the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Spatial heterogeneity of soil organic matter and microbial community composition across ice-wedge polygons and soil layers in Arctic lowland tundra
Victoria Martin
Cornelia Rottensteiner
Hannes Schmidt
Moritz Mohrlok
Julia Horak
Carolina Urbina-Malo
Julia Wagner
Willeke A'Campo
Luca Durstewitz
Niek Jesse Speetjens
Rachele Lodi
Bela Hausmann
Michael Fritz
Gustaf Hugelius
Andreas Richter
Arctic lowland tundra is characterized by pronounced spatial heterogeneity that introduces uncertainty into predictions of permafrost soil carbon dynamics. In these ecosystems, edaphic variability is primarily structured along two spatial axes: ice wedge polygon microtopography at the terrain scale and soil layers at the pedon scale. Here, we investigated how polygon types (low-, flat-, and high-centered polygons) and major soil layers (organic topsoil, mineral subsoil, cryoturbated material, and upper permafrost) jointly shape soil organic matter pools, microbial community composition, and potential extracellular enzyme activities.
Polygon-specific patterns in soil organic matter characteristics and microbial communities persisted across all soil layers, and soil-layer specific differences were consistent across polygon types, while interactive effects were comparatively minor. Low centered polygons showed reduced organic matter bioavailability, lower microbial abundances, and diminished hydrolytic enzyme potential compared to flat- and high-centered polygons. Organic topsoils emerged as pronounced microbial and enzymatic hotspots. The upper permafrost contained substantial amounts of relatively undecomposed organic matter and indicated a considerable potential for hydrolytic degradation upon thaw. Across both spatial axes, patterns in soil organic matter pools, and microbial communities were largely structured along gradients in organic matter inputs and redox conditions, which themselves arise from interactions in surface microtopography, hydrology, and vegetation.
Overall, our findings demonstrate that a limited number of spatial units captures a disproportionate share of edaphic, microbial, and biogeochemical variability in Arctic lowland tundra soils. Explicitly accounting for polygon morphologies and major soil layers therefore provides a tractable framework for upscaling soil processes across spatially heterogeneous ecosystems and improving climate-relevant biogeochemical projections.
- Article
(3876 KB) - Full-text XML
-
Supplement
(2307 KB) - BibTeX
- EndNote
Permafrost-affected landscapes are characterized by pronounced surface and sub-surface variability (Ping et al., 2015; Siewert et al., 2021). Over centennial to millennial timescales, periglacial processes have formed a dynamic mosaic of geomorphological landscape features in close spatial proximity (Washburn, 1956). Among the most widespread in continuous permafrost regions are ice-wedge polygons (French, 2007; Washburn, 1973), which cover approximately one third of the Arctic landmass and are particularly prevalent in ground-ice rich lowland tundra and thermokarst terrains of Siberia and North America (Brown, 1967; Fritz et al., 2016). These polygonal networks originate from cyclic freeze–thaw dynamics and repeated frost cracking that promote the formation of ice wedges within the ground (French, 2007; Washburn, 1973). Depending on the state of these ice wedges, distinct polygonal surface patterns eventually emerge through physical self-organization processes (Krantz, 1990; MacKay, 2000). When ice wedges grow, the plastic deformation of overlying soil layers results in elevated rims that enclose lower-lying areas, forming low-centered polygons (LCPs). Conversely, high-centered polygons (HCPs) arise when ice wedges degrade or when sediment or peat accumulation exceeds ice-wedge growth, resulting in raised centers surrounded by troughs. LCPs and HCPs thus exhibit inverse topographies (French, 2007; Washburn, 1973), whereas so-called flat-centered polygons (FCPs) represent an intergrade type with attributes such as a flat center bordered by shallow drainage channels (Shur et al., 2025; Vaughn and Torn, 2018).
In lowland tundra, polygon morphology constitutes a major axis of edaphic variability at the terrain scale. Their microtopographical characteristics influence soil hydrological and thermal dynamics, affect soil type and texture, shape the composition of microbial and vegetational communities, and impact soil biogeochemical processes and the ecosystem energy balance (Lara et al., 2018; Liljedahl et al., 2016; Nitzbon et al., 2019; Wainwright et al., 2015). HCPs typically exhibit well-drained centers with dry surface conditions, whereas the centers of LCPs regularly experience inundation and ponding (Boike et al., 2008; Nitzbon et al., 2019). Waterlogging strongly shapes soil conditions in LCPs. Low oxygen availability restricts microbial decomposition and facilitates organic matter accumulation (Donner et al., 2012; Kuhry et al., 2020), leading to the development of a prominent organic layer (Organic Cryosoils). Because wetter soils have higher thermal conductivity, summer active layer depths may reach deeper in LCP centers compared to HCP centers (Liljedahl et al., 2016; Speetjens et al., 2022; Walvoord and Kurylyk, 2016). Yet, soils in LCPs are thought to experience the least pronounced seasonal temperature fluctuations among polygon types (Hubbard et al., 2013), owing to the combined insulating effects of summer inundation, preferential snow accumulation during winter, and peat buildup (Abolt et al., 2018, .Grosse et al., 2011). In the less insulated FCPs and HCPs, frost penetrates deeper into the ground, which promotes the mixing of soil layers (Turbic Cryosoils) and leads to the burial of poorly decomposed organic material from the topsoil into the mineral subsoil via cryoturbation (Ping et al., 2008; Wild et al., 2016).
Differences in soil properties across polygon types translate into characteristic shifts in plant and microbial community structure and function (Chu et al., 2011; Taş et al., 2018; Wolter et al., 2016). Dry surfaces of HCPs are typically dominated by dwarf-shrubs, forbs, and lichens (Speetjens et al., 2022; Wainwright et al., 2015), whereas vegetation in LCPs is adapted to water-saturated conditions and consists largely of graminoids and peat- or brown-mosses (Minayeva et al., 2018). Gradients in soil redox conditions associated with surface hydrology likewise influence the composition of microbial communities and the dominant pathways of soil organic matter (SOM) transformation (Ernakovich et al., 2017; Liebner et al., 2008; Lipson et al., 2015). Aerobic communities and processes prevail in FCPs and HCPs, whereas anaerobic pathways are common in LCPs (Frank-Fahle et al., 2014; Roy Chowdhury et al., 2021). Because soil physicochemistry, vegetation-derived SOM inputs, and microbial communities jointly regulate biogeochemical cycling (Joabsson and Christensen, 2001; Sachs et al., 2010; Taş et al., 2018; Wallenstein et al., 2007), polygon morphology explains a substantial portion of the spatial variability in lowland tundra carbon exchange (Arora et al., 2019; Wainwright et al., 2015). For example, LCPs are recognized as significant sources of CH4, whereas CO2 efflux dominates from HCPs (Lara et al., 2015; Sachs et al., 2010).
In permafrost affected systems, another major axis of edaphic variability emerges at the pedon scale (Siewert et al., 2021). Key physicochemical properties such as temperature, redox conditions, ice- and organic matter content, or bulk density show strong vertical stratification along the soil profile. Most notably, the seasonal thaw of the active layer contrasts sharply with the persistently frozen permafrost below. In ice-rich, poorly drained lowland tundra soils, cryogenic processes are especially common. Over time, frost heave and cryoturbation introduce fine-scale irregularities in the vertical arrangement of soil horizons, disrupting the prevailing physicochemical and biotic conditions.
Naturally, these gradients and dynamics carry important implications for the life of associated microbial communities. For instance, the permafrost table imposes a strong physical barrier to the exchange of water, nutrients, and gasses between the active layer and the frozen soil (Wilhelm et al., 2011), but also restricts microbial dispersion (Bottos et al., 2018; Doherty et al., 2020; Ernakovich et al., 2022).The position of the permafrost table is, however, temporally variable, and under sustained summer warming the uppermost portion of frozen permafrost (transient layer) may thaw on decadal timescales (Shur et al., 2005). In tundra, topsoil horizons can experience pronounced seasonal and diurnal temperature fluctuations, while deeper horizons remain thermally more stable (Baker et al., 2023; Barbier et al., 2012). Within the active layer, the most marked changes in texture, bulk density, organic matter content, and soil water capacity occur at the transition between the organic topsoil and the mineral subsoil (Alexander, 1989; Huntington, 2020). The influence of plants also decreases rapidly with depth. Approximately 96 % of the root biomass in tundra is concentrated within the upper 30 cm of the soil profile (Iversen et al., 2015; Jackson et al., 1996). Only a few species, e.g., sedges, possess deeper rooting systems that affect deeper layers via oxygenation, exudation, or litter inputs (Joabsson and Christensen, 2001; Shaver et al., 1979). As contributions of acidifying plant-derived inputs such as peat moss biomass and metabolites (Clymo and Hayward, 1982; Jones, 1998; Vives-Peris et al., 2020), or root-derived organic acids diminish with depth, soil pH tends to increase, accordingly. Owing to these depth-dependent patterns, pronounced gradients in the quantity, quality, and stoichiometry of soil organic matter emerge throughout the soil profile (Weintraub and Schimel, 2003).
Along these two axes of spatial variability a wide range of microbial habitats and ecological niches occurs across comparatively small spatial distances (e.g., Frank-Fahle et al., 2014; Gittel et al., 2014; Malard and Pearce, 2018; Taş et al., 2018). This mosaic of dynamic and contrasting environmental conditions imposes physiological constraints on microbial communities and selects for diverse metabolic strategies and specialized adaptations (Jansson and Taş, 2014; Lipson et al., 2013; Tveit et al., 2013; Waldrop et al., 2025). Frozen permafrost, represents a harsh environment for microorganisms, characterized by sub-zero temperatures, limited water availability and constrained nutrient resources (Ernakovich et al., 2022; Mackelprang et al., 2017). Under these conditions, microbial activity is largely confined to thin films and channels of unfrozen saline water (Gilichinsky et al., 2003). In contrast, active layer horizons, particularly organic topsoils, support substantially higher microbial biomass and diversity (Doherty et al., 2025; Waldrop et al., 2025), due to less harsh conditions, including greater substrate availability, oxygenation, and hydrological connectivity. The functional and spatial turnover in microbial communities thus links edaphic heterogeneity to microbial metabolic potential and, ultimately, to variability in soil carbon transformations. The production of greenhouse gases through microbial processing of previously frozen organic matter (Graham et al., 2012; Knoblauch et al., 2018; Kwon et al., 2019; Mackelprang et al., 2011; Xue et al., 2016) may shift tundra ecosystems from a net carbon sink to a source (IPCC, 2022; Schuur et al., 2008, 2015; Voigt et al., 2016). This coupling to the permafrost carbon–climate feedback has made permafrost microbial ecology a central focus of research over recent decades (e.g., Ernakovich et al., 2022; Hultman et al., 2015; Jansson and Taş, 2014; Johnston et al., 2019; Keuper et al., 2020; Waldrop et al., 2023, 2025), yet ecosystem-scale biogeochemical models still inadequately capture the effects of edaphic variability (Sturtevant and Oechel, 2013).
We therefore build on a conceptual framework in which different polygon morphologies and soil layers represent the two primary axes of edaphic variability in ice-wedge polygon tundra. Polygon microtopography generates lateral gradients in soil redox conditions, soil types, and organic matter inputs which are associated with distinct hydrologic and vegetation patterns. Vertical gradients along the soil profile impose additional physicochemical controls through shifts in temperature regimes, oxygen availability, pH, and plant inputs. Together, these two spatial dimensions structure organic matter quantity and quality, influence microbial abundance, diversity, and community composition, constrain dominant pathways of organic matter transformation, and shape climate-relevant biogeochemical dynamics. An integrative characterization of polygonal lowland tundra soils across both scales is therefore essential for understanding the functioning of these landscapes and for assessing their future trajectories under climate change. Although effects of polygon morphology on edaphic properties, vegetation, and trace gas exchange have been well described (e.g., Lara et al., 2015; Liljedahl et al., 2016; Sachs et al., 2010; Wainwright et al., 2015), and depth-dependent controls on organic matter composition and microbial communities are also well documented (e.g., (Kuhry et al., 2020; Lynch et al., 2023; Müller et al., 2018; Schnecker et al., 2015; Wild et al., 2016), only few studies have considered both spatial dimensions together (Lipson et al., 2015; Taş et al., 2018).
To address this gap, we characterized physicochemical properties, organic matter composition, bacterial, archaeal, and fungal community structure, and potential enzyme activities across both axes of edaphic variability in ice-wedge polygon tundra. Specifically, we tested whether (i) polygon morphology (low-, flat-, and high-centered polygons) and soil layers (organic topsoil, mineral subsoil, cryoturbated material, upper permafrost) exert predominantly independent main effects, or interacting controls, and whether (ii) consistent patterns emerge across these scales that may inform scalable representations of Arctic lowland tundra heterogeneity in future ecosystem and land-surface models.
2.1 Study area
We studied Arctic lowland ice-wedge polygon tundra, located on the coastal plain of the Yukon, Western Canada, (Fig. 1). The first focus area comprised two small lagoons called Ptarmigan Bay (69°27′ N, 139°05′ W) and Whale Bay (69°25′ N, 138°59′ W). The second focus area, approximately 40 km further towards the west called Komakuk Beach (69°35′ N, 140°10′ W), is a small coastal catchment positioned between two alluvial fans. The periglacial landscape in this ecosystem is characterized by a mosaic of ice-wedge polygon networks, mires, beaded streams, and thermokarst lakes (Fritz et al., 2012; Rampton, 1982; Speetjens et al., 2022), underlain by continuous permafrost with a high ground ice content (Couture and Pollard, 2017; Westerveld et al., 2023). The climate is classified as Polar Tundra (Beck et al., 2018), and the vegetation as bioclimatic subzone E/low Arctic shrub tundra (Walker et al., 2005). Microtopography and relief are strong determinants for the identity of the prevailing soil suborder, and plant species composition. Turbic Cryosols were present in the drier centers of HCPs and FCPs (Soil Classification Working Group, 1998), where also dwarf-shrubs, forbs, and lichens dominated the flora (Table S1a). FCPs were mainly characterized by graminoid tussocks and dwarf-shrubs. Inundated centers of LCPs harbored organic Cryosols. The dominant plant groups were graminoids, brown mosses, and peat mosses (Brooks and Lane, 2011; Rampton, 1982; Walker et al., 2005). A more detailed description of the study area, their surface geology, glaciation history, climate, soil suborders and vegetation, can be found in the Supplement Sect. S1, and in Wagner et al. (2023).
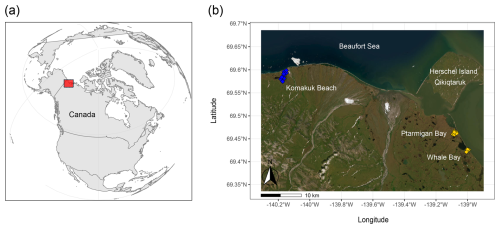
Figure 1Study area. Panel (a) marks the study area along the Yukon Coast, western Canadian Arctic. Panel (b) provides an aerial overview of the sampling locations at Ptarmigan Bay and Whale Bay (yellow, sampled in 2018) and at Komakuk Beach (blue, sampled in 2019). To capture edaphic variability in lowland tundra ecosystems, sampling was conducted across the main ice-wedge polygon types (low-, flat-, and high-centered polygons) and across major soil layers (organic topsoil, mineral subsoil, cryoturbated material, and upper permafrost). Maps were created using code available on GitHub (Irwin, 2021). Basemap: Esri | Powered by Esri.
2.2 Soil sampling and sample storage
Soil sampling was conducted during two field campaigns. Ptarmigan Bay and Whale Bay were sampled in August 2018, and Komakuk Beach in August 2019. Sampling took place in late summer, when active layer depths typically approach their seasonal maximum. In the field, we identified larger networks of low-centered (LCPs), flat-centered (FCPs), and high-centered polygons (HCPs) and selected six polygons of each type for sampling (Table S1 a). For sampling the active layer in HCPs and FCPs, we excavated 1–2.5 m wide soil pits until the permafrost table was reached. We recorded active layer depths, in situ soil temperatures at a distance of 10 cm (Table S1b), classified soil horizons following Schoeneberger et al. (2012), and documented their distribution and thickness (Fig. S1). We collected 100–200 g of fresh material from each horizon by compositing subsamples from several positions within the soil profile. Organic horizons and cryoturbated material were sampled by cutting blocks of known dimensions using a knife. For mineral subsoils we inserted steel cylinders (5.5 cm diameter) horizontally into the exposed profile. Excavating soil pits was not possible for the mostly waterlogged LCPs. We therefore restricted active layer sampling to retrieving two replicate cores per plot using a gas-powered SIPRE corer (diameter 7.5 cm). Documentation, identification and sampling of soil horizons were done in the same manner as described for HCPs and FCPs. In all types of polygons, we sampled the frozen part of the permafrost by using a gas powered SIPRE corer, or by hammering a steel pipe (diameter 4.2 cm) into the ground with a sledgehammer (Hugelius et al., 2010). For each core, we recorded the identity and dimensions of respective horizons and estimated visible ice contents. As within this study only the upper 10 cm of the extracted permafrost cores were used, we strictly refer to the transient layer when discussing characteristics of the permafrost layer.
Soil sampling and sample processing followed contamination-minimizing procedures feasible under field conditions: nitrile gloves were worn throughout and changed between samples, tools (e.g., knives, bulk-density cylinders, SIPRE corer) were cleaned with water and sterilized using alcohol-based disinfectant wipes between samples. Permafrost cores were handled on cutting boards lined with fresh aluminum foil, and the outer rind of each core was removed with a sterilized knife before subsampling. Within 24 h after sampling, we carefully removed visible roots, green litter, and coarse solid organic matter fragments from active layer samples and homogenized them by hand. Sample aliquots for DNA extraction were preserved using RNAlater™ Stabilization Solution (ThermoFisher Scientific). Active layer samples were stored and transported at 4 °C, and permafrost samples frozen. The samples arrived approximately two weeks after each respective sampling campaign at the University of Vienna and were processed immediately. Prior analysis, frozen permafrost samples were thawed for two days at 4 °C and homogenized inside the sterile sampling bags. The samples of both field campaigns were treated with the same protocols, analyzed by the same methods and combined into one dataset. For more details see Supplement Sect. S1.
In total, 81 soil samples were collected (Ptarmigan Bay and Whale Bay n=39; Komakuk Beach n=42). Samples were grouped by polygon type (LCP_n=20, FCP_n=32; HCP_n=29), and by soil layer category (organic topsoil_n=35 including O, Oi, Oe, Oa horizons; mineral subsoil_n=14 including B, Bg horizons, cryoturbated material_n=13 including Ojj, Oijj, Oajj, Ajj horizons, and upper permafrost_n=19 including Off, Bff, Cff horizons). Due to natural heterogeneity in the field and soil-pit specific differences in soil horizon development, an imbalanced sampling design emerged (organic: LCP_n=12, FCP_n=12, HCP_n=11; mineral: LCP_n=2, FCP_n=6, HCP_n=6; cryoturbated: LCP_n=0, FCP_n=7, HCP_n=6; permafrost LCP_n=6, FCP_n=7, HCP_n=6; see also Table S1c).
2.3 Physicochemical soil parameters and nutrient pools
The samples were analyzed for bulk density, pH (ultra-clean water), and gravimetric water content (80 °C for 48 h). We measured total soil Carbon (Soil C), Nitrogen (Soil N), plus their isotopic composition by an elemental analyzer (EA 1110, CE Instruments, Italy) coupled to a continuous-flow isotope ratio mass spectrometer (IRMS, DeltaPlus, Finnigan MAT). Following a modified ignition method (Kuo, 1996) to convert organic phosphorous (P) to inorganic P, soil total P (Soil P) was determined photometrically in 0.5 M H2SO4 extracts via malachite-green-assay (D'Angelo and Crutchfield, 2001). Dissolved organic carbon (DOC) and total dissolved nitrogen (TDN) concentrations were quantified in 1 M KCl extracts via TOC/TN-Analyzer (Shimadzu, TOC-VCPH/CPNTNM-1 analyzer). For more details see Supplement Sect. S2, and https://doi.org/10.5281/zenodo.18631833 (Martin, 2026).
2.4 Chemical composition of soil organic matter
The chemical composition of soil organic matter (SOM) was characterized by Pyrolysis-Gas Chromatography/Mass Spectrometry (CDS Pyroprobe 6200, CDS Analytical coupled to Pegasus BT, LECO; with the polar column Supelcowax™ 10 Fused Silica Capillary Column, 30 m × 0.25 mm × 0.25 µm film thickness, Sigma Aldrich), using the semi-automated approach that is described in Martin et al. (2024) with minor modifications. For the qualitative investigation of the SOM pool, we performed Principal Component Analysis (PCA) on center-log-ratio (clr) – transformed abundances (mg C per g soil DW) of 534 pyrolysis products. We further grouped these pyrolysis products into six SOM compound groups (aromatics and phenols, carbohydrates, lignins and lignin-derived compounds, lipids, N-containing substances, compounds of general and unknown origin and explored differences in their absolute and relative abundances among polygon types and soil layer categories. For more details see Supplement Sect. S3, and https://doi.org/10.5281/zenodo.18631833 (Martin, 2026).
2.5 Soil microbial communities – DNA extraction, amplicon sequencing, digital droplet (dd)PCR
In this study “microbial communities” refer to bacterial, archaeal, and fungal taxa profiled by amplicon sequencing of the V4 region of the 16S rRNA gene (bacteria and archaea) and the ITS1 region (fungi). We extracted microbial DNA (250 mg FW soil from the organic topsoil layer and 400 mg FW soil from all other soil layers) using the FastDNA™ SPIN Kit for Soil (MP Biomedicals, Santa Ana, USA). We followed the manufacturers' instructions but added minor modifications for the removal of the RNAlater™ Stabilization Solution (addition of 1 mL of provided sodium phosphate buffer to soil pellet, vortex, brief centrifugation, discard supernatant, repeat procedure 5 times). Extraction blanks were included and subjected to subsequent quantification and sequencing steps. Amplicon sequencing and raw data processing was performed at the Joint Microbiome Facility of the Medical University of Vienna and the University of Vienna (JMF project ID JMF-2008-5). A two-step barcoding approach was employed to generate amplicon libraries of archaeal, bacterial, and fungal communities using Illumina MiSeq (V3 Kit, 2×300 bp configuration, 1 % PhiX spike-in), following Pjevac et al. (2021). We used the primer pairs 515F (GTGYCAGCMGCCGCGGTAA, (Parada et al., 2016) and 806R (GGACTACNVGGGTWTCTAAT, (Apprill et al., 2015) for amplifying the V4 hypervariable region of the 16S rRNA gene and the primer pairs ITS1F (CTTGGTCATTTAGAGGAAGTAA, (Smith and Peay, 2014) and ITS2 (GCTGCGTTCTTCATCGATGC, (White et al., 1990) for amplifying the fungal ITS1 region (amplification conditions in Supplement Sect. S4). Amplicon pools were extracted from the raw sequencing data using the FASTQ workflow in BaseSpace (Illumina) with default parameters. Demultiplexing was performed with the python package demultiplex (Laros JFJ, github.com/jfjlaros/demultiplex), allowing one mismatch for barcodes and two mismatches for linkers and primers (Pjevac et al., 2021). Amplicon sequence variants (ASVs) were inferred using the DADA2 R package applying the recommended workflow (Callahan et al., 2016a, b). FASTQ reads 1 and 2 were trimmed at 150 nt with allowed expected errors of 2 (16S rRNA gene) and 230 nt with allowed expected errors of 4 and 6 (ITS1 region), respectively. Bacterial and archaeal ASV sequences were classified using SINA version 1.6.1 (Pruesse et al., 2012) and the SILVA database SSU Ref NR 99 release 138.1 (Quast et al., 2013) using default parameters. Fungal ASVs were classified using DADA2 and the UNITE general FASTA release for eukaryotes (v.8.2), using default parameters (Abarenkov et al., 2020). We note that applying ASV-based approaches to fungal ITS regions may be affected by high sequence variability and intra-genomic variation, potentially influencing taxonomic resolution. Datasets were deposited in the NCBI Sequence Read Archive under BioProject accession number (PRJNA1274918).
Prior downstream analyses, we cleaned the amplicon sequencing datasets from non-archaeal, -bacterial, or -fungal sequences and excluded samples with less than 500 obtained reads. Contaminant sequences were removed on an ASV-specific basis by subtracting the highest observed read number in one of the DNA extraction blanks from the corresponding sample reads.
Rarefaction was applied to these datasets prior to assessing α-diversity (using the rarefy_even_depth-function implemented in phyloseq with replacement-argument = F; and determined cut-offs at 2650 16S rRNA reads and at 543 ITS1 reads, respectively). We note that rarefaction may remove rare taxa and is subject to ongoing debate (McMurdie and Holmes, 2014) but was applied here to standardize sequencing depth across samples. We assessed α-diversity as richness (number of observed ASVs) and Shannon diversity. We note that α-diversity estimates based on fungal ITS ASVs may be inflated due to intra-genomic and intra-specific variability and should therefore be interpreted with caution.
We performed digital droplet PCR (ddPCR) to quantify 16S rRNA genes and ITS1 regions with the same primers used for sequencing. Each ddPCR reaction had a volume of 22 µL and consisted of 1× QX200 ddPCR EvaGreen Supermix (BioRad), 0.1 µmol L−1 of each primer and 0.05 and 0.025 ng of template for the quantification of 16S rRNA genes and ITS1 regions, respectively. Droplets were generated on a QX200™ Droplet Generator (BioRad) and directly subjected to PCR amplification (amplification conditions in Table S4). PCR products in droplets were kept at 4 °C over night to increase their separation before measuring their fluorescence intensity (on a QX200™ Droplet Reader, BioRad). Gene copy numbers were calculated using the QX ONE Software Standard Edition (v. 1.2, BioRad) where thresholds between positive and negative droplet populations were set consistently for each sample using histograms as a guide. We expressed final ddPCR results as 16S rRNA and ITS1 gene copy numbers g−1 DW soil and used them as abundance proxies for bacteria and archaea, and fungi, respectively. We note that we did not explicitly correct for taxon-specific ribosomal gene copy number variation, and that ddPCR-derived abundance estimates therefore do not directly reflect microbial cell numbers.
To derive ddPCR-informed abundance estimates, we followed the framework of quantitative microbiome profiling (Vandeputte et al., 2017), in which amplicon-based relative abundances are scaled using gene copy numbers. Accordingly, we calculated ASV-level abundance estimates (gene copy number corrected reads per g soil DW) by multiplying the 16S rRNA or ITS1 gene copy numbers measured in ddPCR assays with their respective relative abundances from the amplicon sequencing datasets (based on the raw reads-dataset after removal of non-archaeal, -bacterial, or -fungal sequences, exclusion of samples with less than 500 obtained reads, and blank-correction). Rare bacterial, archaeal, and fungal taxa (defined as containing less than 0.05 % of all gene copy number corrected reads per sample) were excluded, resulting in 3643 bacterial, 137 archaeal, and 1604 fungal ASVs to be considered in the follow-up analyses. We explored quantitative differences of certain phyla (ddPCR-corrected reads per g DW aggregated on phylum level) between polygon types and soil layer categories. We visualized microbial community composition (β-diversity) by following a widely acknowledged approach for handling compositional data (e.g., Alteio et al., 2021; Barlow et al., 2020; Gloor et al., 2017): Principal Component Analysis (PCA) was performed on center-log-ratio-(clr-) transformed gene copy number corrected reads per g DW soil (corresponding to the “Aitchison distance”; Aitchison, 1984) at ASV level. For more details see Supplement Sect. S4, and https://doi.org/10.5281/zenodo.18631833 (Martin, 2026).
2.6 Microbial extracellular enzymatic activity
We measured the potential activities of six hydrolytic extracellular enzymes involved in carbon-, nitrogen-, phosphorus-, and sulfur-cycling: β-D-1,4-cellobiosidase (exoglucanase), β-D-1,4-glucosidase (glucosidase), β-1,4-N-acetyl-glucosaminidase (exochitinase), acid phosphatase, leucine-aminopeptidase (protease) and sulfatase, using microplate fluorometric assays as described in Canarini et al. (2021). For more details see Supplement Sect. S5.
2.7 Statistical analyses and data visualization
All analyses were performed in R Studio Version 4.1.2 (R Core Team, 2017, version 4.1.2). Significances of relationships were tested against a p < 0.05 threshold. Plots were generated using ggplot2 (Wickham, 2016) and partly edited using Inkscape (Inkscape, 2020).
We employed linear-mixed-effects models (lmes) to test all univariate variables for the fixed effects of “ice-wedge polygon type” and “soil layer category” plus their interaction. Therefore, we used the packages lme4 (Bates et al., 2015), lmerTest (Kuznetsova et al., 2017), emmeans (Lenth et al., 2022), and car (Fox and Weisberg, 2019). Due to the sites' very similar landscape, climate, soil, and vegetation, we determined the random effect in the lme model as specific soil pit ID blocked within the sampling site. Model results were inspected using the anova function with the default being a type III analysis of variance (ANOVA). In the case of no interactive effect being observed we used type II ANOVA to account for potential effects of different treatment replicates (Langsrud, 2003). We used the Estimated Marginal Means post hoc test to perform multiple comparisons (p.adjust = “tukey”) on the fixed effects of polygon type and soil layer category. In the case of an interactive effect being observed by ANOVA result and /or visual investigation of the data, we compared (a) differences between soil layers per polygon type and (b) differences between polygon types per soil layer category. If homogeneity of variances and normality of model residuals were not given, log or sqrt transformations were applied. In case of no agreement with model assumptions after transformation, we conducted nonparametric tests (for variable-specific details see https://doi.org/10.5281/zenodo.18631833, Martin, 2026). Kruskal Wallis tests were used to test the effects of polygon type and soil layer category, followed by pairwise two-sided Wilcoxon tests (function pairwise.wilcox.test, p.adjust = “bonferroni”). To check for possible interactive effects in a comparable manner as described for the lme models, we applied Wilcoxon tests on respectively subsetted parts of the dataset and visually checked the distribution of the examined parameter among the soil layer categories of each polygon type.
We employed the phyloseq package (McMurdie and Holmes, 2013) for handling the multivariate datasets on amplicon sequencing and SOM chemical composition. For visualization, we performed Principal Component Analyses (PCAs; function phyloseq::ordinate) on centered log-ratio (clr)- normalized data (microbiome::transform phyloseq.object, “clr”).
Prior to Permutational Multivariate Analysis of Variance (PERMANOVA) testing, we first checked for homogeneity of multivariate dispersions using Permutation Tests for Multivariate Dispersion Homogeneity (PERMDISP), implemented in vegan (vegan::betadisper function) with 999 permutations and the argument bias.adjust = TRUE to account for unequal sample sizes (Anderson, 2017). Subsequently, we performed PERMANOVAs (on euclidean distances) to explore the effects of polygon type and soil layer category, as well as their possible interaction (adonis function implemented in vegan with 999 permutations and p.adjust.m = “bonferroni”, vegan version 2.5–7; Oksanen et al., 2020). Differences between polygon types and/or soil layer categories were assessed using pairwise multilevel comparisons (pairwise.adonis function implemented in vegan with 999 permutations and p.adjust.m = “bonferroni”; Martinez Arbizu, 2020). In case of interactive effects, we used subsetted datasets for making pairwise tests. Analogous to the approach applied for the linear mixed-effects models, we tested (a) differences between soil layers within each polygon type and (b) differences between polygon types within each soil layer. Venn diagrams (get_vennlist(phyloseq.object)) were used for visualizing the fraction of shared versus unique pyrolysis products and/or microbial ASVs among polygon types and soil layers respectively (MicrobiotaProcess package, (Xu et al., 2022). We explored potential ecological roles of fungal communities using the FungalTraits database (Põlme et al., 2020), with trait-based annotations performed at the genus level. For more details see also Supplement Sect. S6, and https://doi.org/10.5281/zenodo.18631833 (Martin, 2026).
3.1 Physicochemical soil parameters and stoichiometry
We characterized soil properties either by polygon type (averaged across all soil layers) or by soil layer (averaged across all polygon types), respectively. LCPs differed from the other polygon types mainly in their soil C, N, and P contents. Soils of this polygon type had higher C and N concentrations across all layers, whereas P concentration were lower, especially in the organic layer (Table S2a). Consequently, C : P and N : P ratios were on average double as high in LCP soils compared to FCP or HCP soils. Based on mean values, LCPs were also characterized by the lowest soil bulk density, highest gravimetric soil water content, and deepest active layer among polygon types (Tables S1b, 2a), yet these differences were not statistically significant.
Physicochemical properties also exhibited pronounced shifts along the soil profile. In situ temperatures decreased steadily from approximately 5.6 °C at the surface to 1.4 °C at the permafrost table (Table S1b). The strongest contrasts often occurred between organic topsoils and mineral subsoils. For instance, mineral subsoils had a sixfold higher bulk density than the organic layer, but eightfold lower gravimetric water content. Organic topsoils further contained approximately fivefold higher soil C and N contents and twelvefold higher dissolved organic carbon (DOC) and total dissolved nitrogen (TDN) concentrations than mineral subsoils (Table S2b), whereas cryoturbated material and upper permafrost soils showed intermediate and relatively similar values. Soil C : N ratios remained largely consistent across soil layers, but C : P and N : P ratios were significantly lower in mineral subsoils. The DOC : TDN ratio was nearly twice as high in organic and mineral horizons compared to cryoturbated material and permafrost soils. While soil δ13C remained largely stable across layers, soil δ15N signals shifted with depth. The organic and mineral layers had more enriched signatures, the permafrost layer more depleted values, and cryoturbated material displayed an intermediate isotopic signature.
3.2 Soil organic matter composition
We assessed the chemical composition of organic matter pools in different polygon types and soil layer categories using pyrolysis-GC/MS. We noted a particularly distinct fingerprint pattern of LCP soils, whereas those of FCP and HCP soils were similar (Fig. 2a). Correspondingly, LCPs also shared much less pyrolysis products with the other polygon types than were shared among FCPs and HCPs (Fig. S2a). At the same time, LCP soils also had the smallest fraction of polygon-type specific pyrolysis products. Comparing SOM compound class abundances between polygon types revealed that LCP soils harbored significantly more lignin- derived substances than the other polygon types in absolute and relative terms, and higher absolute abundances of aromatics and phenols, lipids, and general and unknown compounds than FCP soils by trend (Figs. S3a, S4a).
Shifts in the chemical composition of SOM also occurred between soil layers, and this effect was comparatively stronger than the effect of polygon type (Fig. 2b). Organic topsoils and mineral subsoils were characterized by rather distinct SOM pools. Their chemical fingerprints differed significantly from those of all other soil layers and included a notable proportion of layer-specific pyrolysis products (10 % and 7 % of all considered pyrolysis products, respectively; Fig. S2b). The SOM fingerprints from the cryoturbated material and the permafrost layer could not be distinguished from another and only contained a small fraction of unique pyrolysis products (2.5 % of all pyrolysis products, respectively). Absolute abundances of SOM compound groups closely reflected the underlying soil carbon concentrations (Table S4). The highest absolute abundances across all six SOM groups were found in the organic topsoil, followed by intermediate levels in cryoturbated and permafrost layers, and the lowest abundances in the mineral subsoil, accordingly (Fig. S3b). To account for differences in total carbon content, it was hence more suitable to compare the relative abundances of SOM compound classes across soil layers. In relative proportions, aromatic and phenolic compounds were for example highest in mineral subsoils, whilst lowest in organic topsoils (Fig. S4b). Similarly, N-containing compounds were most scarce in the mineral layer in absolute terms, while in relative terms, cryoturbated material was the most limited.
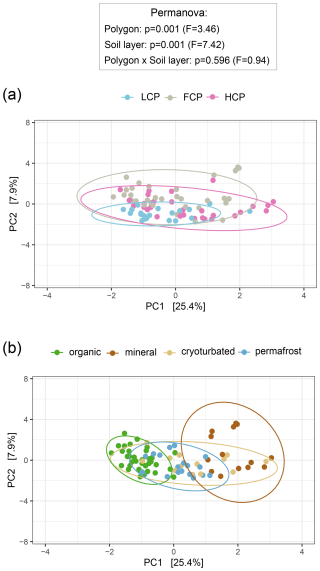
Figure 2Soil organic matter (SOM) composition across ice-wedge polygon types (a) and soil layers (b). Principal component analysis (PCA) was performed on center-log-ratio (clr)-transformed abundances (mg C g−1 DW) of 534 pyrolysis products to visualize variation in SOM composition. Statistical significance was assessed using PERMANOVA and pairwise comparisons (based on Euclidean distance matrices). Ellipses represent 95 % confidence intervals. (a) SOM composition differed between polygon types, with low-centered polygon (LCP) soils showing a distinct fingerprint compared to flat-centered (FCP) and high-centered polygon (HCP) soils (LCP vs. FCP: p = 0.003, F = 3.62; LCP vs. HCP: p = 0.018, F = 2.87; FCP vs. HCP: p = 0.072, F = 2.06). Group dispersions were homogeneous (Betadisper_polygons: p = 0.102, F = 2.36). Sample sizes: LCP = 20, FCP = 32, HCP = 29. (b) SOM composition differed across soil layers, with organic topsoils and mineral subsoils showing distinct profiles from each other and from all other soil layers (organic vs. mineral: p = 0 006, F = 17.01; organic vs. cryoturbated: p = 0.006, F = 6.43; organic vs. permafrost: p = 0.006, F = 5.47; mineral vs. cryoturbated: p = 0.012, F = 3.12; mineral vs. permafrost: p = 0.006, F = 7.04). Profiles of the cryoturbated and the permafrost layers could not be distinguished (p = 0.252, F = 1.64). Group dispersions were homogeneous (Betadisper_soil layers: p = 0.061, F = 2.57). Sample sizes: organic = 35; mineral = 14; cryoturbated = 13, permafrost = 19.
3.3 Microbial Communities
We analyzed microbial community composition by sequencing the bacterial and archaeal 16S rRNA gene and fungal ITS1 region. We used ddPCR-derived gene copy numbers to estimate microbial abundances and to assess differences in specific phyla of interest.
3.3.1 Bacterial and archaeal abundance proxies, alpha-, and beta- diversity
The dataset comprised 41 bacterial and six archaeal phyla. Bacteroidota (28.6 %), Proteobacteria (19.8 %), Verrumicrobiota (16.5 %), Acidobacteriota (14.2 %), and Actinobacteriota (4.5 %), represented the five most abundant phyla, and together accounted for 84 % of all obtained ddPCR-corrected reads. Archaea, by comparison, only comprised 1.8 % of the overall community. Taxonomic resolution was limited for a substantial proportion of the prokaryotic community members, as approximately more than a third (1173 out of 3780) of all bacterial and archaeal ASVs remained unclassified at the family level.
Compared to other polygon types, LCP soils exhibited lower richness, Shannon diversity, and reduced abundance of bacteria and archaea (Fig. S5). When 16S rRNA gene copy numbers were expressed per gram of dry soil, the lower abundance in LCP soils was only visible in the organic layer (interactive effect). However, when normalized to differences in soil carbon content, bacterial and archaeal abundance was consistently lower across all soil layers of LCPs. The structure of bacterial and archaeal communities in LCP soils also differed significantly from those in FCP and HCP soils (Fig. 3a). No significant difference was, however, found between FCP and HCP communities. The distinctiveness of LCP communities was also reflected in other observations. FCPs and HCPs, for example, shared 32 % of the total number of detected ASVs, whereas LCPs shared only 5 % with FCPs, and 2.5 % with HCPs (Fig. S6a). Furthermore, LCP soils also had the highest proportion of polygon-specific ASVs relative to total ASVs per polygon type (31 % for LCPs, 24 % for FCPs, and 18 % for HCPs), despite harboring a much lower total number of bacterial and archaeal ASVs (1449 for LCPs, 2791 for FCPs, and 2471 for HCPs). When comparing ddPCR-derived abundance estimates across polygon morphologies, we found that three of the five most dominant bacterial phyla, namely Proteobacteria, Verrucomicrobiota, and Actinobacteriota were significantly less abundant in soils of LCPs compared to FCPs and HCPs (Table S6, Fig. S8). Less abundant compared to either FCPs or HCPs were also Armatimonadota, Bdellovibrionata, Cyanobacterota, and Gemmatimonadota, and phyla, such as RCP2-54, or WPS-2 were nearly absent from LCP soils. By contrast, archaea were notably enriched in LCP soils, particularly in the topsoil layer (Fig. S7). LCP topsoils accounted for 65 % of ddPCR corrected archaeal reads in the dataset and were characterized by a high abundance of Euryarchaeota, Crenarchaeota, Micrarchaeota, Nanoarchaeota, and Halobacterota (Table S6, Fig. S8).This enrichment was evident in relative abundances as well: archaea comprised, on average, 6 % of the total community in LCP topsoils – a substantially higher fraction than in all other soil layers and polygon types. That LCP topsoils harbored particularly distinct communities, was also indicated by the significant interaction between polygon type and soil layer in the ordination (Fig. 3). For instance, Desulfobacterota (sulfate reducers), and Methylomirabilota (methane oxidizers), occurred in elevated abundances, whereas Acidobacteriota, Myxococcota (predators), and Planctomycetota were comparatively scarce (Table S6; Fig. S8).
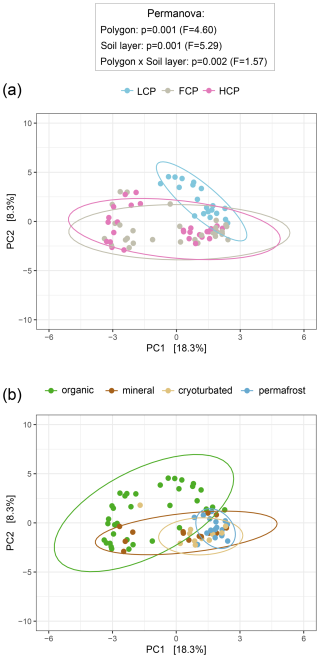
Figure 3Bacterial and archaeal community composition across ice-wedge polygon types (a) and soil layers (b). Principal component analysis (PCA) was performed on center-log-ratio (clr)-transformed abundance estimates (gene copy number corrected reads g−1 DW) of 3780 bacterial and archaeal ASVs to visualize community structure. Statistical significance was assessed using PERMANOVA and pairwise comparisons (based on Euclidean distance matrices). Ellipses represent 95 % confidence intervals. (a) Bacterial and archaeal community composition differed between polygon types, with low-centered polygon (LCP) soils hosting distinct assemblages compared to flat-centered (FCP) and high-centered polygon (HCP) soils (LCP vs. FCP: p = 0.003, F = 4.76; LCP vs. HCP: p = 0.003, F = 5.93; FCP vs. HCP: p = 0.183, F = 1.56). A significant interaction between polygon type and soil layer was detected: organic and permafrost layers in LCPs hosted distinct communities compared with their FCP and HCP counterparts (statistical details in Table S5). Note that heterogeneous dispersions between polygon types may have influenced these results (Betadisper_polygons: p = 0.006, F = 6.32). Sample sizes: LCP = 20, FCP = 30, HCP = 29. (b) Bacterial and archaeal community composition differed across soil layers, with organic topsoils and the permafrost layer showing distinct profiles in all polygon types (LCP: p = 0.006, F = 3.37; FCP: p = 0.006, F = 4.59; HCP: p = 0.006, F = 5.71), whereas those of the cryoturbated and the mineral layer could not be distinguished (FCP: p = 0.972, F = 1.51; HCP: p = 0.426, F = 1.31). A significant interaction between polygon type and soil layer was detected: in FCPs and HCPs, topsoil and permafrost communities also differed from those in other soil layers (statistical details in Table S5). Note that heterogeneous dispersions between soil layers may have influenced these results (Betadisper_soil layers: p = 0.001, F = 31.72). Sample sizes: organic = 35, mineral = 14, cryoturbated = 11, permafrost = 19).
Overall, soil layer had a stronger influence on microbial richness, alpha diversity, and abundance patterns than polygon morphology. All metrics declined significantly from the organic topsoil to the permafrost layer (Fig. S5). For instance, the organic layer harbored twice as many bacterial and archaeal ASVs as the permafrost layer and accounted for 75 % of all 16S rRNA gene copies per gram of dry soil in the dataset (compared to 3.6 % in the mineral subsoil, 11.7 % in cryoturbated material, and 10 % in the permafrost layer). Notably, organic topsoils remained a clear microbial abundance hotspot even after accounting for differences in soil carbon content.
Across all polygon types, bacterial and archaeal community structure differed significantly between the organic and permafrost layers, whereas communities in cryoturbated material and adjacent mineral soils were statistically indistinguishable (Fig. 3b). The communities in organic topsoils were particularly distinct, with approximately 40 % of the total number of bacterial and archaeal ASVs being unique to this layer (mineral layer 5 %, cryoturbated material 2%, and permafrost 3.5 %; Fig. S6b). The proportion of bacterial and archaeal ASVs that the organic layer shared with other layers declined with increasing soil depth (the organic layer shared 15 % with the mineral, 4 % with the cryoturbated, and 2.5 % with the permafrost layer, respectively). Comparing phylum-level ddPCR- informed abundances across soil layers showed that all five most abundant phyla occurred at substantially higher abundances in the organic topsoil than in the permafrost layer (Table S6; Fig. S8). The phyla that were mostly associated with the permafrost layer were Campylobacterota (93 % of all ddPCR corr. reads assigned to this phylum), Caldisericota (81 %), Cloacimonadota (68 %), and Firmicutes (54 %), but also the fraction of unknown taxa was notably high (43 %).
3.3.2 Fungal abundance proxies, alpha-, and beta-diversity
The fungal dataset comprised seven phyla, with Ascomycota and Basidiomycota being the most abundant, together accounting for approximately two thirds of all fungal ddPCR-corrected reads. Taxonomic resolution was limited for a substantial fraction of the dataset. More than 50 % of fungal ASVs (873 of 1604) could not be assigned at the phylum level, representing roughly one third of total fungal reads. Functional annotation using FUNGuild matched ecological roles for approximately 18 % of all fungal ASVs in the dataset, with ectomycorrhizal fungi (6.4 %), saprotrophs (7.7 %, including litter-, wood-, and soil-associated taxa), and root endophytes (1 %) being most prevalent (see also https://doi.org/10.5281/zenodo.18631833, Martin, 2026).
Fungal community patterns largely mirrored those observed for bacteria and archaea. LCP soils showed lower fungal richness, Shannon diversity, and abundances compared to FCPs and HCPs (Fig. S9). This lower fungal abundance was restricted to the organic layer when ITS1 gene copy numbers were expressed per gram dry soil (interactive effect) but became evident across all layers after normalization to soil carbon content. Fungal community structure also differed significantly in LCP soils (Fig. 4a), with only ∼ 5 % of taxa being shared between LCPs and the other polygon types (Fig. S10a). Although LCP soils harbored fewer fungal taxa overall, they contained the highest proportion of polygon-specific ASVs relative to total ASVs (LCPs 60 %, FCPs 55 %, HCPs 53 %). The comparison of phylum-level ddPCR- informed abundances further supported the presence of less rich, less diverse, and compositionally distinct fungal communities in LCP soils. Compared to FCP and HCP soils, Ascomycota, Basidiomycota, Chytridiomycota, and unclassified fungi occurred in lower abundances, Kickxellomycota were nearly absent, and Mortierellomycota, Rozellomycota and Zoopagomycota were not detected in LCP soils (Table S7, Fig. S11). Fungal Guilds analysis also suggested that LCP soils harbored a lower range of fungal lifestyles compared to the other polygon types, and that especially the fraction of ectomycorrhizal fungi was smaller.
Along the soil profile, fungal richness, diversity, and abundance declined sharply with depth (Fig. S9). The vast majority of fungal biomass was concentrated in the organic topsoil, which accounted for 96.2 % of all ITS1 gene copies in the dataset. In contrast, the contribution of fungi below the topsoil layer was minimal (2.4 % in cryoturbated material and <2 % combined in mineral and permafrost layers). Even after accounting for differences in soil carbon content, the organic layer remained a pronounced hotspot for fungal abundance. As indicated by ddPCR- informed abundance proxies, all fungal phyla occurred in an order of magnitude higher abundance levels in the organic topspoil layer compared to the other layers (Table S7, Fig. S11). Consistent with this, fungal community structure in organic topsoils was clearly distinct from that in deeper soil layers, independent of polygon type (Fig. 4b). This distinction was further reflected in taxon turnover along the soil profile. Half of all fungal ASVs in the dataset occurred exclusively in the organic layer, and the fraction of shared taxa between the organic layer and other layers also decreased with increasing depth (Fig. S10b). While all seven fungal phyla were present in the organic layer, six occurred in cryoturbated material, and only four could be detected in the mineral and permafrost layers, respectively (Table S7; Fig. S11). Fungal Guilds analysis also suggested shifts in fungal lifestyles along the soil profile. The contribution of ectomycorrhizal fungi and root endophytes decreased with depth, while some saprotrophic guilds (i.e., wood and soil saprotrophs) increased slightly.
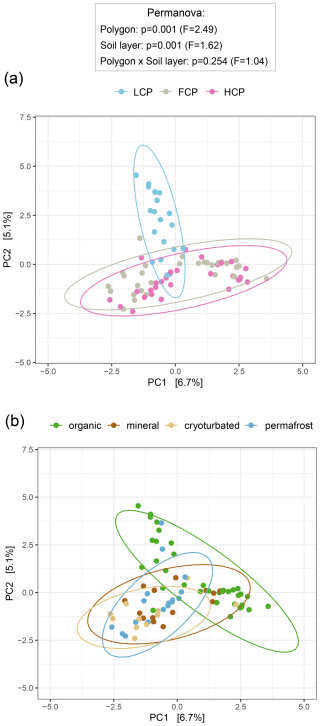
Figure 4Fungal community composition across ice-wedge polygon types (a) and soil layers (b). Principal component analysis (PCA) was performed on center-log-ratio (clr) -transformed abundance estimates (gene copy number corrected reads g−1 DW) of 1604 fungal ASVs to visualize community structure. Statistical significance was assessed using PERMANOVA and pairwise comparisons (based on Euclidean distance matrices). Ellipses represent 95 % confidence intervals. (a) Fungal community composition differed across polygons, with low-centered polygon (LCP) soils hosting distinct fungal assemblages compared to flat-centered (FCP) and high-centered polygon (HCP) soils (LCP vs. FCP: p = 0.003, F = 2.70; LCP vs. HCP: p = 0.003, F = 3.13; FCP vs. HCP: p = 0.009, F = 1.72). Note that heterogeneous dispersions between polygon types may have influenced the results (Betadisper_polygons: p = 0.039, F = 3.59). Sample sizes: LCP = 19, FCP = 30, HCP = 29. (b) Fungal community composition differed between soil layers, with a distinct fingerprint in the organic layer (organic vs. mineral: p = 0.024, F = 1.51; organic vs. cryoturbated: p = 0.006, F = 2.01; organic vs. permafrost: p = 0.006, F = 1.91). Note that heterogeneous dispersions between soil layers may have influenced the results (Betadisper_soil layers: p = 0.003, F = 5.83). Sample sizes: organic = 35, mineral = 14, cryoturbated = 12, permafrost = 17.
3.4 Potential extracellular enzymatic activity
We expressed enzyme rates per unit of soil carbon to account for a potential effect by diverging soil C concentrations (Table S8, Fig. S12). Overall, potential enzymatic activity per unit soil C varied greater between soil layers than between polygon types (Fig. 5). While activities of P- and S-cycling enzymes did not differ consistently between polygon types, C- and N-cycling enzyme rates were lower in LCP soils compared to FCP and HCP soils. However, this was mainly driven by differences in specific soil layers. LCPs exhibited reduced enzyme activities, with lower rates of betaglucosidase, exoglucanase, and leucine aminopeptidase in the organic layer, and diminished exochitinase activity in the permafrost layer (Table S9). Generally, enzyme activity profiles were relatively uniform across soil layers in LCPs but showed pronounced vertical variation in other polygon types. In FCPs and HCPs, activities of betaglucosidase, exoglucanase, and phosphatase still peaked in the organic layer despite normalization to soil carbon. In HCPs, exochitinase activity however reached its maximum in the cryoturbated and permafrost layers, whereas no layer-specific differences occurred in LCPs and FCPs.
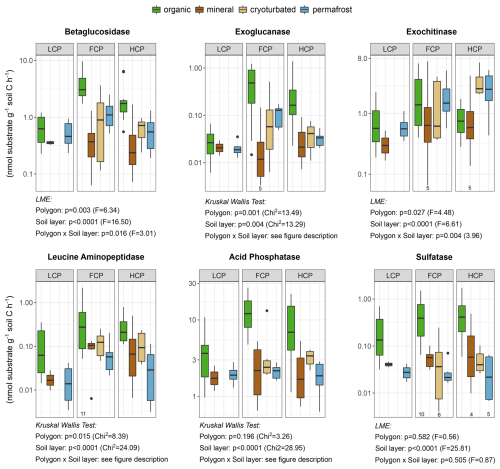
Figure 5Extracellular enzymatic activities across soil layers and ice-wedge polygon types. Potential activities of C-, N-, P- and S-acquiring enzymes are shown as rates (nmol substrate g−1 soil C h−1) on a logarithmic scale for improved readability. Boxplots depict median values and interquartile ranges. Sample sizes: LCP_organic = 12, FCP_organic = 12, HCP_organic = 11; LCP_mineral = 2, FCP_mineral = 6, HCP_mineral = 6; FCP_cryoturbated = 7, HCP_cryoturbated = 6; LCP_permafrost = 6, FCP_permafrost = 7, HCP_permafrost = 6. Individual deviations are indicated below the respective boxplots. Effects of polygon type and soil layer category are indicated below the respective panels (LME ANOVA type III or Kruskal Wallis test results). Overall, low-centered polygon (LCP) soils exhibited lower activities of C- and N-cycling enzymes compared to flat-centered (FCP) and high-centered polygon (HCP) soils. This effect was largely driven by layer-specific differences (i.e., interaction effects), including reduced betaglucosidase, exoglucanase, leucine-aminopeptidase activities in the LCP organic layer, and reduced exochitinase activity in the LCP permafrost layer. A further interaction showed that enzyme activities, including betaglucosidase, exoglucanase, exochitinase, acid phosphatase were more constant across soil layers in LCPs, but varied considerably in FCPs or HCPs. All statistical details are provided in Table S9. Please note that N-, P-, and S-depolymerizing enzymes are functionally linked to carbon cycling, which limits a strict distinction between microbial nutrient acquisition and C acquisition.
In this study, we asked how ice-wedge polygon microtopography and soil layers jointly shape soil organic matter pools and microbial communities, and whether the resulting spatial patterns can guide the upscaling of soil processes across spatially heterogeneous Arctic lowland tundra ecosystems.
4.1 Effects of Polygon Morphology
Across most measured characteristics, FCP and HCP soils were broadly similar, whereas LCP soils consistently stood out as markedly different. This disparity reflects intrinsic features of LCPs, including their distinctive vegetation cover, peaty soils, and persistent summer water saturation. Together, these factors affect a wide range of physical, chemical, and biological processes, and likely shaped the pronounced differences in soil properties, SOM characteristics, and microbial communities observed in this polygon type.
Prolonged waterlogging in LCPs strongly restricts microbial decomposition (Dungait et al., 2012; Schädel et al., 2014) and likely contributed to the elevated soil C and N concentrations through the accumulation of plant-derived material. However, for biogeochemical modeling, organic matter quality is considered as important as its quantity (Jansson and Taş, 2014; Mackelprang et al., 2016; Treat et al., 2014). Although soil C : N ratios are used as convenient and easily available proxy for OM availability (Malmer and Holm, 1984; Schädel et al., 2013, 2014; Weiss et al., 2016), they may fail to capture important compositional differences. For our study, we therefore relied on pyrolysis-GC/MS, a method that has been successfully applied in permafrost studies (Folhas et al., 2025; Keskitalo et al., 2021; Verret et al., 2025), and which provides compound-level details that bulk indices like C : N ratios cannot capture. Indeed, soil C : N ratios were similar across polygon types (Table S2b), but pyrolysis-GC/MS fingerprinting revealed pronounced differences in SOM quality (Fig. 2a). LCP soils showed the least variability in SOM composition along both PCA axes and contained the lowest absolute and relative shares of polygon-specific pyrolysis products (Fig. S2a). This points to a more uniform SOM profile, consistent with the relatively homogeneous nature of graminoid-, or moss-derived peat. In contrast, the more dispersed SOM fingerprints of FCPs and HCPs may reflect their more diverse plant cover and a broader spectrum of litter and root-derived inputs.
Beyond vegetation effects, the distinct SOM pattern in LCP soils likely also mirrors the water saturated, low-oxygen conditions that are typical for this polygon type during summer. Being strongly regulated by moisture and oxygen availability (Schmidt et al., 2011; Weintraub and Schimel, 2003), microbial degradation is less efficient under anaerobic than under aerobic conditions (Brune et al., 2000). In anaerobic environments, microbes preferentially consume readily bioavailable substrates, such as carbohydrates and organic acids, while structurally more complex substrates, including long-chained lipids, unsaturated hydrocarbons, lignin, and phenolic compounds tend to accumulate (Tveit et al., 2013, Wilson et al., 2022). Low-oxygen conditions also impede the activity of oxidative enzymes which mediate the breakdown of lignin and phenolic substances (Freeman et al., 2001, 2004; Tveit et al., 2013). Especially in Sphagnum-rich peat soils, these constraints may be further amplified by moss-derived compounds that directly inhibit microbial activity (Fofana et al., 2022; Turetsky, 2003). Together, these mechanisms likely contributed to the elevated relative abundances of lignin-derived compounds, aromatics and phenolics observed in LCP soils (Fig. S3a).
Polygon morphology also strongly influenced microbial abundance, diversity, and enzymatic activity, aligning well with previous reports of distinct microbial communities and metabolic pathways across polygon types (Taş et al., 2018; Wainwright et al., 2015). Although microbial biomass often scales with soil carbon content (Bastida et al., 2021; McGonigle and Turner, 2017), microbial abundance was lowest in the carbon-rich LCP soils, a pattern that even remained after normalizing gene copy numbers to soil carbon content (Fig. S5). This strongly indicates that another factor beyond soil carbon acts as overarching force in shaping the LCP microbiome. Redox conditions have been proposed as primary drivers of microbial communities (Lipson et al., 2015) through their influence on oxygen availability, pH, and organic matter quantity and quality. Long-lasting anaerobic conditions as typical for LCP centers, require adapted communities. In our dataset, this is reflected by LCP soils hosting taxonomically distinct communities with minimal overlap with FCP and HCP communities and the highest proportion of polygon-specific taxa (Figs. S6a, S10a). Typical of anoxic environments (Lynch et al., 2023), microbial richness and diversity were also lower in LCPs (Figs. S5, S9). On the one hand, this likely reflects the scarcity of obligate aerobic taxa. For example, the previously noted enrichment of lignin-derived substances in LCP soils (Figs. S3a, S4a) corresponds with the low abundance of aerobic, lignin-degrading Basidiomycota (Table S7; Fig.S11), (Zak and Kling, 2006). On the other hand, anaerobic metabolic pathways such as fermentation, methanotrophy, and respiration via alternative electron acceptors yield less energy than aerobic respiration (Madigan et al., 2021), which may constrain microbial growth yields and the number of organisms that can be sustained. Archaea likely contributed disproportionately to the distinct prokaryotic community structure of LCP soils (Fig. 3a). Although archaea generally represent a minor fraction of Arctic soil microbiomes (e.g., Gittel et al., 2014; Müller et al., 2018; Wilhelm et al., 2011), they can become prominent in waterlogged and peaty soils. In our dataset, archaea accounted for just 1.8 % of all prokaryotic ddPCR-corrected reads overall but were strongly enriched in LCP soils (7.3 %) compared to FCP (1.2 %) and HCP (0.2 %) soils (Fig. S7). Notably, Crenarchaeota and Euryarchaeota were particularly enriched in LCP soils (Table S6; Fig. S8). Members of these groups are known to participate in key peatland biogeochemical processes, including methanogenesis and anaerobic methane cycling (Tveit et al., 2013). Beyond the briefly noted influence of redox conditions, the distinct fungal community fingerprint observed in LCP soils (Fig. 4a) is likely driven by differences in plant species composition (Chu et al., 2011; Malard and Pearce, 2018; Wallenstein et al., 2007). For instance, LCPs are dominated by sedges and mosses (e.g., Eriophorum sp., Sphagnaceae, Amblystegiaceae), which typically lack mycorrhizal associations (Chen et al., 2020), whereas FCPs and HCPs are richer in dwarf shrubs (e.g., Betula, Salix, Ericaceae) that form mycorrhizal symbioses (Lynch et al., 2018). Consistent with these vegetation differences, the relative abundance of ectomycorrhizal fungi was substantially lower in LCP soils (2.2 %) than in FCP (7.8 %) and HCP (6.8 %) soils (FUNGuild analysis, Supplement Sect. S4).
Finally, polygon-specific differences in microbial communities and SOM characteristics were mirrored in extracellular enzyme activities. Heterotrophic microbes rely on extracellular enzymes to depolymerize high-molecular-weight substrates, with enzyme production being regulated by substrate availability and microbial demand (Burns et al., 2013; Moorhead et al., 2012). Accordingly, the lower potential activity of hydrolytic C- and N-acquiring enzymes in LCP soils, particularly in the organic layer (Fig. 5), corresponds well with the lower abundance of hydrolysable substrates (Figs. S3, S4) and the lower microbial abundance estimates, and diversity (Figs. S5, S9) observed in these soils. Especially fungi are considered key decomposers, producing a wide suite of hydrolytic and oxidative enzymes (Baldrian et al., 2010; Schneider et al., 2012). In high-latitude ecosystems, mycorrhizal fungi play a crucial role in the hydrolytic degradation of proteins (Bending and Read, 1996; Read and Perez-Moreno, 2003), while white-rot fungi, including Basidiomycota, oxidatively degrade lignin and humified SOM (Hatakka, 2005; Lee et al., 2012b). Both groups, however, occurred in lower abundances in LCP soils (Table S7, Fig. S11, FUNGuild analysis). Although our assays targeted only hydrolytic enzymes, the accumulation of lignin and phenolic compounds in LCP soils (Fig. S3a) suggests that oxidative enzyme activity was suppressed too, likely as a consequence of low-oxygen conditions (Freeman et al., 2004; Tveit et al., 2013). Elevated concentrations of polyphenols may further reinforce this finding by exerting a negative feedback on enzyme activity (Kostka et al., 2016). Finally, because enzyme production is energetically costly (Wortel et al., 2018), aerobic communities in FCPs and HCPs may be able to allocate more energy toward enzyme synthesis than the predominantly anaerobic communities in LCPs.
4.2 Effects of Soil Layer
While LCP soils consistently differed from FCP and HCP soils, soil-layer effects were more nuanced, with each layer exhibiting characteristic features. We advocate that many of the observed patterns can be interpreted along gradients in redox conditions, SOM content, or their interplay. Soil pH, for example, was consistently lower in organic topsoils than in the permafrost layer (Table S2b), a pattern also reported by Gentsch et al. (2018). This contrast suggests that the dominant controls on soil pH vary with depth, even though both layers may experience oxygen-limitations at times. In the permafrost layer, inorganic redox reactions and proton-consuming microbial processes (e.g., iron-, manganese-, sulfate-, or nitrate reduction) likely exert a strong influence on soil pH. Contrastingly, pH in the topsoil layer is more strongly shaped by acidifying plant inputs, including organic acids released as root exudates (Vives-Peris et al., 2020), or Sphagnum mosses, which acidify their surrounding via their metabolism and galacturonic acid-rich biomass (Kostka et al., 2016).
Broad-scale indicators for SOM composition and degree of processing, such as bulk soil C : N ratios and δ13C signatures remained remarkably stable across soil layers, (Table S2b), likely due to extensive cryoturbation activity in the study area. However, finer-scale changes in SOM quality became evident from the pyrolysis-GC/MS analysis. The ordination pattern (Fig. 2b) revealed a pronounced compositional shift along the first PCA axis that closely mirrored the concomitant gradient in soil carbon content (two-sided Spearman rank order correlation: Soil C - PCA axis 1: , p<0.0001). Layer-specific differences in SOM compound group abundances further indicated a progressive shift in the degree of organic matter transformation along the soil profile that was consistent across polygon types. Organic topsoils were relatively enriched in lignins, carbohydrates, and general and unknown compounds carbohydrates (Fig. S4b), likely reflecting inputs of little decomposed, labile plant detritus and root-derived substrates (Kuhry et al., 2020). In contrast, mineral subsoils were characterized by the highest relative contributions of less bioavailable compound classes, such as aromatics, phenols, and lipids. This likely reflects reduced inputs of fresh organic matter (Iversen et al., 2015), together with restricted substrate exchange due to limited effective pore space pore water mobility in the dense soil material, alongside mineral stabilization mechanisms (Dao et al., 2022; Prater et al., 2020). Consequently, microbial communities in mineral subsoils rely more strongly on OM recycling, or on metabolizing the accumulated, less bioavailable substrates (Weintraub and Schimel, 2003; Wild et al., 2016). Based on the SOM ordination pattern and relative abundances of compound groups (Figs. 2b, S4b), the permafrost SOM pool appeared structurally most similar to that of organic topsoils. This resemblance aligns well with field observations of abundant structurally intact plant residues in the frozen material, and suggests that the upper permafrost contains a substantial reservoir of relatively undecomposed organic matter that may become microbially accessible upon thaw (Gentsch et al., 2018). Despite their relatively similar SOM profiles topsoils and the upper permafrost layer differed most strongly in their microbial alpha and beta diversity patterns. By comparison, C-rich cryoturbated material and its adjacent C-poor mineral soil hosted statistically indistinguishable microbial communities (Figs. 3b, 4b). Together, these patterns indicate that microbial community structure along the soil profile is more strongly shaped by depth-related physicochemical constraints, such as temperature or oxygen availability, than by SOM composition.
How microbiomes change along a permafrost soil profile has been of interest to a plethora of studies. These, in line with our results (Figs. S5, S9), reported a depth-dependent declines in microbial biomass (Jansson and Taş, 2014; Liebner et al., 2008; Wild et al., 2016; Wilhelm et al., 2011), richness (Lipson et al., 2015), and diversity (Frank-Fahle et al., 2014; Jansson and Taş, 2014; Liebner et al., 2008; Müller et al., 2018; Ping et al., 1998; Taş et al., 2018). Also depth-dependent transitions in community structure and metabolic pathways are well documented (Frank-Fahle et al., 2014; Mackelprang et al., 2011; Müller et al., 2018). In our dataset, a comparable community shift was likely indicated by the steadily decreasing fraction of shared taxa between organic topsoils and deeper layers (Figs. S6b, S10b). Across all investigated layers, microbial richness, diversity, and ddPCR-derived abundance estimates were lowest in the permafrost layer (Figs. S5, S9), as the permafrost table certainly marks a major physical and ecological boundary for microorganisms. In permafrost, subzero temperatures, limited liquid water and oxygen availability, elevated salinity, and restricted exchange with the active-layer microbiome (Doherty et al., 2020; Ernakovich et al., 2022), collectively select for specialized communities (Jansson and Taş, 2014). For example, the permafrost layer harbored a notable fraction of ddPCR corrected reads assigned to Cloacimonadota (68 %), Caldisericota (81 %), and Campylobacterota (93 %), (Table S6, Fig. S8), who comprise members that have been linked to anaerobic C turnover, including fermentation and sulfur/nitrogen redox processes (Liu et al., 2025; Ratnikova et al., 2026; Sun et al., 2023). Firmicutes were also abundant (54 %), consistent with their ability to persist in freeze–thaw transition zones through dormancy and spore formation (Galperin, 2016). In contrast to the permafrost layer, the organic layer emerged as pronounced microbial hotspot, especially for fungi (Figs. S5, S9). Several factors likely contributed to this pattern. First, most fungi are strongly confined to aerobic conditions (Zak and Kling, 2006). Second, root biomass declines rapidly with depth in tundra soils (Iversen et al., 2015), leading to a concomitant decrease in mycorrhizal associations (Gittel et al., 2014), as also observed in our dataset (8 % in organic topsoils vs. 5 % in the permafrost layer; FunGuild analysis). Third, dominant decomposers of plant-derived organic matter, such as Ascomycota and Basidiomycota (Wallenstein et al., 2007), may benefit from cellulose- and lignin-rich substrate inputs into the topsoil layer (Boer et al., 2005).
Finally, the organic layer also emerged as a hotspot for hydrolytic enzyme activity (Fig. 5), likely due to its exceptionally high microbial abundance per unit soil C, its comparatively diverse decomposer community (Figs. S5, S9), and high availability of little decomposed plant-derived substrates (Fig. S4b). Notably, the permafrost layer exhibited relatively high enzyme activities despite harboring the lowest microbial abundance estimates and diversity (Figs. 5; S12). This decoupling of microbial community structure from enzyme potentials suggests that hydrolytic degradation capacity is largely governed by SOM properties and less constrained by microbial abundance alone. At the same time, the substantial enzyme potential observed in permafrost soils could also reflect the persistence of extracellular enzymes under chronically cold conditions and/or a greater per-biomass investment in enzyme production required to access substrates in frozen environments. Nonetheless, the pronounced latent enzymatic potential that we observed indicates that permafrost-associated SOM may be rapidly mobilized following thaw.
4.3 Potential links to carbon–climate feedbacks
Widespread ice-wedge degradation has been documented across the Arctic (Abolt et al., 2020; Fraser et al., 2018; Jorgenson et al., 2006, 2022; Kartoziia, 2019), largely in response to ongoing climate change. As permafrost temperatures rise and disturbances such as thermokarst formation and flooding become more frequent, low-centered polygons may increasingly transition into high-centered ones (Kartoziia, 2019; Kokelj et al., 2014; Liljedahl et al., 2016; Nitzbon et al., 2019). Given that low-centered polygons in our study were characterized by lower microbial abundance, reduced organic matter bioavailability, and diminished hydrolytic enzyme potential, their conversion into high-centered polygons may accelerate soil carbon losses. Although drying tends to shift carbon emissions from CH4 toward the less potent CO2 (Lara et al., 2015; Sachs et al., 2010), cumulative carbon losses may still be higher (Lee et al., 2012a; Schädel et al., 2016).
Topsoils, here identified as hotspots of microbial abundance and hydrolytic degradation potential, are likely to be most affected by future warming. Rising temperatures are expected to stimulate microbial activity (Hutchins et al., 2019; Karhu et al., 2014; Schuur et al., 2015), thereby increasing the potential for carbon release from these carbon-rich horizons. However, warming also promotes active-layer deepening (Solomon et al., 2007; Westerveld et al., 2023) and episodic thaw events, such as active-layer detachments, which expose previously frozen substrates to microbial decomposition (Graham et al., 2012; Schmidt et al., 2011). Presuming that the upper permafrost contained a substantial pool of relatively undecomposed organic matter and that permafrost microbes exhibited considerable hydrolytic enzymatic potential, thaw progression could unlock significant additional carbon losses.
Notably, our study provides a detailed spatial characterization but lacks temporal resolution. However, accurate projections of tundra carbon balance under climate change require the integration of multiple ecosystem processes over time, including vegetation dynamics (Myers-Smith et al., 2020; Phoenix and Treharne, 2022; Wolter et al., 2016), rhizosphere priming effects (Friggens et al., 2025; Keuper et al., 2020; Wild et al., 2014, 2016), microbial community assembly and coalescence (Ernakovich et al., 2022; Monteux et al., 2020), or couplings to other biogeochemical cycles (Burke et al., 2022; Keuper et al., 2012; Treat et al., 2016). Nevertheless, our findings highlight the significance of spatial organization of lowland tundra landscapes for shaping soil organic matter pools, microbial communities, and climate relevant biogeochemical dynamics.
Improving predictions of future water, energy, and carbon fluxes in Arctic lowland tundra requires explicit treatment of its spatial heterogeneity. Here, we show how soil organic matter pools, microbial communities, and hydrolytic enzyme potentials differ across ice-wedge polygon types and soil layers, the two dominant axes of edaphic variability in these ecosystems.
Strong polygon-specific patterns persisted across all soil layers, and soil layer-specific effects were consistent across polygon types. This suggests that these two spatial dimensions largely act independently, with modest interactions. Across both axes, patterns aligned with gradients in organic matter inputs and redox conditions, which themselves arise from the dynamic interplay of microtopography, hydrology, and vegetation. These environmental gradients, in turn, shape microbial communities and constrain characteristic biogeochemical processes at both the polygon and soil layer scales.
Overall, our findings demonstrate that a limited set of spatial units captures a disproportionate share of edaphic, microbial, and biogeochemical variability in Arctic lowland tundra soils. Explicitly accounting for polygon morphology and major soil layers in ecosystem models, with low-centered polygons and organic topsoils as particularly informative and tractable units, therefore provides a practical framework for upscaling soil processes in geomorphologically complex landscapes, and for improving climate-relevant biogeochemical projections.
The data is accessible under: https://doi.org/10.5281/zenodo.18631833 (Martin, 2026).
The supplement related to this article is available online at https://doi.org/10.5194/bg-23-2761-2026-supplement.
VM conducted the field and laboratory work, curated and analyzed the data, prepared visualizations, and wrote the manuscript with input from co-authors. AR led the scientific conceptualization of the study, served as principal supervisor of the research, and secured funding. AR and GH were responsible for project administration; GH also provided scientific guidance and financial support for the expeditions. MF supported project administration and fieldwork logistics, contributed expertise on the research area, and assisted with manuscript preparation. Fieldwork was carried out by VM, JW, WAC, LD, RL, NS, AR, and GH. CR, JH, CUM, and MM assisted VM with laboratory work and sample analyses. AR, VM, CR, and MM collaboratively developed the conceptual framework for the pyrolysis–GC/MS fingerprinting methodology. CR also played a key role in data analysis related to soil organic matter and microbial community composition. HS contributed to amplicon sequencing and ddPCR assays and provided scientific input on microbial community analysis and manuscript preparation. BH was responsible for sequencing methodology and raw data processing.
The contact author has declared that none of the authors has any competing interests.
Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims made in the text, published maps, institutional affiliations, or any other geographical representation in this paper. The authors bear the ultimate responsibility for providing appropriate place names. Views expressed in the text are those of the authors and do not necessarily reflect the views of the publisher.
We gratefully acknowledge the logistical support provided by the team at AWI Potsdam during the Yukon Coast expeditions in the summers of 2018 and 2019. We thank Hugues Lantuit for establishing the foundation and framework that made this research possible, including funding, infrastructure, and permitting. We thank George Tanski for logistical and field support during sample collection in 2018; Alberto Canarini for significant assistance in developing the semi-automated pyrolysis-GC/MS fingerprinting workflow; Leila Jensen for guidance on ddPCR measurements; and Petra Pjevac for coordinating the amplicon sequencing process. We are especially grateful to Samuel McLeod, Frank Dillon, and Peter Archie for their invaluable assistance and contributions in the field. We further acknowledge the support of the Yukon Territorial Government, Yukon Parks (Herschel Island – Qikiqtaruk Territorial Park), and the Aurora Research Institute in Inuvik.
This research has been supported by the European Commission, Horizon 2020 Framework Programme (Project “Nunataryuk” (grant no. 773421)).
This paper was edited by Erika Buscardo and reviewed by three anonymous referees.
Abarenkov, K., Zirk, A., Piirmann, T., Pöhönen, R., Ivanov, F., Nilsson, R. H., and Kõljalg, U.: UNITE general FASTA release for eukaryotes, UNITE Community, PlutoF repository, https://doi.org/10.15156/BIO/786371, 2020.
Abolt, C. J., Young, M. H., Atchley, A. L., and Harp, D. R.: Microtopographic control on the ground thermal regime in ice wedge polygons, The Cryosphere, 12, 1957–1968, https://doi.org/10.5194/tc-12-1957-2018, 2018.
Abolt, C. J., Young, M. H., Atchley, A. L., Harp, D. R., and Coon, E. T.: Feedbacks Between Surface Deformation and Permafrost Degradation in Ice Wedge Polygons, Arctic Coastal Plain, Alaska, J. Geophys. Res.-Earth, 125, e2019JF005349, https://doi.org/10.1029/2019JF005349, 2020.
Aitchison, J.: The statistical analysis of geochemical compositions, J. Int. Assoc. Math. Geol., 16, 531–564, https://doi.org/10.1007/BF01029316, 1984.
Alexander, E. B.: Bulk density equations for southern Alaska soils, Can. J. Soil Sci., 69, 177–180, https://doi.org/10.4141/cjss89-017, 1989.
Alteio, L. V., Séneca, J., Canarini, A., Angel, R., Jansa, J., Guseva, K., Kaiser, C., Richter, A., and Schmidt, H.: A critical perspective on interpreting amplicon sequencing data in soil ecological research, Soil Biol. Biochem., 160, https://doi.org/10.1016/j.soilbio.2021.108357, 2021.
Anderson, M. J.: Permutational Multivariate Analysis of Variance (PERMANOVA), in: Wiley StatsRef: Statistics Reference Online, edited by: Kenett, R. S., Longford, N. T., Piegorsch, W. W., and Ruggeri, F., Wiley, 1–15, https://doi.org/10.1002/9781118445112.stat07841, 2017.
Apprill, A., McNally, S., Parsons, R., and Weber, L.: Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton, Aquat. Microb. Ecol., 75, 129–137, 2015.
Arora, B., Wainwright, H. M., Dwivedi, D., Vaughn, L. J. S., Curtis, J. B., Torn, M. S., Dafflon, B., and Hubbard, S. S.: Evaluating temporal controls on greenhouse gas (GHG) fluxes in an Arctic tundra environment: An entropy-based approach, Sci. Total Environ., 649, 284–299, https://doi.org/10.1016/j.scitotenv.2018.08.251, 2019.
Baker, C. C. M., Barker, A. J., Douglas, T. A., Doherty, S. J., and Barbato, R. A.: Seasonal variation in near-surface seasonally thawed active layer and permafrost soil microbial communities, Environ. Res. Lett., 18, https://doi.org/10.1088/1748-9326/acc542, 2023.
Baldrian, P., Voříšková, J., Dobiášová, P., Merhautová, V., Lisá, L., and Valášková, V.: Production of extracellular enzymes and degradation of biopolymers by saprotrophic microfungi from the upper layers of forest soil, Plant and Soil, 338, 111–125, 2010.
Barbier, B. A., Dziduch, I., Liebner, S., Ganzert, L., Lantuit, H., Pollard, W., and Wagner, D.: Methane-cycling communities in a permafrost-affected soil on Herschel Island, Western Canadian Arctic: Active layer profiling of mcrA and pmoA genes, FEMS Microbiol. Ecol., 82, 287–302, https://doi.org/10.1111/j.1574-6941.2012.01332.x, 2012.
Barlow, J. T., Bogatyrev, S. R., and Ismagilov, R. F.: A quantitative sequencing framework for absolute abundance measurements of mucosal and lumenal microbial communities, Nat. Commun., 11, 2590, https://doi.org/10.1038/s41467-020-16224-6, 2020.
Bastida, F., Eldridge, D. J., García, C., Kenny Png, G., Bardgett, R. D., and Delgado-Baquerizo, M.: Soil microbial diversity–biomass relationships are driven by soil carbon content across global biomes, ISME J., 15, 2081–2091, https://doi.org/10.1038/s41396-021-00906-0, 2021.
Bates, D., Mächler, M., Bolker, B., and Walker, S.: Fitting Linear Mixed-Effects Models Using lme4, J. Stat. Softw., 67, 1–48, https://doi.org/10.18637/jss.v067.i01, 2015.
Beck, H., Zimmermann, N., McVicar, T., Vergopolan, N., Berg, A., and Wood, E.: Present and future Köppen-Geiger climate classification maps at 1-km resolution, Sci. Data, 5, 180214, https://doi.org/10.1038/sdata.2018.214, 2018.
Bending, G. D. and Read, D. J.: Nitrogen mobilization from protein-polyphenol complex by ericoid and ectomycorrhizal fungi, Soil Biol. Biochem., 28, 1603–1612, 1996.
Boer, W. de, Folman, L. B., Summerbell, R. C., and Boddy, L.: Living in a fungal world: impact of fungi on soil bacterial niche development?, FEMS Microbiol. Rev., 29, 795–811, https://doi.org/10.1016/j.femsre.2004.11.005, 2005.
Boike, J., Wille, C., and Abnizova, A.: Climatology and summer energy and water balance of polygonal tundra in the Lena River Delta, Siberia, J. Geophys. Res.-Biogeo., 113, https://doi.org/10.1029/2007JG000540, 2008.
Bottos, E. M., Kennedy, D. W., Romero, E. B., Fansler, S. J., Brown, J. M., Bramer, L. M., Chu, R. K., Tfaily, M. M., Jansson, J. K., and Stegen, J. C.: Dispersal limitation and thermodynamic constraints govern spatial structure of permafrost microbial communities, FEMS Microbiol. Ecol., 94, fiy110, https://doi.org/10.1093/femsec/fiy110, 2018.
Brooks, G. R. and Lane, L. S.: A guide to the landscape of the Firth River Valley, Ivvavik National Park, Parks Canada, https://publications.gc.ca/site/eng/9.840548/publication.html (last access: 17 August 2025), 2011.
Brown, J.: Tundra Soils Formed over Ice Wedges, Northern Alaska, Soil Sci. Soc. Am. J., 31, 686–691, https://doi.org/10.2136/sssaj1967.03615995003100050022x, 1967.
Brune, A., Frenzel, P., and Cypionka, H.: Life at the oxic–anoxic interface: microbial activities and adaptations, FEMS Microbiol. Rev., 24, 691–710, https://doi.org/10.1111/j.1574-6976.2000.tb00567.x, 2000.
Burke, E., Chadburn, S., and Huntingford, C.: Thawing Permafrost as a Nitrogen Fertiliser: Implications for Climate Feedbacks, Nitrogen, 3, https://doi.org/10.3390/nitrogen3020023, 2022.
Burns, R. G., DeForest, J. L., Marxsen, J., Sinsabaugh, R. L., Stromberger, M. E., Wallenstein, M. D., Weintraub, M. N., and Zoppini, A.: Soil enzymes in a changing environment: Current knowledge and future directions, Soil Biol. Biochem., 58, 216–234, https://doi.org/10.1016/j.soilbio.2012.11.009, 2013.
Callahan, B. J., Sankaran, K., Fukuyama, J. A., McMurdie, P. J., and Holmes, S. P.: Bioconductor Workflow for Microbiome Data Analysis: from raw reads to community analyses, Research, 5, F1000, https://doi.org/10.12688/f1000research.8986.2, 2016a.
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., and Holmes, S. P.: DADA2: High-resolution sample inference from Illumina amplicon data, Nat. Method., 13, 581–583, https://doi.org/10.1038/nmeth.3869, 2016b.
Canarini, A., Schmidt, H., Fuchslueger, L., Martin, V., Herbold, C. W., Zezula, D., Gündler, P., Hasibeder, R., Jecmenica, M., Bahn, M., and Richter, A.: Ecological memory of recurrent drought modifies soil processes via changes in soil microbial community, Nat. Commun., 12, 1–14, https://doi.org/10.1038/s41467-021-25675-4, 2021.
Chen, W., Tape, K. D., Euskirchen, E. S., Liang, S., Matos, A., Greenberg, J., and Fraterrigo, J. M.: Impacts of Arctic Shrubs on Root Traits and Belowground Nutrient Cycles Across a Northern Alaskan Climate Gradient, Front. Plant Sci., 11, 588098, https://doi.org/10.3389/fpls.2020.588098, 2020.
Chu, H., Neufeld, J. D., Walker, V. K., and Grogan, P.: The Influence of Vegetation Type on the Dominant Soil Bacteria, Archaea, and Fungi in a Low Arctic Tundra Landscape, Soil Sci. Soc. Am. J., 75, 1756–1765, https://doi.org/10.2136/sssaj2011.0057, 2011.
Clymo, R. S. and Hayward, P. M.: The Ecology of Sphagnum – Bryophyte Ecology, edited by: Smith, A. J. E., Springer Netherlands, Dordrecht, 229–289, https://doi.org/10.1007/978-94-009-5891-3_8, 1982.
Couture, N. J. and Pollard, W. H.: A Model for Quantifying Ground-Ice Volume, Yukon Coast, Western Arctic Canada, Permafrost Periglac., 28, 534–542, https://doi.org/10.1002/ppp.1952, 2017.
D'Angelo, E. and Crutchfield, J.: Rapid, Sensitive, Microscale Determination of Phosphate in Water and Soil, J. Environ. Qual., 30, 2206, 2001.
Dao, T. T., Mikutta, R., Sauheitl, L., Gentsch, N., Shibistova, O., Wild, B., Schnecker, J., Bárta, J., Čapek, P., Gittel, A., Lashchinskiy, N., Urich, T., Šantrůčková, H., Richter, A., and Guggenberger, G.: Lignin Preservation and Microbial Carbohydrate Metabolism in Permafrost Soils, J. Geophys. Res.-Biogeo., 127, https://doi.org/10.1029/2020JG006181, 2022.
Doherty, S. J., Barbato, R. A., Grandy, A. S., Thomas, W. K., Monteux, S., Dorrepaal, E., Johansson, M., and Ernakovich, J. G.: The Transition From Stochastic to Deterministic Bacterial Community Assembly During Permafrost Thaw Succession, Front. Microbiol., 11, 596589, https://doi.org/10.3389/fmicb.2020.596589, 2020.
Doherty, S. J., Thurston, A. K., and Barbato, R. A.: Active layer and permafrost microbial community coalescence increases soil activity and diversity in mixed communities compared to permafrost alone, Front. Microbiol., 16, 1579156, https://doi.org/10.3389/fmicb.2025.1579156, 2025.
Donner, N., Minke, M., de Klerk, P., Sofronov, R., and Joosten, H.: Patterns in polygon mires in north-eastern Yakutia, Siberia: the role of vegetation and water, Finnish Environ., 38, 19–30, 2012.
Dungait, J. A. J., Hopkins, D. W., Gregory, A. S., and Whitmore, A. P.: Soil organic matter turnover is governed by accessibility not recalcitrance, Glob. Change Biol., 18, 1781–1796, https://doi.org/10.1111/j.1365-2486.2012.02665.x, 2012.
Ernakovich, J. G., Lynch, L. M., Brewer, P. E., Calderon, F. J., and Wallenstein, M. D.: Redox and temperature-sensitive changes in microbial communities and soil chemistry dictate greenhouse gas loss from thawed permafrost, Biogeochemistry, 134, 183–200, https://doi.org/10.1007/s10533-017-0354-5, 2017.
Ernakovich, J. G., Barbato, R. A., Rich, V. I., Schädel, C., Hewitt, R. E., Doherty, S. J., Whalen, E. D., Abbott, B. W., Barta, J., Biasi, C., Chabot, C. L., Hultman, J., Knoblauch, C., Mackelprang, R., Onstott, T. C., Richter, A., Vishnivetskaya, T. A., Waldrop, M. P., and Winkel, M.: Microbiome assembly in thawing permafrost and its feedbacks to climate, Glob. Change Biol., 28, 5007–5026, https://doi.org/10.1111/gcb.16231, 2022.
Fofana, A., Anderson, D., McCalley, C. K., Hodgkins, S., Wilson, R. M., Cronin, D., Raab, N., Torabi, M., Varner, R. K., Crill, P., Saleska, S. R., Chanton, J. P., Tfaily, M. M., and Rich, V. I.: Mapping substrate use across a permafrost thaw gradient, Soil Biol. Biochem., 175, 108809, https://doi.org/10.1016/j.soilbio.2022.108809, 2022.
Folhas, D., Couture, R.-M., Laurion, I., Vieira, G., and Canário, J.: Natural organic matter dynamics in permafrost peatlands: Critical overview of recent findings and characterization tools, TrAC Trend. Anal. Chem., 184, 118153, https://doi.org/10.1016/j.trac.2025.118153, 2025.
Fox, J. and Weisberg, S.: An R Companion to Applied Regression, Third Edition, Sage: Thousand Oaks, CA, USA, https://cran.r-project.org/package=car (last access: 2 September 2025), 2019.
Frank-Fahle, B. A., Yergeau, É., Greer, C. W., Lantuit, H., and Wagner, D.: Microbial functional potential and community composition in permafrost-affected soils of the NW Canadian Arctic, PLoS ONE, 9, https://doi.org/10.1371/journal.pone.0084761, 2014.
Fraser, R. H., Kokelj, S. V., Lantz, T. C., McFarlane-Winchester, M., Olthof, I., and Lacelle, D.: Climate Sensitivity of High Arctic Permafrost Terrain Demonstrated by Widespread Ice-Wedge Thermokarst on Banks Island, Remote Sens., 10, https://doi.org/10.3390/rs10060954, 2018.
Freeman, C., Ostle, N., and Kang, H.: An enzymic “latch” on a global carbon store, Nature, 409, 149–149, https://doi.org/10.1038/35051650, 2001.
Freeman, C., Ostle, N. J., Fenner, N., and Kang, H.: A regulatory role for phenol oxidase during decomposition in peatlands, Soil Biol. Biochem., 36, 1663–1667, https://doi.org/10.1016/j.soilbio.2004.07.012, 2004.
French, H. M.: The Periglacial Environment, Third Edition, John Wiley & Sons Ltd, West Sussex PO19 8SQ, England, https://doi.org/10.1002/9781118684931.ch8, 2007.
Friggens, N. L., Hugelius, G., Kokelj, S. V., Murton, J. B., Phoenix, G. K., and Hartley, I. P.: Positive rhizosphere priming accelerates carbon release from permafrost soils, Nat. Commun., 16, 3576, https://doi.org/10.1038/s41467-025-58845-9, 2025.
Fritz, M., Wetterich, S., Schirrmeister, L., Meyer, H., Lantuit, H., Preusser, F., and Pollard, W. H.: Eastern Beringia and beyond: Late Wisconsinan and Holocene landscape dynamics along the Yukon Coastal Plain, Canada, Palaeogeogr. Palaeocl., 319/320, 28–45, https://doi.org/10.1016/j.palaeo.2011.12.015, 2012.
Fritz, M., Wolter, J., Rudaya, N., Palagushkina, O., Nazarova, L., Obu, J., Rethemeyer, J., Lantuit, H., and Wetterich, S.: Holocene ice-wedge polygon development in northern Yukon permafrost peatlands (Canada), Quaternary Sci. Rev., 147, 279–297, https://doi.org/10.1016/j.quascirev.2016.02.008, 2016.
Galperin, M.: Genome Diversity of Spore-Forming Firmicutes, in: The Bacterial Spore, 1–18, https://doi.org/10.1128/9781555819323.ch1, 2016.
Gentsch, N., Wild, B., Mikutta, R., Čapek, P., Diáková, K., Schrumpf, M., Turner, S., Minnich, C., Schaarschmidt, F., Shibistova, O., Schnecker, J., Urich, T., Gittel, A., Šantrůčková, H., Bárta, J., Lashchinskiy, N., Fuß, R., Richter, A., and Guggenberger, G.: Temperature response of permafrost soil carbon is attenuated by mineral protection, Glob. Change Biol., 24, 3401–3415, https://doi.org/10.1111/gcb.14316, 2018.
Gilichinsky, D., Rivkina, E., Shcherbakova, V., Laurinavichuis, K., and Tiedje, J.: Supercooled water brines within permafrost-an unknown ecological niche for microorganisms: a model for astrobiology, Astrobiology, 3, 331–341, https://doi.org/10.1089/153110703769016424, 2003.
Gittel, A., Bárta, J., Kohoutová, I., Mikutta, R., Owens, S., Gilbert, J., Schnecker, J., Wild, B., Hannisdal, B., Maerz, J., Lashchinskiy, N., Čapek, P., Šantrůčková, H., Gentsch, N., Shibistova, O., Guggenberger, G., Richter, A., Torsvik, V. L., Schleper, C., and Urich, T.: Distinct microbial communities associated with buried soils in the siberian tundra, ISME J., 8, 841–853, https://doi.org/10.1038/ismej.2013.219, 2014.
Gloor, G. B., Macklaim, J. M., Pawlowsky-Glahn, V., and Egozcue, J. J.: Microbiome Datasets Are Compositional: And This Is Not Optional, Front. Microbiol., 8, 2224, https://doi.org/10.3389/fmicb.2017.02224, 2017.
Graham, D. E., Wallenstein, M. D., Vishnivetskaya, T. A., Waldrop, M. P., Phelps, T. J., Pfiffner, S. M., Onstott, T. C., Whyte, L. G., Rivkina, E. M., Gilichinsky, D. A., Elias, D. A., Mackelprang, R., VerBerkmoes, N. C., Hettich, R. L., Wagner, D., Wullschleger, S. D., and Jansson, J. K.: Microbes in thawing permafrost: the unknown variable in the climate change equation, ISME J., 6, 709–712, https://doi.org/10.1038/ismej.2011.163, 2012.
Grosse, G., Harden, J., Turetsky, M., McGuire, A. D., Camill, P., Tarnocai, C., Frolking, S., Schuur, E. A. G., Jorgenson, T., Marchenko, S., Romanovsky, V., Wickland, K. P., French, N., Waldrop, M., Bourgeau-Chavez, L., and Striegl, R. G.: Vulnerability of high-latitude soil organic carbon in North America to disturbance, J. Geophys. Res.-Biogeo., 116, https://doi.org/10.1029/2010JG001507, 2011.
Hatakka, A.: Biodegradation of Lignin, in: Biopolymers Online, https://doi.org/10.1002/3527600035.bpol1005, 2005.
Hubbard, S. S., Gangodagamage, C., Dafflon, B., Wainwright, H., Peterson, J., Gusmeroli, A., Ulrich, C., Wu, Y., Wilson, C., Rowland, J., Tweedie, C., and Wullschleger, S. D.: Quantifying and relating land-surface and subsurface variability in permafrost environments using LiDAR and surface geophysical datasets, Hydrogeol. J., 21, 149–169, https://doi.org/10.1007/s10040-012-0939-y, 2013.
Hugelius, G., Kuhry, P., Tarnocai, C., and Virtanen, T.: Soil organic carbon pools in a periglacial landscape: a case study from the central Canadian Arctic, Permafrost Periglac., 21, 16–29, https://doi.org/10.1002/ppp.677, 2010.
Hultman, J., Waldrop, M. P., Mackelprang, R., David, M. M., McFarland, J., Blazewicz, S. J., Harden, J., Turetsky, M. R., McGuire, A. D., Shah, M. B., VerBerkmoes, N. C., Lee, L. H., Mavrommatis, K., and Jansson, J. K.: Multi-omics of permafrost, active layer and thermokarst bog soil microbiomes, Nature, 521, 208–212, https://doi.org/10.1038/nature14238, 2015.
Huntington, T. G.: Soil: Organic Matter and Available Water Capacity, in: Landscape and Land Capacity, edited by: Wang, Y., CRC Press, 273–281, https://doi.org/10.1201/9780429445552-36, 2020.
Hutchins, D. A., Jansson, J. K., Remais, J. V., Rich, V. I., Singh, B. K., and Trivedi, P.: Climate change microbiology – problems and perspectives, Nat. Rev. Microbiol., 17, 391–396, https://doi.org/10.1038/s41579-019-0178-5, 2019.
Inkscape Project: Inkscape, https://inkscape.org (last access: 10 February 2025), 2020.
IPCC (Ed.): Polar Regions, in: The Ocean and Cryosphere in a Changing Climate: Special Report of the Intergovernmental Panel on Climate Change, Cambridge University Press, Cambridge, 203–320, https://doi.org/10.1017/9781009157964.005, 2022.
Irwin, A.: Data Visualization - Making Maps, 2021.
Iversen, C. M., Sloan, V. L., Sullivan, P. F., Euskirchen, E. S., Mcguire, A. D., Norby, R. J., Walker, A. P., Warren, J. M., and Wullschleger, S. D.: The unseen iceberg: Plant roots in arctic tundra, New Phytol., 205, 34–58, https://doi.org/10.1111/nph.13003, 2015.
Jackson, R. B., Canadell, J., Ehleringer, J. R., Mooney, H. A., Sala, O. E., and Schulze, E. D.: A global analysis of root distributions for terrestrial biomes, Oecologia, 108, 389–411, https://doi.org/10.1007/BF00333714, 1996.
Jansson, J. and Taş, N.: The microbial ecology of permafrost, Nat. Rev. Microbiol., 12, 414–425, https://doi.org/10.1038/nrmicro3262, 2014.
Joabsson, A. and Christensen, T. R.: Methane emissions from wetlands and their relationship with vascular plants: an Arctic example, Glob. Change Biol., 7, 919–932, https://doi.org/10.1046/j.1354-1013.2001.00044.x, 2001.
Johnston, E. R., Hatt, J. K., He, Z., Wu, L., Guo, X., Luo, Y., Schuur, E. A. G., Tiedje, J. M., Zhou, J., and Konstantinidis, K. T.: Responses of tundra soil microbial communities to half a decade of experimental warming at two critical depths, P. Natl. Acad. Sci. USA, 116, 15096–15105, https://doi.org/10.1073/pnas.1901307116, 2019.
Jones, D. L.: Organic acids in the rhizosphere – a critical review, Plant Soil, 205, 25–44, https://doi.org/10.1023/A:1004356007312, 1998.
Jorgenson, M. T., Shur, Y. L., and Pullman, E. R.: Abrupt increase in permafrost degradation in Arctic Alaska, Geophys. Res. Lett., 33, https://doi.org/10.1029/2005GL024960, 2006.
Jorgenson, M. T., Kanevskiy, M. Z., Jorgenson, J. C., Liljedahl, A., Shur, Y., Epstein, H., Kent, K., Griffin, C. G., Daanen, R., Boldenow, M., Orndahl, K., Witharana, C., and Jones, B. M.: Rapid transformation of tundra ecosystems from ice-wedge degradation, Glob. Planet. Change, 216, 103921, https://doi.org/10.1016/j.gloplacha.2022.103921, 2022.
Karhu, K., Auffret, M. D., Dungait, J. A. J., Hopkins, D. W., Prosser, J. I., Singh, B. K., Subke, J.-A., Wookey, P. A., Agren, G. I., Sebastià, M.-T., Gouriveau, F., Bergkvist, G., Meir, P., Nottingham, A. T., Salinas, N., and Hartley, I. P.: Temperature sensitivity of soil respiration rates enhanced by microbial community response, Nature, 513, 81–84, https://doi.org/10.1038/nature13604, 2014.
Kartoziia, A.: Assessment of the ice wedge polygon current state by means of UAV imagery analysis (Samoylov Island, the Lena Delta), Remote Sens., 11, https://doi.org/10.3390/rs11131627, 2019.
Keskitalo, K. H., Bröder, L., Shakil, S., Zolkos, S., Tank, S. E., van Dongen, B. E., Tesi, T., Haghipour, N., Eglinton, T. I., Kokelj, S. V., and Vonk, J. E.: Downstream Evolution of Particulate Organic Matter Composition From Permafrost Thaw Slumps, Front. Earth Sci., 9, 642675, https://doi.org/10.3389/feart.2021.642675, 2021.
Keuper, F., Bodegom, P., Dorrepaal, E., Weedon, J., van Hal, J., Logtestijn, R., and Aerts, R.: A frozen feast: Thawing permafrost increases plant-available nitrogen in subarctic peatlands, Glob. Change Biol., 18, 1998–2007, https://doi.org/10.1111/j.1365-2486.2012.02663.x, 2012.
Keuper, F., Wild, B., Kummu, M., Beer, C., Blume-Werry, G., Fontaine, S., Gavazov, K., Gentsch, N., Guggenberger, G., Hugelius, G., Jalava, M., Koven, C., Krab, E. J., Kuhry, P., Monteux, S., Richter, A., Shahzad, T., Weedon, J. T., and Dorrepaal, E.: Carbon loss from northern circumpolar permafrost soils amplified by rhizosphere priming, Nat. Geosci., 13, 560–565, https://doi.org/10.1038/s41561-020-0607-0, 2020.
Knoblauch, C., Beer, C., Liebner, S., Grigoriev, M. N., and Pfeiffer, E. M.: Methane production as key to the greenhouse gas budget of thawing permafrost, Nat. Clim. Change, 8, 1–4, https://doi.org/10.1038/s41558-018-0095-z, 2018.
Kokelj, S. V., Lantz, T. C., Wolfe, S. A., Kanigan, J. C., Morse, P. D., Coutts, R., Molina-Giraldo, N., and Burn, C. R.: Distribution and activity of ice wedges across the forest-tundra transition, western Arctic Canada, J. Geophys. Res.-Earth, 119, 2032–2047, https://doi.org/10.1002/2014JF003085, 2014.
Kostka, J., Weston, D., Glass, J., Lilleskov, E., Shaw, A., and Turetsky, M.: The Sphagnum microbiome: New insights from an ancient plant lineage, New Phytol., 211, https://doi.org/10.1111/nph.13993, 2016.
Krantz, W. B.: Self-organization manifest as patterned ground in recurrently frozen soils, Earth-Sci. Rev., 29, 117–130, https://doi.org/10.1016/0012-8252(0)90031-P, 1990.
Kuhry, Peter., Barta, Jiri., Blok, Daan., Elberling, Bo., Faucherre, Samuel., Hugelius, Gustaf., Jørgensen, C. J., Richter, Andreas., ŠantrÅ?čková, Hana., and Weiss, Niels.: Lability classification of soil organic matter in the northern permafrost region, Biogeosciences, 17, 361–379, https://doi.org/10.5194/bg-17-361-2020, 2020.
Kuo, S.: Phosphorus, in: Methods of Soil Analysis, Vol. 3, Chemical Methods, edited by: Bartels, J. M. and Bigham, J. M., Soil Science Society of America, Madison, 869–919, 1996.
Kuznetsova, A., Brockhoff, P. B., and Christensen, R. H. B.: lmerTest Package: Tests in Linear Mixed Effects Models, J. Stat. Softw., 82, 1–26, https://doi.org/10.18637/jss.v082.i13, 2017.
Kwon, M. J., Jung, J. Y., Tripathi, B. M., Göckede, M., Lee, Y. K., and Kim, M.: Dynamics of microbial communities and CO2 and CH4 fluxes in the tundra ecosystems of the changing Arctic, J. Microbiol., 57, 325–336, https://doi.org/10.1007/s12275-019-8661-2, 2019.
Langsrud, Ø.: ANOVA for unbalanced data: Use Type II instead of Type III sums of squares, Stat. Comput., 13, 163–167, 2003.
Lara, M. J., Mcguire, A. David., Euskirchen, E. S., Tweedie, C. E., Hinkel, K. M., Skurikhin, A. N., Romanovsky, V. E., Grosse, Guido., Bolton, W. Robert., and Genet, Helene.: Polygonal tundra geomorphological change in response to warming alters future CO2 and CH4 flux on the Barrow Peninsula, Glob. Change Biol., 21, 1634–1651, https://doi.org/10.1111/gcb.12757, 2015.
Lara, M. J., Nitze, I., Grosse, G., and McGuire, A. D.: Tundra landform and vegetation productivity trend maps for the Arctic Coastal Plain of northern Alaska, Sci. Data, 5, 180058, https://doi.org/10.1038/sdata.2018.58, 2018.
Lee, H., Schuur, E. A. G., Inglett, K. S., Lavoie, M., and Chanton, J. P.: The rate of permafrost carbon release under aerobic and anaerobic conditions and its potential effects on climate, Glob. Change Bio., 18, 515–527, https://doi.org/10.1111/j.1365-2486.2011.02519.x, 2012a.
Lee, S., Jang, I., Chae, N., Choi, T., and Kang, H.: Organic Layer Serves as a Hotspot of Microbial Activity and Abundance in Arctic Tundra Soils, Microb. Ecol., 65, https://doi.org/10.1007/s00248-012-0125-8, 2012b.
Lenth, R. V., Buerkner, P., Herve, M., Love, J., Miguez, F., Riebl, H., and Singmann, H.: emmeans: Estimated Marginal Means, aka Least-Squares Means, R package version 1.7.2, https://cran.r-project.org/package=emmeans (last access: 2 September 2025), 2022.
Liebner, S., Harder, J., and Wagner, D.: Bacterial diversity and community structure in polygonal tundra soils from Samoylov Island, Lena Delta, Siberia, Int. Microbiol., 11, 195–202, https://doi.org/10.2436/20.1501.01.60, 2008.
Liljedahl, A. K., Boike, J., Daanen, R. P., Fedorov, A. N., Frost, G. V., Grosse, G., Hinzman, L. D., Iijma, Y., Jorgenson, J. C., Matveyeva, N., Necsoiu, M., Raynolds, M. K., Romanovsky, V. E., Schulla, J., Tape, K. D., Walker, D. A., Wilson, C. J., Yabuki, H., and Zona, D.: Pan-Arctic ice-wedge degradation in warming permafrost and its influence on tundra hydrology, Nat. Geosci., 9, 312–318, https://doi.org/10.1038/ngeo2674, 2016.
Lipson, D. A., Haggerty, J. M., Srinivas, A., Raab, T. K., Sathe, S., and Dinsdale, E. A.: Metagenomic Insights into Anaerobic Metabolism along an Arctic Peat Soil Profile, PLoS ONE, 8, https://doi.org/10.1371/journal.pone.0064659, 2013.
Lipson, D. A., Raab, T. K., Parker, M., Kelley, S. T., Brislawn, C. J., and Jansson, J.: Changes in microbial communities along redox gradients in polygonized Arctic wet tundra soils, Environ. Microbiol. Rep., 7, 649–657, https://doi.org/10.1111/1758-2229.12301, 2015.
Liu, Y., Yu, M., Chen, X., Ran, L., and Zhang, X.-H.: Diversity, metabolic potential and global distribution of the anaerobic fermentative bacteria Phylum Candidatus Cloacimonadota, Environ. Microb., 20, 136, https://doi.org/10.1186/s40793-025-00796-1, 2025.
Lynch, L. M., Machmuller, M. B., Cotrufo, M. F., Paul, E. A., and Wallenstein, M. D.: Tracking the fate of fresh carbon in the Arctic tundra: Will shrub expansion alter responses of soil organic matter to warming?, Soil Biol. Biochem., 120, 134–144, https://doi.org/10.1016/j.soilbio.2018.02.002, 2018.
Lynch, L. M., Calderon, F., and Ernakovich, J.: Greater regulation of permafrost organic matter composition by enzymes and redox than temperature, Soil Biol. Biochem., 180, 108991, https://doi.org/10.1016/j.soilbio.2023.108991, 2023.
MacKay, J.: Thermally induced movements in ice-wedge polygons, Western Arctic Coast: A long-term study, Géographie physique et Quaternaire, 54, 41, https://doi.org/10.7202/004846ar, 2000.
Mackelprang, R., Waldrop, M. P., Deangelis, K. M., David, M. M., Chavarria, K. L., Blazewicz, S. J., Rubin, E. M., and Jansson, J. K.: Metagenomic analysis of a permafrost microbial community reveals a rapid response to thaw, Nature, 480, 368–371, https://doi.org/10.1038/nature10576, 2011.
Mackelprang, R., Saleska, S. R., Jacobsen, C. S., Jansson, J. K., and Taş, N.: Permafrost Meta-Omics and Climate Change, Annu. Rev. Earth Pl. Sci., 44, 439–462, https://doi.org/10.1146/annurev-earth-060614-105126, 2016.
Mackelprang, R., Burkert, A., Haw, M., Mahendrarajah, T., Conaway, C. H., Douglas, T. A., and Waldrop, M. P.: Microbial survival strategies in ancient permafrost: Insights from metagenomics, ISME J., 11, 2305–2318, https://doi.org/10.1038/ismej.2017.93, 2017.
Madigan, M., Sattley, W., Aiyer, J., Stahl, D., and Buckley, D.: Brock Biology of Microorganisms, Global Edition, Pearson Deutschland, 1128 pp., ISBN 978-1-292-40479-0, 2021.
Malard, L. A. and Pearce, D. A.: Microbial diversity and biogeography in Arctic soils, Environ. Microbiol. Rep., 10, 611–625, https://doi.org/10.1111/1758-2229.12680, 2018.
Malmer, N. and Holm, E.: Variation in the C/N-Quotient of Peat in Relation to Decomposition Rate and Age Determination with 210Pb, Oikos, 43, 171–182, https://doi.org/10.2307/3544766, 1984.
Martin, V.: Spatial heterogeneity of soil organic matter and microbial community composition across ice-wedge polygons and soil layers in Arctic lowland tundra, Zenodo [data set], https://doi.org/10.5281/zenodo.18631833, 2026.
Martin, V., Schmidt, H., Canarini, A., Koranda, M., Hausmann, B., Müller, C. W., and Richter, A.: Soil cover shapes organic matter pools and microbial communities in soils of maritime Antarctica, Geoderma, 446, 116894, https://doi.org/10.1016/j.geoderma.2024.116894, 2024.
Martinez Arbizu, P.: pairwiseAdonis: Pairwise multilevel comparison using adonis, R package, https://github.com/pmartinezarbizu/pairwiseAdonis (last access: 2 September 2025), 2020.
McGonigle, T. P. and Turner, W. G.: Grasslands and Croplands Have Different Microbial Biomass Carbon Levels per Unit of Soil Organic Carbon, Agriculture, 7, https://doi.org/10.3390/agriculture7070057, 2017.
McMurdie, P. J. and Holmes, S.: phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data, PLOS ONE, 8, 1–11, https://doi.org/10.1371/journal.pone.0061217, 2013.
McMurdie, P. J. and Holmes, S.: Waste Not, Want Not: Why Rarefying Microbiome Data Is Inadmissible, PLOS Comput. Biol., 10, e1003531, https://doi.org/10.1371/journal.pcbi.1003531, 2014.
Minayeva, T., Sirin, A., Kershaw, P., and Bragg, O.: Arctic Peatlands, in: The Wetland Book: II: Distribution, Description, and Conservation, edited by: Finlayson, C. M., Milton, G. R., Prentice, R. C., and Davidson, N. C., Springer Netherlands, Dordrecht, 275–288, https://doi.org/10.1007/978-94-007-4001-3_109, 2018.
Monteux, S., Keuper, F., Fontaine, S., Gavazov, K., Hallin, S., Juhanson, J., Krab, E. J., Revaillot, S., Verbruggen, E., Walz, J., Weedon, J. T., and Dorrepaal, E.: Carbon and nitrogen cycling in Yedoma permafrost controlled by microbial functional limitations, Nat. Geosci., 13, 794–798, https://doi.org/10.1038/s41561-020-00662-4, 2020.
Moorhead, D., Lashermes, G., and Sinsabaugh, R.: A theoretical model of C- and N-acquiring exoenzyme activities, which balances microbial demands during decomposition, Soil Biol. Biochem., 53, 133–141, https://doi.org/10.1016/j.soilbio.2012.05.011, 2012.
Müller, O., Bang-Andreasen, T., White, R. A., Elberling, B., Taş, N., Kneafsey, T., Jansson, J. K., and Øvreås, L.: Disentangling the complexity of permafrost soil by using high resolution profiling of microbial community composition, key functions and respiration rates, Environ. Microbiol., 20, 4328–4342, https://doi.org/10.1111/1462-2920.14348, 2018.
Myers-Smith, I. H., Kerby, J. T., Phoenix, G. K., Bjerke, J. W., Epstein, H. E., Assmann, J. J., John, C., Andreu-Hayles, L., Angers-Blondin, S., Beck, P. S. A., Berner, L. T., Bhatt, U. S., Bjorkman, A. D., Blok, D., Bryn, A., Christiansen, C. T., Cornelissen, J. H. C., Cunliffe, A. M., Elmendorf, S. C., Forbes, B. C., Goetz, S. J., Hollister, R. D., De Jong, R., Loranty, M. M., Macias-Fauria, M., Maseyk, K., Normand, S., Olofsson, J., Parker, T. C., Parmentier, F.-J. W., Post, E., Schaepman-Strub, G., Stordal, F., Sullivan, P. F., Thomas, H. J. D., Tømmervik, H., Treharne, R., Tweedie, C. E., Walker, D. A., Wilmking, M., and Wipf, S.: Complexity revealed in the greening of the Arctic, Nat. Clim. Chang., 10, 106–117, https://doi.org/10.1038/s41558-019-0688-1, 2020.
Nitzbon, J., Langer, M., Westermann, S., Martin, L., Aas, K. S., and Boike, J.: Pathways of ice-wedge degradation in polygonal tundra under different hydrological conditions, The Cryosphere, 13, 1089–1123, https://doi.org/10.5194/tc-13-1089-2019, 2019.
Oksanen, J., Blanchet, F. G., Friendly, M., Kindt, R., Legendre, P., McGlinn, D., Minchin, P. R., O’Hara, R. B., Simpson, G. L., Solymos, P., Stevens, M. H. H., Szoecs, E., and Wagner, H.: vegan: Community Ecology Package, R package version 2.5-7, https://CRAN.R-project.org/package=vegan (last access: 3 September 2025), 2020.
Parada, A. E., Needham, D. M., and Fuhrman, J. A.: Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples, Environ. Microbiol., 18, 1403–1414, https://doi.org/10.1111/1462-2920.13023, 2016.
Phoenix, G. K. and Treharne, R.: Arctic greening and browning: Challenges and a cascade of complexities, Glob. Change Biol., 28, 3481–3483, https://doi.org/10.1111/gcb.16118, 2022.
Ping, C. L., Bockheim, J. G., Kimble, J. M., Michaelson, G. J., and Walker, D. A.: Characteristics of cryogenic soils along a latitudinal transect in arctic Alaska, J. Geophys. Res.-Atmos., 103, 28917–28928, https://doi.org/10.1029/98JD02024, 1998.
Ping, C. L., Michaelson, G. J., Kimble, J. M., Romanovsky, V. E., Shur, Y. L., Swanson, D. K., and Walker, D. A.: Cryogenesis and soil formation along a bioclimate gradient in Arctic North America, J. Geophys. Res.-Biogeo., 113, https://doi.org/10.1029/2008JG000744, 2008.
Ping, C. L., Jastrow, J. D., Jorgenson, M. T., Michaelson, G. J., and Shur, Y. L.: Permafrost soils and carbon cycling, SOIL, 1, 147–171, https://doi.org/10.5194/soil-1-147-2015, 2015.
Pjevac, P., Hausmann, B., Schwarz, J., Kohl, G., Herbold, C. W., Loy, A., and Berry, D.: An Economical and Flexible Dual Barcoding, Two-Step PCR Approach for Highly Multiplexed Amplicon Sequencing, Front. Microbiol., 12, https://doi.org/10.3389/fmicb.2021.669776, 2021.
Põlme, S., Abarenkov, K., Henrik Nilsson, R., Lindahl, B. D., Clemmensen, K. E., Kauserud, H., Nguyen, N., Kjøller, R., Bates, S. T., Baldrian, P., Frøslev, T. G., Adojaan, K., Vizzini, A., Suija, A., Pfister, D., Baral, H.-O., Järv, H., Madrid, H., Nordén, J., Liu, J.-K., Pawlowska, J., Põldmaa, K., Pärtel, K., Runnel, K., Hansen, K., Larsson, K.-H., Hyde, K. D., Sandoval-Denis, M., Smith, M. E., Toome-Heller, M., Wijayawardene, N. N., Menolli, N., Reynolds, N. K., Drenkhan, R., Maharachchikumbura, S. S. N., Gibertoni, T. B., Læssøe, T., Davis, W., Tokarev, Y., Corrales, A., Soares, A. M., Agan, A., Machado, A. R., Argüelles-Moyao, A., Detheridge, A., De Meiras-Ottoni, A., Verbeken, A., Dutta, A. K., Cui, B.-K., Pradeep, C. K., Marín, C., Stanton, D., Gohar, D., Wanasinghe, D. N., Otsing, E., Aslani, F., Griffith, G. W., Lumbsch, T. H., Grossart, H.-P., Masigol, H., Timling, I., Hiiesalu, I., Oja, J., Kupagme, J. Y., Geml, J., Alvarez-Manjarrez, J., Ilves, K., Loit, K., Adamson, K., Nara, K., Küngas, K., Rojas-Jimenez, K., Bitenieks, K., Irinyi, L., Nagy, L. G., Soonvald, L., Zhou, L.-W., Wagner, L., Aime, M. C., Öpik, M., Mujica, M. I., Metsoja, M., Ryberg, M., Vasar, M., Murata, M., Nelsen, M. P., Cleary, M., Samarakoon, M. C., Doilom, M., Bahram, M., Hagh-Doust, N., Dulya, O., Johnston, P., Kohout, P., Chen, Q., Tian, Q., Nandi, R., Amiri, R., and Perera, R. H.: FungalTraits: a user-friendly traits database of fungi and fungus-like stramenopiles, Fungal Diversity, 105, 1–16, https://doi.org/10.1007/s13225-020-00466-2, 2020.
Prater, I., Zubrzycki, S., Buegger, F., Zoor-Füllgraff, L. C., Angst, G., Dannenmann, M., and Mueller, C. W.: From fibrous plant residues to mineral-associated organic carbon – the fate of organic matter in Arctic permafrost soils, Biogeosciences, 17, 3367–3383, https://doi.org/10.5194/bg-17-3367-2020, 2020.
Pruesse, E., Peplies, J., and Glöckner, F. O.: SINA: Accurate high-throughput multiple sequence alignment of ribosomal RNA genes, Bioinformatics, 28, 1823–1829, https://doi.org/10.1093/bioinformatics/bts252, 2012.
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., Peplies, J., and Glöckner, F. O.: The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools, Nucl. Acid. Res., 41, 590–596, https://doi.org/10.1093/nar/gks1219, 2013.
Rampton, V. N.: Quaternary geology of the Yukon Coastal Plain, Geological Survey of Canada, Bulletin 317, 49 pp., Ottawa, Canada, https://doi.org/10.4095/111347, 1982.
Ratnikova, N. M., Merkel, A. Y., Novikov, A. A., and Slobodkin, A. I.: Isolation and characterization of anaerobic thermophilic bacterium Caldisericum insulae sp.nov., the second cultivated representative of the phylum Caldisericota, Extremophiles, 30, 7, https://doi.org/10.1007/s00792-026-01418-5, 2026.
R Core Team: R.: A language and environment for statistical computing, R Found. Stat. Comput. Vienna, Austria, 2017.
Read, D. J. and Perez-Moreno, J.: Mycorrhizas and nutrient cycling in ecosystems – a journey towards relevance?, New Phytol., 157, 475–492, https://doi.org/10.1046/j.1469-8137.2003.00704.x, 2003.
Roy Chowdhury, T., Berns, E. C., Moon, J.-W., Gu, B., Liang, L., Wullschleger, S. D., and Graham, D. E.: Temporal, Spatial, and Temperature Controls on Organic Carbon Mineralization and Methanogenesis in Arctic High-Centered Polygon Soils, Front. Microbiol., 11, https://doi.org/10.3389/fmicb.2020.616518, 2021.
Sachs, T., Giebels, M., Boike, J., and Kutzbach, L.: Environmental controls on CH4 emission from polygonal tundra on the microsite scale in the Lena river delta, Siberia, Glob. Change Biol., 16, 3096–3110, https://doi.org/10.1111/j.1365-2486.2010.02232.x, 2010.
Schädel, C., Luo, Y., David Evans, R., Fei, S., and Schaeffer, S. M.: Separating soil CO2 efflux into C-pool-specific decay rates via inverse analysis of soil incubation data, Oecologia, 171, 721–732, https://doi.org/10.1007/s00442-012-2577-4, 2013.
Schädel, C., Schuur, E. A. G., Bracho, R., Elberling, B., Knoblauch, C., Lee, H., Luo, Y., Shaver, G. R., and Turetsky, M. R.: Circumpolar assessment of permafrost C quality and its vulnerability over time using long-term incubation data., Glob. Change Biol., 20, 641–652, https://doi.org/10.1111/gcb.12417, 2014.
Schädel, C., Bader, M. K.-F., Schuur, E. A. G., Biasi, C., Bracho, R., Čapek, P., De Baets, S., Diáková, K., Ernakovich, J., Estop-Aragones, C., Graham, D. E., Hartley, I. P., Iversen, C. M., Kane, E., Knoblauch, C., Lupascu, M., Martikainen, P. J., Natali, S. M., Norby, R. J., O’Donnell, J. A., Chowdhury, T. R., Šantrůčková, H., Shaver, G., Sloan, V. L., Treat, C. C., Turetsky, M. R., Waldrop, M. P., and Wickland, K. P.: Potential carbon emissions dominated by carbon dioxide from thawed permafrost soils, Nat. Clim. Change, 6, 950–953, https://doi.org/10.1038/nclimate3054, 2016.
Schmidt, M. W. I., Torn, M. S., Abiven, S., Dittmar, T., Guggenberger, G., Janssens, I. a., Kleber, M., Kögel-Knabner, I., Lehmann, J., Manning, D. a. C., Nannipieri, P., Rasse, D. P., Weiner, S., and Trumbore, S. E.: Persistence of soil organic matter as an ecosystem property, Nature, 478, 49–56, https://doi.org/10.1038/nature10386, 2011.
Schnecker, J., Wild, B., Takriti, M., Eloy Alves, R. J., Gentsch, N., Gittel, A., Hofer, A., Klaus, K., Knoltsch, A., Lashchinskiy, N., Mikutta, R., and Richter, A.: Microbial community composition shapes enzyme patterns in topsoil and subsoil horizons along a latitudinal transect in Western Siberia, Soil Biol. Biochem., 83, 106–115, https://doi.org/10.1016/j.soilbio.2015.01.016, 2015.
Schneider, T., Keiblinger, K. M., Schmid, E., Sterflinger-Gleixner, K., Ellersdorfer, G., Roschitzki, B., Richter, A., Eberl, L., Zechmeister-Boltenstern, S., and Riedel, K.: Who is who in litter decomposition? Metaproteomics reveals major microbial players and their biogeochemical functions, ISME J., 6, 1749–1762, https://doi.org/10.1038/ismej.2012.11, 2012.
Schoeneberger, P. J., Wysocki, D. A., Benham, E. C., and Soil Survey Staff: Field book for describing and sampling soils, Version 3.0, Natural Resources Conservation Service, National Soil Survey Center, Lincoln, NE, 2012.
Schuur, E. A. G., Bockheim, J., Canadell, J. G., Euskirchen, E., Field, C. B., Goryachkin, S. V., Hagemann, S., Kuhry, P., Lafleur, P. M., Lee, H., Mazhitova, G., Nelson, F. E., Rinke, A., Romanovsky, V. E., Shiklomanov, N., Tarnocai, C., Venevsky, S., Vogel, J. G., and Zimov, S. A.: Vulnerability of Permafrost Carbon to Climate Change: Implications for the Global Carbon Cycle, BioScience, 58, 701–714, https://doi.org/10.1641/B580807, 2008.
Schuur, E. A. G., McGuire, A. D., Schädel, C., Grosse, G., Harden, J. W., Hayes, D. J., Hugelius, G., Koven, C. D., Kuhry, P., Lawrence, D. M., Natali, S. M., Olefeldt, D., Romanovsky, V. E., Schaefer, K., Turetsky, M. R., Treat, C. C., and Vonk, J. E.: Climate change and the permafrost carbon feedback, Nature, 520, 171–179, https://doi.org/10.1038/nature14338, 2015.
Shaver, G. R., Chapin, F. S., and Billings, W. D.: Ecotypic Differentiation in Carex Aquatilis on Ice-Wedge Polygons in the Alaskan Coastal Tundra, J. Ecol., 67, 1025–1045, https://doi.org/10.2307/2259226, 1979.
Shur, Y., Hinkel, K. M., and Nelson, F. E.: The transient layer: implications for geocryology and climate-change science, Permafrost Periglac., 16, 5–17, https://doi.org/10.1002/ppp.518, 2005.
Shur, Y., Jones, B. M., Jorgenson, M. T., Kanevskiy, M. Z., Liljedahl, A., Walker, D. A., Ward Jones, M. K., Fortier, D., and Vasiliev, A.: Formation of Low-Centered Ice-Wedge Polygons and Their Orthogonal Systems: A Review, Geosciences, 15, 249, https://doi.org/10.3390/geosciences15070249, 2025.
Siewert, M. B., Lantuit, H., Richter, A., and Hugelius, G.: Permafrost causes unique fine-scale spatial variability across tundra soils, Global Biogeochem. Cy., 1–19, https://doi.org/10.1029/2020gb006659, 2021.
Smith, D. P. and Peay, K. G.: Sequence depth, not PCR replication, improves ecological inference from next generation DNA sequencing, PloS one, 9, e90234, https://doi.org/10.1371/journal.pone.0090234, 2014.
Soil Classification Working Group: The Canadian System of Soil Classification, 187 pp., Ottawa, Agriculture Canada, 1998.
Solomon, S., Qin, D., Manning, M., Chen, Z., Marquis, M., Averyt, K. B., Tignor, M., and Miller, H. L.: Climate Change 2007: The Physical Science Basis. Contribution of Working Group I to the Fourth Assessment Report of the Intergovernmental Panel on Climate Change, Cambridge University Press, Cambridge, UK, ISBN 978-0521880096, 2007.
Speetjens, N. J., Tanski, G., Martin, V., Wagner, J., Richter, A., Hugelius, G., Boucher, C., Lodi, R., Knoblauch, C., Koch, B. P., Wünsch, U., Lantuit, H., and Vonk, J. E.: Dissolved organic matter characterization in soils and streams in a small coastal low-Arctic catchment, Biogeosciences, 19, 3073–3097, https://doi.org/10.5194/bg-19-3073-2022, 2022.
Sturtevant, C. S. and Oechel, W. C.: Spatial variation in landscape-level CO2 and CH4 fluxes from arctic coastal tundra: influence from vegetation, wetness, and the thaw lake cycle, Glob. Change Biol., 19, 2853–2866, https://doi.org/10.1111/gcb.12247, 2013.
Sun, Q., Xu, K., Cao, L., Du, Z., Wang, M., and Sun, L.: Nitrogen and sulfur cycling driven by Campylobacterota in the sediment–water interface of deep-sea cold seep: a case in the South China Sea, mBio, 14, e00117-23, https://doi.org/10.1128/mbio.00117-23, 2023.
Taş, N., Prestat, E., Wang, S., Wu, Y., Ulrich, C., Kneafsey, T., Tringe, S. G., Torn, M. S., Hubbard, S. S., and Jansson, J. K.: Landscape topography structures the soil microbiome in arctic polygonal tundra, Nat. Commun., 9, https://doi.org/10.1038/s41467-018-03089-z, 2018.
Treat, C. C., Wollheim, W. M., Varner, R. K., Grandy, A. S., Talbot, J., and Frolking, S.: Temperature and peat type control CO2 and CH4 production in Alaskan permafrost peats, Glob. Change Biol., 20, 2674–2686, https://doi.org/10.1111/gcb.12572, 2014.
Treat, C. C., Wollheim, W. M., Varner, R. K., and Bowden, W. B.: Longer thaw seasons increase nitrogen availability for leaching during fall in tundra soils, Environ. Res. Lett., 11, https://doi.org/10.1088/1748-9326/11/6/064013, 2016.
Turetsky, M. R.: The Role of Bryophytes in Carbon and Nitrogen Cycling, The Bryologist, 106, 395–409, 2003.
Tveit, A., Schwacke, R., Svenning, M. M., and Urich, T.: Organic carbon transformations in high-Arctic peat soils: Key functions and microorganisms, ISME Journal, 7, 299–311, https://doi.org/10.1038/ismej.2012.99, 2013.
Vandeputte, D., Kathagen, G., D'hoe, K., Vieira-Silva, S., Valles-Colomer, M., Sabino, J., Wang, J., Tito, R. Y., De Commer, L., Darzi, Y., Vermeire, S., Falony, G., and Raes, J.: Quantitative microbiome profiling links gut community variation to microbial load, Nature, 551, 507–511, https://doi.org/10.1038/nature24460, 2017.
Vaughn, L. J. S. and Torn, M. S.: Radiocarbon measurements of ecosystem respiration and soil pore-space CO2 in Utqiagvik (Barrow), Alaska, Earth Syst. Sci. Data, 10, 1943–1957, https://doi.org/10.5194/essd-10-1943-2018, 2018.
Verret, M., Naeher, S., Lacelle, D., Ginnane, C., Dickinson, W., Norton, K., Turnbull, J., and Levy, R.: Preservation and degradation of ancient organic matter in mid-Miocene Antarctic permafrost, Biogeosciences, 22, 5771–5786, https://doi.org/10.5194/bg-22-5771-2025, 2025.
Vives-Peris, V., de Ollas, C., Gómez-Cadenas, A., and Pérez-Clemente, R. M.: Root exudates: from plant to rhizosphere and beyond, Plant Cell Reports, 39, 3–17, https://doi.org/10.1007/s00299-019-02447-5, 2020.
Voigt, C., Lind, S. E., Lamprecht, R. E., Biasi, C., Martikainen, P. J., Marushchak, M. E., Novakovskiy, A., and Aurela, M.: Warming of subarctic tundra increases emissions of all three important greenhouse gases – carbon dioxide, methane, and nitrous oxide, Glob. Change Biol., 23, 3121–3138, https://doi.org/10.1111/gcb.13563, 2016.
Wagner, J., Martin, V., Speetjens, N. J., A'Campo, W., Durstewitz, L., Lodi, R., Fritz, M., Tanski, G., Vonk, J. E., Richter, A., Bartsch, A., Lantuit, H., and Hugelius, G.: High resolution mapping shows differences in soil carbon and nitrogen stocks in areas of varying landscape history in Canadian lowland tundra, Geoderma, 438, 116652, https://doi.org/10.1016/j.geoderma.2023.116652, 2023.
Wainwright, H., Dafflon, B., Smith, L., Hahn, M., Curtis, J., Wu, Y., Ulrich, C., Peterson, J., Torn, M., and Hubbard, S..: Identifying multiscale zonation and assessing the relative importance of polygon geomorphology on carbon fluxes in an Arctic tundra ecosystem, J. Geophys. Res.-Biogeo., 788–808, https://doi.org/10.1002/2014JG002799, 2015.
Waldrop, M. P., Chabot, C. L., Liebner, S., Holm, S., Snyder, M. W., Dillon, M., Dudgeon, S. R., Douglas, T. A., Leewis, M.-C., Walter Anthony, K. M., McFarland, J. W., Arp, C. D., Bondurant, A. C., Taş, N., and Mackelprang, R.: Permafrost microbial communities and functional genes are structured by latitudinal and soil geochemical gradients, ISME J., 17, 1224–1235, https://doi.org/10.1038/s41396-023-01429-6, 2023.
Waldrop, M. P., Ernakovich, J. G., Vishnivetskaya, T. A., Schaefer, S. R., Mackelprang, R., Barta, J., O'Brien, J. M., Winkel, M., Barbato, R. A., Heffernan, L., Leewis, M. C., Hewitt, R. E., Hultman, J., Sun, Y., Biasi, C., Bradley, J. A., Liebner, S., Ricketts, M. P., Muscarella, M. E., Schütte, U., Abuah, F., Whalen, E., Timling, I., Voigt, C., Taş, N., Lloyd, K. G., Siljanen, H. M. P., Rivkina, E. M., Voříšková, J., Tao, J., Liang, R., Li, Z., Lennon, J. T., and Onstott, T. C.: Microbial Ecology of Permafrost Soils: Populations, Processes, and Perspectives, Permafrost Periglac., 1–14, https://doi.org/10.1002/ppp.2264, 2025.
Walker, D. A., Raynolds, M. K., Daniëls, F. J. A., Einarsson, E., Elvebakk, A., Gould, W. A., Katenin, A. E., Kholod, S. S., Markon, C. J., Melnikov, E. S., Moskalenko, N. G., Talbot, S. S., Yurtsev, B. A., and The other members of the CAVM Team: The Circumpolar Arctic vegetation map, J. Veg. Sci., 16, 267–282, https://doi.org/10.1111/j.1654-1103.2005.tb02365.x, 2005.
Wallenstein, M. D., McMahon, S., and Schimel, J.: Bacterial and fungal community structure in Arctic tundra tussock and shrub soils, FEMS Microbiol. Ecol., 59, 428–435, https://doi.org/10.1111/j.1574-6941.2006.00260.x, 2007.
Walvoord, M. A. and Kurylyk, B. L.: Hydrologic Impacts of Thawing Permafrost – A Review, Vadose Zone J., 15, vzj2016.01.0010, https://doi.org/10.2136/vzj2016.01.0010, 2016.
Washburn, A. L.: Classification of patterned ground and review of suggested origins, GSA Bull., 67, 823–866, https://doi.org/10.1130/0016-7606(1956)67[823:COPGAR]2.0.CO;2, 1956.
Washburn, A. L.: Periglacial processes and environments, St. Martin's Press, New York, ISBN 0-7131-5653-8, 1973.
Weintraub, M. N. and Schimel, J. P.: Interactions between carbon and nitrogen mineralization and soil organic matter chemistry in arctic tundra soils, Ecosystems, 6, 129–143, https://doi.org/10.1007/s10021-002-0124-6, 2003.
Weiss, N., Blok, D., Elberling, B., Hugelius, G., Jørgensen, C. J., Siewert, M. B., and Kuhry, P.: Thermokarst dynamics and soil organic matter characteristics controlling initial carbon release from permafrost soils in the Siberian Yedoma region, Sediment. Geol., 340, 38–48, https://doi.org/10.1016/j.sedgeo.2015.12.004, 2016.
Westerveld, L., Kurvits, T., Schoolmeester, T., Eckhoff, T., Overduin, P., Fritz, M., Alfthan, B., Sinisalo, A., and Mulelid, O.: Arctic Permafrost Atlas, https://doi.org/10.61523/KPJI4549, 2023.
White, T. J., Bruns, T., Lee, S., and Taylor, J.: Amplification and Direct Sequencing of Fungal Ribosomal RNA Genes for Phylogenetics: In PCR – Protocols and Applications – A Laboratory Manual, Academic Press, 315–322, ISBN 0-12-372180-6, 1990.
Wickham, H.: ggplot2: Elegant Graphics for Data Analysis, R package version 3.3.6, https://cran.r-project.org/package=ggplot2 (last access: 3 September 2025), 2016.
Wild, B., Schnecker, J., Alves, R. J. E., Barsukov, P., Bárta, J., Čapek, P., Gentsch, N., Gittel, A., Guggenberger, G., Lashchinskiy, N., Mikutta, R., Rusalimova, O., Šantrůčková, H., Shibistova, O., Urich, T., Watzka, M., Zrazhevskaya, G., and Richter, A.: Input of easily available organic C and N stimulates microbial decomposition of soil organic matter in arctic permafrost soil, Soil Biol. Biochem., 75, 143–151, https://doi.org/10.1016/j.soilbio.2014.04.014, 2014.
Wild, B., Gentsch, N., Capek, P., Diáková, K., Alves, R. J. E., Bárta, J., Gittel, A., Hugelius, G., Knoltsch, A., Kuhry, P., Lashchinskiy, N., Mikutta, R., Palmtag, J., Schleper, C., Schnecker, J., Shibistova, O., Takriti, M., Torsvik, V. L., Urich, T., Watzka, M., Šantrūcková, H., Guggenberger, G., and Richter, A.: Plant-derived compounds stimulate the decomposition of organic matter in arctic permafrost soils, Sci. Rep., 6, 1–11, https://doi.org/10.1038/srep25607, 2016.
Wilhelm, R. C., Niederberger, T. D., Greer, C., and Whyte, L. G.: Microbial diversity of active layer and permafrost in an acidic wetland from the Canadian high arctic, Can. J. Microbiol., 57, 303–315, https://doi.org/10.1139/w11-004, 2011.
Wilson, R. M., Hough, M. A., Verbeke, B. A., Hodgkins, S. B., Tyson, G., Sullivan, M. B., Brodie, E., Riley, W. J., Woodcroft, B., McCalley, C., Dominguez, S. C., Crill, P. M., Varner, R. K., Frolking, S., Cooper, W. T., Chanton, J. P., Saleska, S. D., Rich, V. I., and Tfaily, M. M.: Plant organic matter inputs exert a strong control on soil organic matter decomposition in a thawing permafrost peatland, Sci. Total Environ., 820, 152757, https://doi.org/10.1016/j.scitotenv.2021.152757, 2022.
Wolter, J., Lantuit, H., Fritz, M., Macias-Fauria, M., Myers-Smith, I., and Herzschuh, U.: Vegetation composition and shrub extent on the Yukon coast, Canada, are strongly linked to ice-wedge polygon degradation, Polar Res., 35, 27489, https://doi.org/10.3402/polar.v35.27489, 2016.
Wortel, M. T., Noor, E., Ferris, M., Bruggeman, F. J., and Liebermeister, W.: Metabolic enzyme cost explains variable trade-offs between microbial growth rate and yield, PLOS Computat. Biol., 14, e1006010, https://doi.org/10.1371/journal.pcbi.1006010, 2018.
Xu, S., Li, Z., Tang, W., Dai, Z., Zhou, L., Feng, T., Chen, M., Liu, S., Fu, X., Wu, T., Hu, E., and Yu, G.: MicrobiotaProcess: A comprehensive R package for managing and analyzing microbiome and other ecological data within the tidy framework, R package, https://doi.org/10.21203/rs.3.rs-1284357/v1, 2022.
Xue, K., Yuan, M. M., Shi, Z. J., Qin, Y., Deng, Y., Cheng, L., Wu, L., He, Z., Van Nostrand, J. D., Bracho, R., Natali, S., Schuur, E. A. G., Luo, C., Konstantinidis, K. T., Wang, Q., Cole, J. R., Tiedje, J. M., Luo, Y., and Zhou, J.: Tundra soil carbon is vulnerable to rapid microbial decomposition under climate warming, Nat. Clim. Change, 6, 595–600, https://doi.org/10.1038/nclimate2940, 2016.
Zak, D. R. and Kling, G. W.: Microbial community composition and function across an arctic tundra landscape, Ecology, 87, 1659–1670, https://doi.org/10.1890/0012-9658(2006)87[1659:mccafa]2.0.co;2, 2006.




