejzerowicz, F., Esling, P., Szczuciński, W., Zajączkowski, M., and Pawlowski, J.: Ancient DNA sheds new light on the Svalbard foraminiferal fossil record of the last millennium, Geobiology, 12, 277–288, https://doi.org/10.1111/gbi.12087, 2014.
Pawłowska, J., Łącka, M., Kucharska, M., Pawlowski, J., and Zajączkowski, M.: Multiproxy evidence of the Neoglacial expansion of Atlantic Water to eastern Svalbard, Clim. Past, 16, 487–501, https://doi.org/10.5194/cp-16-487-2020, 2020.
Perret-Gentil, L. A., Cordonier, A., Straub, F., Iseli, J., Esling, P., and Pawlowski, J.: Taxonomy-free molecular diatom index for high-throughput eDNA biomonitoring, Mol. Ecol. Resour., 17, 1231–1242, https://doi.org/10.1111/1755-0998.12668, 2017.
Perret-Gentil, L. A., Bouchez, A., Cordier, T., Cordonier, A., Guéguen, J., Rimet, F., Vasselon, V., and Pawlowski, J.: Monitoring the ecological status of rivers with diatom eDNA metabarcoding: A comparison of taxonomic markers and analytical approaches for the inference of a molecular diatom index, Mol. Ecol., 30, 2959–2968, https://doi.org/10.1111/mec.15646, 2021.
Petersen, G. H.: Studies on some Arctic and baltic Astarte species (Bivalvia, Mollusca), Museum Tusculanum Press, 2001.
Petz, W., Song, W., and Wilbert, N.: Taxonomy and ecology of the ciliate fauna (Protozoa, Ciliophora) in the endopagial and pelagial of the Weddell Sea, Antarctica, Land Oberösterreich, OÖ Landesmuseum, 1995.
Polyakov, I. V., Pnyushkov, A. V., Alkire, M. B., Ashik, I. M., Baumann, T. M., Carmack, E. C., Goszczko, I., Guthrie, J., Ivanov, V. V., Kanzow, T., Krishfield, R., Kwok, R., Sundfjord, A., Morison, J., Rember, R., and Yulin, A.: Greater role for Atlantic inflows on sea-ice loss in the Eurasian Basin of the Arctic Ocean, Science, 356, 285–291, https://doi.org/10.1126/science.aai8204, 2017.
Polyakov, I. V., Rippeth, T. P., Fer, I., Alkire, M. B., Baumann, T. M., Carmack, E. C., Ingvaldsen, R., Ivanov, V. V., Janout, M., Lind, S., Padman, L., Pnyushkov, A. V., and Rember, R.: Weakening of Cold Halocline Layer Exposes Sea Ice to Oceanic Heat in the Eastern Arctic Ocean, J. Climate, 33, 8107–8123, https://doi.org/10.1175/JCLI-D-19-0976.1, 2020.
Ribeiro, C. G., Lopes dos Santos, A., Trefault, N., Marie, D., Lovejoy, C., and Vaulot, D.: Arctic phytoplankton microdiversity across the marginal ice zone: Subspecies vulnerability to sea-ice loss, Elem. Sci. Anth., 12, https://doi.org/10.1525/elementa.2023.00109, 2024.
Rintala, J. M., Hällfors, H., Hällfors, S., Hällfors, G., Majaneva, M., and Blomster, J.: Heterocapsa Arctica Subsp. Frigida Subsp. Nov. (Peridiniales, Dinophyceae)-Description of a New Dinoflagellate and Its Occurrence in the Baltic Sea, J. Phycol., 46, 751–762, https://doi.org/10.1111/j.1529-8817.2010.00868.x, 2010.
Risebrobakken, B., Moros, M., Ivanova, E. V., Chistyakova, N., and Rosenberg, R.: Climate and oceanographic variability in the SW Barents Sea during the Holocene, Holocene, 20, 609–621, https://doi.org/10.1177/0959683609356586, 2010.
Roberts, D. W.: Statistical analysis of multidimensional fuzzy set ordinations, Ecology, 89, 1246–1260, https://doi.org/10.1890/07-0136.1, 2008.
Rohart, F., Gautier, B., Singh, A., and Lê Cao, K.: mixOmics: An R package for `omics feature selection and multiple data integration, PLOS Comput. Biol., 13, e1005752, https://doi.org/10.1371/journal.pcbi.1005752, 2017.
Rueckert, S., Wakeman, K. C., Jenke-Kodama, H., and Leander, B. S.: Molecular systematics of marine gregarine apicomplexans from Pacific tunicates, with descriptions of five novel species of Lankesteria, Int. J. Syst. Evol. Microbiol., 65, 2598–2614, https://doi.org/10.1099/ijs.0.000300, 2015.
Schweikert, M. and Schnepf, E.: Light and electron microscopical observations on Pirsonia punctigerae spec, nov., a nanoflagellate feeding on the marine centric diatom Thalassiosira punctigera, Eur. J. Protistol., 33, 168–177, https://doi.org/10.1016/S0932-4739(97)80033-8, 1997.
Sinniger, F., Pawlowski, J., Harii, S., Gooday, A. J., Yamamoto, H., Chevaldonné, P., Cedhagen, T., Carvalho, G., and Creer, S.: Worldwide Analysis of Sedimentary DNA Reveals Major Gaps in Taxonomic Knowledge of Deep-Sea Benthos, Front. Mar. Sci., 3, https://doi.org/10.3389/fmars.2016.00092, 2016.
Skogseth, R., Olivier, L., Nilsen, F., Falck, E., Fraser, N., Tverberg, V., Ledang, A. B., Vader, A., Jonassen, M. O., and Søreide, J.: Variability and decadal trends in the Isfjorden (Svalbard) ocean climate and circulation–An indicator for climate change in the European Arctic, Prog. Oceanogr., 187, 102394, https://doi.org/10.1016/j.pocean.2020.102394, 2020.
Stoecker, D. K. and Lavrentyev, P. J.: Mixotrophic Plankton in the Polar Seas: A Pan-Arctic Review, Front. Mar. Sci., 5, https://doi.org/10.3389/fmars.2018.00292, 2018.
Sundfjord, A., Albretsen, J., Kasajima, Y., Skogseth, R., Kohler, J., Nuth, C., Skarðhamar, J., Cottier, F., Nilsen, F., and Asplin, L.: Effects of glacier runoff and wind on surface layer dynamics and Atlantic Water exchange in Kongsfjorden, Svalbard; a model study, Estuar. Coast. Shelf S., 187, 260–272, https://doi.org/10.1016/j.ecss.2017.01.015, 2017.
R Core Team: R: A language and environment for statistical computing, Vienna, Austria, https://www.R-project.org/ (last access: 10 March 2026), 2025.
Telesiński, M., Kucharska, M., Łącka, M., and Zajączkowski, M.: A late response of the sea-ice cover to Neoglacial cooling in the western Barents Sea, Holocene, 34, 1088–1096, https://doi.org/10.1177/09596836241247305, 2024.
Telesiński, M. M., Przytarska, J. E., Sternal, B., Forwick, M., Szczuciński, W., Łącka, M., and Zajączkowski, M.: Palaeoceanographic evolution of the SW Svalbard shelf over the last 14 000 years, Boreas, 47, 410–422, https://doi.org/10.1111/bor.12282, 2018.
Thomsen, H. A. and Østergaard, J. B.: Acanthoecid choanoflagellates from the Atlantic Arctic Region – a baseline study, Heliyon, 3, https://doi.org/10.1016/j.heliyon.2017.e00345, 2017.
Tillmann, U., Wietkamp, S., Gottschling, M., and Hoppenrath, M.: Prorocentrum pervagatum sp. nov. (Prorocentrales, Dinophyceae): A new, small, planktonic species with a global distribution, Phycol. Res., 71, 56–71, https://doi.org/10.1111/pre.12502, 2022.
Wassmann, P., Duarte, C. M., AgustÍ, S., and Sejr, M. K.: Footprints of climate change in the Arctic marine ecosystem, Glob. Change Biol., 17, 1235–1249, https://doi.org/10.1111/j.1365-2486.2010.02311.x, 2010.
Wei, T. and Simko, V.: R package 'corrplot': Visualization of a Correlation Matrix, R package version 0.95, https://doi.org/10.32614/CRAN.package.corrplot, 2024.
Wickham, H.: ggplot2: Elegant Graphics for Data Analysis, Springer-Verlag New York, https://doi.org/10.1002/wics.147, 2016.
Wickham, H.: stringr: Simple, Consistent Wrappers for Common String Operations, Springer, 59–107, https://doi.org/10.32614/CRAN.package.stringr, 2025.
Wilbert, N. and Song, W.: A further study on littoral ciliates (Protozoa, Ciliophora) near King George Island, Antarctica, with description of a new genus and seven new species, J. Nat. Hist., 42, 979–1012, https://doi.org/10.1080/00222930701877540, 2008.
Wollenburg, J. E., Knies, J., and Mackensen, A.: High-resolution paleoproductivity fluctuations during the past 24 kyr as indicated by benthic foraminifera in the marginal Arctic Ocean, Palaeogeogr. Palaeocl., 204, 209–238, https://doi.org/10.1016/s0031-0182(03)00726-0, 2004.
Wu, X., Liu, Y., Weng, Y., Li, L., and Lin, S.: Isolation, identification and toxicity of three strains of Heterocapsa (Dinophyceae) in a harmful event in Fujian, China, Harmful Algae, 120, 102355, https://doi.org/10.1016/j.hal.2022.102355, 2022.
Xu, D., Song, W., and Hu, X.: Morphology of Cyclotrichium taniguchii sp. nov. and C. cyclokaryon with establishment of a new genus, Dicyclotrichium gen. nov.(Ciliophora: Haptorida), J. Mar. Biol. Assoc. UK, 85, 787–794, https://doi.org/10.1017/S0025315405011719, 2005.
Zajączkowski, M.: Sediment supply and fluxes in glacial and outwash fjords, Kongsfjorden and Adventfjorden, Svalbard, Pol. Polar Res., 29, 59–72, 2008.
Zimmermann, H. H., Stoof-Leichsenring, K. R., Kruse, S., Nürnberg, D., Tiedemann, R., and Herzschuh, U.: Sedimentary Ancient DNA From the Subarctic North Pacific: How Sea Ice, Salinity, and Insolation Dynamics Have Shaped Diatom Composition and Richness Over the Past 20,000 Years, Paleoceanogr. Paleoclimatol., 36, https://doi.org/10.1029/2020pa004091, 2021.
Zimmermann, H. H., Stoof-Leichsenring, K. R., Dinkel, V., Harms, L., Schulte, L., Hutt, M. T., Nurnberg, D., Tiedemann, R., and Herzschuh, U.: Marine ecosystem shifts with deglacial sea-ice loss inferred from ancient DNA shotgun sequencing, Nat. Commun., 14, 1650, https://doi.org/10.1038/s41467-023-36845-x, 2023.