the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Imprint of eutrophication on methane-cycling microbes in freshwater sediment
Alice Bosco-Santos
Eulalie Rose Beyala Bekono
Santona Khatun
Marie-Ève Monchamp
Joana Séneca
Petra Pjevac
Jasmine S. Berg
Eutrophication can alter methane (CH4) cycling in lakes, yet its long-term effect on sediment microbial communities remains unclear. We analyzed a 400 year-old sediment record from the historically eutrophied Lake Joux, Switzerland, combining porewater and solid-phase geochemistry with 16S rRNA gene amplicon analyses to elucidate the effects of nutrient and carbon loading on methanogenic and methanotrophic sediment communities. Lithological and chemical stratification defined three intervals (deep eutrophic, middle carbonate, upper eutrophic) correlated with changes in organic matter sources. Methanogens were clearly depth-partitioned: methylotrophic Methanomassiliicoccales dominated deep eutrophic sediments, whereas hydrogenotrophic Methanomicrobiales and Methanobacteriales increased upward in shallower, more recent sediments with fresher organic matter. Paired isotopic data support this substrate-driven shift in CH4 production. Although O2 was not detected below ∼ 0.4 cm, sequences of aerobic gammaproteobacterial methanotrophs (Crenothrix and Methylobacter) were abundant in surface sediments down to ∼ 20 cm sediment depth, correlating with and concentrations. The absence of anaerobic methanotrophs and C-isotopic evidence for ongoing methane oxidation suggest that these O2-requiring, methane monooxygenase-utilizing Methylococcales constitute the dominant CH4 sink in surface sediments. These findings reveal that eutrophication can cause a stratification of methane-cycling microbial communities, highlighting the role of sedimentary legacies in regulating benthic CH4 emissions from freshwater ecosystems.
- Article
(2023 KB) - Full-text XML
-
Supplement
(956 KB) - BibTeX
- EndNote
Freshwater ecosystems are significant sources of the greenhouse gas methane (CH4), with natural lakes estimated to contribute more than 70 % to freshwater CH4 emissions (Sanches et al., 2019). Despite their substantial contribution to atmospheric CH4, the mechanisms regulating CH4 emissions from lakes at regional and global scales remain poorly understood (Bastviken et al., 2011; Sanches et al., 2019). In freshwater sediments, CH4 is abundantly produced via anaerobic methanogenesis by archaea (Bastviken et al., 2004; Bastviken et al., 2011; Conrad, 2020; Dean et al., 2018; Saunois et al., 2020; Tranvik et al., 2009). Methanogens can respire different substrates produced during organic matter remineralization and are classified according to three known pathways for CH4 production: hydrogenotrophic (carbon dioxide reduction using hydrogen), acetoclastic (splitting acetate), and methylotrophic (using methylated compounds like methanol) (Garcia et al., 2000). Environmental factors such as substrate concentration, temperature, salinity, and pH influence the predominance of these pathways, with methylotrophic methanogenesis, for instance, being favored at higher salinity and acidity (Bueno De Mesquita et al., 2023; Yvon-Durocher et al., 2014).
Much of the CH4 produced in lake sediments is oxidized through both aerobic and anaerobic microbial processes before it can reach the atmosphere (Bastviken et al., 2004, 2008; Martinez-Cruz et al., 2017, 2018; Oswald et al., 2016). Aerobic oxidation, predominantly performed by methane-oxidizing bacteria (MOB) from the Gammaproteobacteria and Alphaproteobacteria classes, occurs at the sediment-water interface and in the water column (Hanson and Hanson, 1996; Knief, 2015). All MOB known to date rely on O2-dependent methane monooxygenase enzymes and oxidize up to 90 % of sediment-derived CH4, thus helping to mitigate greenhouse gas emissions from freshwater ecosystems (Bastviken et al., 2004; Bastviken et al., 2008). Anaerobic oxidation of methane (AOM) is performed by methanotrophic archaea (ANMEs), often in partnership with bacteria that use electron acceptors other than oxygen (Knittel and Boetius, 2009; Milucka et al., 2012; Wegener et al., 2015). In marine environments where sulfate concentrations are high, sulfate-AOM is the dominant process (Jørgensen et al., 2001; Wegener and Boetius, 2009). In contrast, the electron acceptors sulfate, nitrate/nitrite, humic substances, and diverse metal oxides contribute in various degrees to AOM in freshwater sediments (Chen et al., 2023; Deutzmann and Schink, 2011; Martinez-Cruz et al., 2018; Zhao et al., 2024).
Recently, it has been proposed that eutrophication induced by anthropogenic nutrient inputs (e.g., nitrates and phosphates) into lake ecosystems influences methanogen and methanotroph community structure by altering organic matter quality and quantity (Beaulieu et al., 2019; Yang et al., 2021, 2019, 2020; Zhu et al., 2022). The influx of organic carbon from phytoplankton blooms enhances organic matter mineralization in lake bottom waters and sediments, depleting electron acceptors such as oxygen (O2), nitrate (), sulfate (), and metal oxides (Fe(III), Mn(IV)). The decomposition of phytoplankton biomass also releases significant amounts of methyl-sulfur compounds, favoring methylotrophic CH4 production (Penger et al., 2012; Tebbe et al., 2023; Tsola et al., 2021; Yan et al., 2017; Zhou et al., 2022). Anaerobic conditions, combined with increased organic matter availability, are expected to boost methanogenesis, resulting in elevated CH4 release following eutrophication (Fiskal et al., 2019; Sanches et al., 2019; Zhou et al., 2022).
Conversely, nutrient addition can stimulate microbial CH4 oxidation (Yang et al., 2019). Some aerobic MOB can oxidize CH4 while respiring (denitrification), and a growing body of evidence supports the widespread occurrence and activity of these bacteria in O2-limited environments (Almog et al., 2024; Kits et al., 2015; Reis et al., 2024; Schorn et al., 2024). Importantly, MOB exhibit niche partitioning along O2–CH4 and nutrients gradients with Gammaproteobacteria, Alphaproteobacteria, and nitrite-dependent taxa that produce O2 intracellularly, such as Candidatus Methylomirabilis, occupying distinct layers (Mayr et al., 2020; Reis et al., 2020). Gammaproteobacterial MOB are generally associated with fast-growing life strategies in resource-rich conditions, whereas alphaproteobacterial MOB are adapted to resource-limited or stable environments (Ho et al., 2013). Indeed, P and N enrichment, for instance, can increase CH4 oxidation rates and favor Gammaproteobacterial over Alphaproteobacterial MOB (Nijman et al., 2022; Veraart et al., 2015). Together, these findings highlight that nutrients modulate both CH4 production and consumption, adding complexity to how eutrophication shapes lacustrine CH4 dynamics (Nijman et al., 2022; Reis et al., 2020; Veraart et al., 2015; Wei et al., 2022).
Eutrophication of lakes in Switzerland reached critical levels during the mid-20th century, particularly in the 1950–1970s, due to rapid industrialization, urbanization, and agricultural intensification. Public outcry and scientific research prompted the introduction of wastewater treatment plants and stricter regulations on phosphate detergents, leading to significant improvements in water quality by the late 20th century. Nevertheless, sediments retain a legacy of this eutrophication in the form of increased organic matter content (Fiskal et al., 2019), which continues to shape microbial community structure (Han, 2020) long after lake waters recovered. Some studies have reported clear vertical zonation of methanogenic and methanotrophic communities in relation to trophic history and electron acceptor distributions (Rissanen et al., 2023; Van Grinsven et al., 2022), while others found little to no stratification (Meier et al., 2024). Consequently, the cascading impacts of anthropogenic nutrient inputs on the balance of CH4 production and oxidation in lake sediments remain poorly constrained.
Here, we test whether historical eutrophication has left a sedimentary legacy that structures contemporary methane-cycling communities in Lake Joux (Vaud, Switzerland), a site with a well-documented history of trophic regime shifts and phytoplankton bloom deposits (Lavrieux et al., 2017; Monchamp et al., 2021). Using a ∼ 400 year, 55 cm sediment archive, we combine porewater and solid-phase geochemistry, 16S rRNA gene amplicon profiling, and stable-carbon-isotope measurements to resolve depth zonation of methanogens and methanotrophs, and link community shifts to organic-matter sources, lithology, and redox conditions. By explicitly coupling the depositional record to present microbial community structure and isotope compositions of different carbon pools, we provide process-level constraints relevant for forecasting benthic CH4 in eutrophying and recovering lakes.
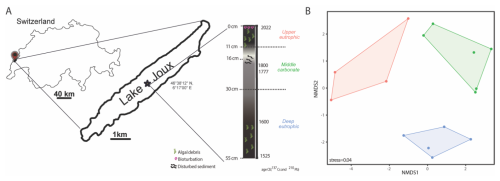
Figure 1(A) Location of Lake Joux (Switzerland; sampling in 2023) and 55 cm core schematic representation with the three stratigraphic intervals identified from lithology and age markers: deep eutrophic, middle carbonate, and upper eutrophic. (B) Non-metric multidimensional scaling (NMDS) of the porewater and solid-phase geochemical dataset, showing separation of samples by stratigraphic interval (upper eutrophic = red, middle carbonate = green, deep eutrophic = blue). Points are individual depth samples; colored polygons outline the convex hull for each interval and symbols mark group centroids. Ordination was performed on z-scored variables using Bray–Curtis dissimilarities.
2.1 Study Area
Lake Joux is a perialpine lake in the Joux Valley in the Swiss Jura Mountains (Fig. 1A). The valley developed in a Jura syncline, marked by glacial erosion and Quaternary deposits, and it lies mainly on Upper Jurassic and Tertiary limestones. The lake has an average depth of 32 m, a surface area of approximately 9 km2 (maximum 9 km in length and 1 km in width), and a watershed covering around 211 km2. Situated at an altitude of 1183 m, the lake is subject to intense seasonal variations and meteorological events, which drive the runoff of both natural and anthropogenic materials.
Human activity in the watershed dates back over 6850 years (Lavrieux et al., 2017; Mitchell et al., 2001; Monchamp et al., 2021). By the 16th century, the area around Lake Joux became more densely populated, leading to land drainage and deforestation for livestock farming (Piguet, 1946). Frequent crop failures and food shortages during the late 17th century spurred the growth of glassmaking and lapidary industries. Horology, introduced in the 18th century, became the region's dominant economic activity by the 19th century. This period of industrialization resulted in a transition from cultivated farmland to pastures and fallow fields (Lavrieux et al., 2017).
Agricultural intensification and urban expansion during the 20th century significantly increased nutrient inputs to Lake Joux, resulting in pronounced eutrophication. Phosphorus levels peaked at 35 µg L−1 in 1979 (Lods-Crozet et al., 2006), triggering major ecological changes, including rapid shifts in phytoplankton communities as eutrophication-adapted taxa outcompeted the lake's original species (Monchamp et al., 2021). A re-oligotrophication phase began in 1988–1989 following improved nutrient management and mitigation efforts. However, despite these reductions in external nutrient loading, the lake has not returned to its pre-eutrophication conditions. More than 70 years after the documented episode of eutrophication, the water column remains altered, suggesting that the system has shifted to an alternative stable state with a biological configuration resistant to reversal (Lods-Crozet et al., 2006; Monchamp et al., 2021).
2.2 Sampling
In May 2023, three gravity cores (45–55 cm long) were recovered from the lakebed of Lake Joux using a Uwitec gravity corer. The cores were taken from one of the deepest parts of the lake (46°38′12′′ N, 6°17′00′′ E; 28 m depth) and sealed with rubber caps. One core was pre-drilled and taped at 3 cm intervals to facilitate rapid CH4 sampling using cut-off 3 mL syringes on shore. The other two cores, one for porewater and the other for sediment chemistry and microbiome analyses, were processed in the laboratory within 24 h. Porewater was extracted via N2 flushed syringes attached to 0.2 µm Rhizons (Rhizosphere), inserted every 3 cm along the core, stored at 4 °C and analyzed within 48 h. The third sediment core was opened with a handheld saw and sectioned every 3 cm. Each solid sample was split into an acid-cleaned vial and a sterile vial, then frozen at −20 °C.
2.3 Porewater chemistry
Porewater samples for dissolved anions (, , , ) were transferred to plastic vials while flushing with N2, capped, and analyzed using an ion chromatograph (DX-ICS-1000, DIONEX) equipped with an AS11-HC column. For dissolved inorganic carbon (DIC) porewater samples were filled into 1.5 mL borosilicate vials and capped headspace-free to prevent CO2 degassing. DIC concentration (mmol L−1) was obtained from the CO2 yield after acid conversion of aliquots transferred to helium-flushed Exetainers containing 200 µL of 99 % H3PO4, and the resulting CO2 peak areas were quantified on a GasBench II (Thermo Fisher Scientific). A response factor (µmol CO2 per peak-area unit) was derived from identically processed Carrara Marble (CM) standards and applied to the second CO2 peak of each sample; moles of CO2 were converted to DIC using the injected sample volume. External uncertainty on DIC concentration, based on CM reproducibility, was < 7 %. For δ13CDIC, the headspace CO2 was analyzed by GasBench II coupled to a Delta V Plus IRMS; each sample was measured 6 times, and we report the mean with 1σ (typically < 0.10 ‰). Values were normalized to the in-house CM standard (δ13C = +2.05 ‰) calibrated against NBS-19 and NBS-18; external reproducibility from CM replicates (n = 12) was < 0.05 ‰ (1σ). Carbon isotopes are reported in delta (δ) notation relative to the Vienna Pee Dee Belemnite (VPDB) standard, defined as × 1000 ‰, with R the ratio; positive δ indicates enrichment in 13C relative to Vienna Pee Dee Belemnite (VPDB) standard. For dissolved sulfide analysis, porewater was fixed with 0.05 M Zn-acetate (Zn(CH3COO)2 ⋅ 2H2O) solution at a 1:2 ratio immediately after extraction, and dissolved sulfide was quantified photometrically using the methylene blue method (Cline, 1969).
2.4 Sediment description and chemistry
Sediment from the opened core was visually assessed (using standard charts) for color and granulometry based on observable differences in particle size, texture, and sorting within the sediment layers. Lithological boundaries in our core were aligned to the dated Lake Joux record of Lavrieux et al. (2017) using their carbonate sediment interval (whiter sediments) as reference. Ages were transferred from their model; uncertainties are those reported therein.
For total phosphorus (P), ∼ 1 g of wet sediment was digested in 9 mL of 4:2 HNO3:HCl using an Anton Paar microwave system, filtered (0.45 µm glass fiber), and analyzed by inductive coupled plasma–optical emission spectrometry (ICP-OES, Agilent 5800). Calibration used a multi-element standard, with certified reference materials yielding 85 %–102 % recovery.
Elemental C, N, H, and S were measured on 1–3 mg of freeze-dried sediment using a UNICUBE (Elementar®) at EPFL's ISIC-MSEAP. Total organic carbon (TOC) and total inorganic carbon (TIC) were estimated by loss on ignition (500 and 1200 °C). δ13Corg was determined by Elemental Analyzer–Isotope Ratio Mass Spectrometry (EA-Isolink IRMS, Thermo Fisher) after 48 h treatment with 6 N HCl to remove carbonates. Results are reported in delta notation related to VPDB, as described above for δ13CDIC), with a reproducibility better than 0.2 ‰.
Acid-volatile sulfur (AVS) and chromium-reducible sulfur (CRS) were extracted from 1–2 g of frozen sediment as per Spangenberg and Bosco-Santos (2024). Sulfide in AVS and CRS fractions was measured colorimetrically (Cline, 1969) and CRS sulfur isotopic composition (δ34Scrs) by IRMS (Spangenberg and Bosco-Santos, 2024). These measurements help distinguish easily mobilized sulfide pools (AVS) from more stable sulfur forms (CRS) in sediments.
2.5 Dissolved oxygen and methane
Oxygen concentrations were measured using a 200 µm-tip glass microsensor (Unisense) after 2-point calibration in Na-dithionite and air-saturated water. Seven vertical profiles from the same core were recorded at 250 µm steps with a motorized controller and Field Multimeter (Unisense).
For CH4 analysis, 3 cm3 of sediment was transferred into 100 mL serum bottles with 5 mL of 10 % NaOH, sealed, and homogenized. Dissolved CH4 was extracted by headspace displacement and quantified via gas chromatography (Joint Analytical Systems) equipped with an FID at the Eawag (Khatun et al., 2024). was measured using Gas Chromatography–Combustion–Isotope Ratio Mass Spectrometry (GCC-IRMS, Agilent 6890N with Thermo Finnigan IRMS) and analyzed with IonVantage software (Khatun et al., 2024). Results are reported in delta notation relative to VPDB with an analytical error < 1.1 ‰.
Carbon isotopic fractionation factors (α) between organic carbon (δ13Corg, substrate) and methane (, product, were calculated as: ). The corresponding isotopic fractionation (ε, ‰) was then determined by the relationship , allowing interpretation of trends in dominant methanogenic pathways.
In order to determine sediment zonation by environmental variables, we performed non-metric multidimensional scaling (NMDS) on a Euclidean distance matrix of z-scored environmental data for samples between 0.5 and 43.5 cm sediment depth, using the function metaMDS() in the R package vegan. The NMDS stress value was 0.04. Differences among depth-defined clusters based on environmental variables were tested using a permutational multivariate analysis of variance (PERMANOVA) on smoothed Euclidean distance matrices.
2.6 DNA extraction and 16S rRNA gene amplicon analysis
DNA was extracted from Lake Joux sediments using the PowerSoil Pro Kit (Qiagen). Extraction, sequencing, and raw data processing were conducted at the Joint Microbiome Facility (Medical University of Vienna and University of Vienna; project ID JMF-2310-14). The V4 hypervariable region of the 16S rRNA gene was amplified and sequenced to assess the total microbial diversity in the collected samples. Amplification was performed with linker-modified 515F and 806R (Apprill et al., 2015; Parada et al., 2016) primers, and amplicons were barcoded, multiplexed, sequenced on an Illumina MiSeq (v3 chemistry, 2x 300 bp), and extracted from the raw sequencing data as described in detail in Pjevac et al. (2021). Amplicon Sequence Variants (ASVs) were inferred using the DADA2 R package v1.42 (Callahan et al., 2016b), applying the recommended workflow (Callahan et al., 2016a). FASTQ reads 1 and 2 were trimmed at 220 and 150 nt with allowed expected errors of 2. ASV sequences were subsequently classified using DADA2 and the SILVA database SSU Ref NR 99 release 138.1 (Quast et al., 2012; Mclaren and Callahan, 2021) using a confidence threshold of 0.5. ASVs without classification or classified as eukaryotes, mitochondria, or chloroplasts, as well as well-known buffer contaminations, were removed. After filtering, only samples with at least 7000 read pairs were kept for further analyses, and relative abundances of ASVs grouped at higher taxonomic levels were calculated in relation to all remaining data. The relative abundance of chloroplast sequences, which were removed from the microbial community dataset, was examined separately to assess phytoplankton debris abundance across the sediment profile.
Downstream analyses were performed using R v4.3.2 and Bioconductor v3.16 packages SummarizedExperiment v1.32, SingleCellExperiment v1.24, TreeSummarizedExperiment v2.8 (Huang et al., 2021), mia v1.8 (https://github.com/microbiome/mia), LMdist (Hoops and Knights, 2023), vegan 2.6-8, phyloseq v1.44 (Mcmurdie and Holmes, 2013) (Vegan R package; phyloseq R package), microbiome v1.22 (http://microbiome.github.io, last access: 2 February 2026), microViz v0.10.8 (Barnett et al., 2021), and corrplot (Wei and Simko, 2024). Microbial community alpha diversity indices were calculated on rarified 16S rRNA gene amplicon data using R packages vegan and mia. For community dissimilarity analysis, microbial 16S rRNA gene amplicon sequence count data was centered log ratio (CLR) transformed, a pairwise Aitchison distances matrix was computed, and oversaturated distances in the dissimilarity matrix were corrected and smoothed using LMdist with default settings prior to ordination using principal coordinates analysis (PCoA). Differences in 16S rRNA gene amplicon community composition among three depth-defined clusters were tested using permutational multivariate analysis of variance (PERMANOVA) on an Aitchison distance-based dissimilarity matrix.
To identify the environmental variables that significantly contributed to the variation in microbial community structure, correlations between microbial community composition and environmental variables were assessed using Mantel tests, based on Euclidean distances calculated from Z-score standardized environmental variables and LMdist corrected and smoothed Aitchison distances of 16S rRNA gene amplicon sequencing data. Prior to correlation analysis, five samples from the deep eutrophic layer without corresponding environmental data were excluded. Mantel tests were performed using Spearman's rank correlation as implemented in the R package vegan. The resulting p values were adjusted for multiple testing using the false discovery rate (FDR) method. Highly correlated environmental variables (Spearman's r > 0.8, Fig. S1 in the Supplement), as assessed by the function cor() in the R package corrplot, were removed before the Mantel tests.
3.1 Sediment description
The 55 cm deep sediment record of Lake Joux could be classified into three main intervals based on distinct lithological and chemical features (Figs. 1 and S1 in the Supplement). The “deep eutrophic” interval, from 55 to 30 cm, comprises black and silty sediments, indicating a period of higher lake productivity and low oxygen conditions. Occasional fine sand and organic fibers are also present. In the “middle carbonate” interval, from 30 to 11 cm, the sediments transition from a murky gray with heterogeneous brownish features, suggesting changes in organic matter quality and oxidation states (Fig. 1) to whitish silty-sandy sediments, with the contribution of shells above 13.5 cm, indicating the dominant deposition of carbonates (Fig. 1). The “upper eutrophic” interval, from 11 to 0 cm, contains intensely black sediment with frequent plant debris, reflecting recent environmental changes.
To assign approximate ages to our sedimentary profile, we correlated our lithological intervals to other Lake Joux sedimentary sequences previously published and dated using 237Cs and 210Pb (Lavrieux et al., 2017; Magny et al., 2008) (Fig. 1). The middle carbonate unit (30–11 cm) in our core aligns with their U3–U4 carbonate interval, including the distinctive pale “white” boundary at ∼ 16–11 cm. The overlying upper eutrophic black, organic-rich sediments (11–0 cm) correspond to U5, which spans the 20th-century eutrophication phase and includes the 137Cs markers at ∼ 1954 and ∼ 1963 in the Lavrieux record. By transfer of their age–depth model, the base of our deep eutrophic interval (below 30 cm) falls in the late 16th–early 17th century. Reported sedimentation rates in Lavrieux (≈ 0.04–0.11 cm yr−1 before the 18th century, a short-lived peak ≈ 0.83 cm yr−1 in the late 18th century, and ≈ 0.18 cm yr−1 over recent decades) are consistent with the thicknesses of our corresponding units (Lavrieux et al., 2017).
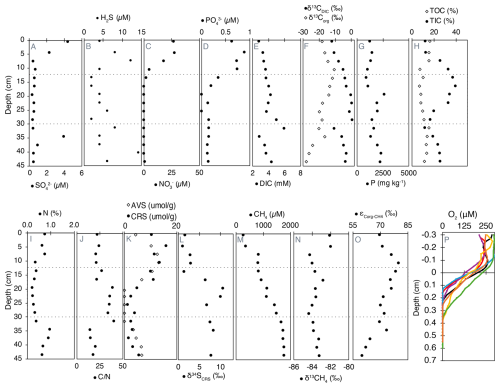
Figure 2Geochemical profiles of porewater, solid-phase compounds, and dissolved gases in Lake Joux sediments. Dashed lines represent the transitions between the “deep eutrophic” interval from 55 to 30.5 cm, the “middle carbonate” interval from 28.5 to 11 cm, and the “upper eutrophic” from 11 to 0 cm. All data are available in Tables S1 (panels A to N) and S2 (panel O) in the Supplement.
3.2 Porewater chemistry
Sulfate (, between 0.35 and 4.5 µM) and dissolved sulfide (H2S, between 2 and 14.3 µM) were measurable throughout the entire sedimentary profile. In the upper eutrophic interval (0–11 cm), opposing gradients of and H2S from the surface to 7.5 cm are evidence of sulfate reduction (Fig. 2A and B). Below this depth, sulfate is absent, but a broad H2S maximum in the deep eutrophic sediments (∼ 40 cm) could be associated with organic sulfur degradation.
Dissolved nitrate () and phosphate () first appeared at 19.5 and 16.5 cm, respectively, with concentrations progressively increasing toward the surface, reaching maximum values of 27 µM for and 0.62 µM for (Fig. 2C and D). Nitrite concentrations were close to the detection limit (0.03 µM) and uniformly low (range 0.038–0.087 µM, median ≈ 0.043 µM) with no systematic depth trend (Fig. S2 in the Supplement).
Dissolved inorganic carbon (DIC) concentrations with isotopically heavier composition (δ13CDIC) were highest in the middle carbonate interval between 16.5 and 31.5 cm (Fig. 2E and F). Above 7.5 cm depth, δ13CDIC values became progressively lighter toward the surface, reaching a minimum of −12 ‰ (Fig. 2F).
3.3 Sediment chemistry
3.3.1 Phosphorous and organic matter characterization
Total phosphorus (P) content, ranging from 860 to 2612 mg kg−1, was generally higher in the deeper sediments and progressively decreased towards the surface, except for a sharp peak at 19.5 cm depth (Fig. 2G). Organic carbon (TOC) exhibited higher concentrations in both the deep eutrophic and upper eutrophic sediments, contrasting with TIC content, which peaked in the middle carbonate interval (Fig. 2H). The δ13Corg was lightest in the deep eutrophic sediments (−28.22 ‰ at 43.5 cm) and heaviest in the upper eutrophic sediments (−10.76 ‰ at 10.5 cm) (Fig. 2F).
Nitrogen content followed the same pattern as TOC, with higher N in the deep and upper eutrophic sediments compared to the middle carbonate region (Fig. 2I). The ratio between C and N, a qualitative parameter of organic matter source (Meyers, 1994), exhibited relatively lower values in the deep eutrophic sediments, increasing in the middle carbonate sediments and decreasing again in the upper eutrophic sediments (Fig. 2J).
3.3.2 Solid-phase sulfides
Acid volatile sulfides (AVS) were measurable in the deep eutrophic sediments between 43.5 and 34.5 cm and within the upper eutrophic sediments above 19.5 cm depth. The maximum concentrations of AVS in the upper eutrophic sediments (around 418 µg kg−1 at 10.5 cm) were about twice as high as in deep eutrophic sediments (200 µg kg−1 at 40.5 cm) (Fig. 2K). Chromium reducible sulfur (CRS) also exhibited higher concentrations in the shallower sediments, becoming more prominent from 16.5 cm depth to the surface. CRS concentrations were more variable than AVS, varying from 2 to 510 µg kg−1 (Fig. 2K). The isotopic composition δ34S of CRS was positive throughout the profile, ranging from ∼ 1 ‰ near the surface to a maximum of 10.5 ‰ at 19.5 cm depth. Values remained elevated in the middle carbonate and deep eutrophic zones (e.g., 8.3 ‰ at 34.5 cm and 7.7 ‰ at 43.5 cm), indicating that the reduced sulfur pool is isotopically enriched in 34S across the sediment column (Fig. 2L).
3.3.3 Dissolved oxygen and methane
Methane (CH4) concentrations were highest in the deep eutrophic sediments, with a maximum of approximately 1760 µM at 45 cm depth. From 31.5 cm depth, CH4 exhibited a clear decreasing trend, reaching the lowest concentration (253 µM) at the surface (Fig. 2M). The most significant drop in CH4 concentrations occurred between 7.5 and 4.5 cm depth, where the concentrations decreased by half (Fig. 2M). The exhibited minimal variation along the profile, averaging −83.0 ± 0.7 ‰. The most pronounced isotopic shift (> 2.2 ‰) towards heavier values occurred at the same depth as the sharp decline in CH4 concentration (Fig. 2N).
Fractionation factors (α) between Corg and CH4 ranged from 1.069 to 1.080 across sediment depths, corresponding to carbon isotope fractionations (ε) of 69 ‰ to 80 ‰ (Fig. 2O). These values reflect the measurable discrimination between 13C and 12C during CH4 production from organic substrates, which arises from the enzymatic pathways and substrates utilized. Lower ε values were consistently observed in deeper sediments compared to shallower layers.
Oxygen concentrations were measured across seven different profiles, and free O2 was detectable only in the uppermost sediments, between 0.165 and 0.365 cm depth (Fig. 2P). Below 0.4 cm, sediments were consistently anoxic. The heterogeneous penetration of O2 into the sediments is attributed to bioturbation, which was confirmed by visual observations of worm castings.
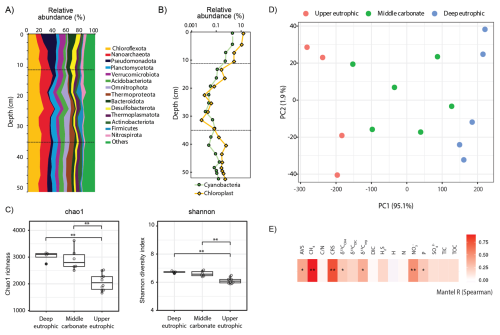
Figure 3(A) Relative 16S rRNA gene amplicon sequence abundances of bacterial and archaeal phyla and (B) of cyanobacteria and chloroplasts (algae and plants) in the 55 cm sedimentary profile of Lake Joux; (C) Alpha diversity (chao1 richness and Shannon diversity) of methanogens and methanotrophs. (D) Principal component ordination of centered log ratio (CLR) transformed 16S rRNA gene amplicon data, based on an Aitchison distance for which oversaturated distances were corrected and smoothed using LMdist. (E) Mantel tests results (Spearman's rank correlation) of community dissimilarity (corrected and smoothed Aitchison distance) and environmental parameters (z-scored). P values were adjusted for multiple testing using the false discovery rate (FDR) method. p < = 0.01; ∗ p < = 0.05.
3.3.4 Microbial community composition and chloroplast relative sequence abundances
The microbial community in Lake Joux sediments was dominated by the phyla Chloroflexota, Nanoarchaeota, and Pseudomonadota (Fig. 3A). In the upper eutrophic sediments (below 30 cm), microbial species richness and evenness (Chao1 and Shannon alpha diversity indices) were significantly lower than in overlying layers (Fig. 3C). In this zone, Nanoarchaeota reached their highest relative sequence abundances (> 10 %), decreasing to ∼ 7 % in the shallower sediments. These elevated abundances, also reported in freshwater (Chen et al., 2023; Xie et al., 2024) and marine environments (Brick et al., 2025), likely reflect their wide environmental tolerance and host associations (Jarett et al., 2018) (Fig. 3A).
In the middle carbonate-rich interval (30–11 cm), microbial diversity increased, and Bacteroidota appeared, consistently representing > 5 % of the microbial community. Reduced relative abundances of chloroplast sequences in this layer (Fig. 3B) also indicate limited input from photosynthetic organisms during this depositional phase. Cyanobacteria-related ASVs displayed similar depth trends to chloroplast sequences, but with lower overall abundance, reaching a maximum of 2 % at 4.5 cm depth (Fig. 3B). In the upper eutrophic sediments (11–0 cm), Pseudomonadota became more abundant (> 20 %) and chloroplast sequences markedly increased, reflecting enhanced sedimentation of photosynthetic organisms.
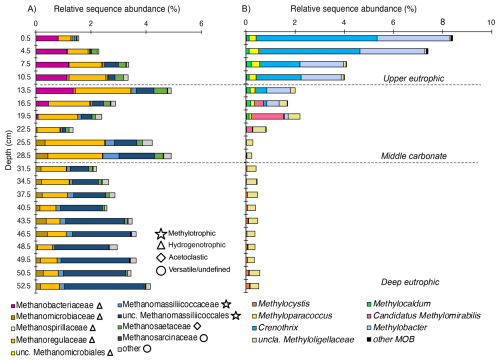
Figure 4Depth-resolved composition of methanotrophic and methanogenic taxa in Lake Joux sediments. (A) Methanogenic archaea clustered by family/order (relative sequence abundance of total community), and grouped by inferred pathways (methylotrophic, hydrogenotrophic, acetoclastic, versatile/undefined). (B) Methanotrophic bacteria, including canonical MOB and Candidatus Methylomirabilis (NC10), were expressed as relative sequence abundance of the total community.
Microbial community composition was more similar within sedimentary intervals than between them (Fig. 3D). The separation of samples into three depth clusters based on sediment biogeochemistry was statistically supported (PERMANOVA: F = 25.05, R2 = 0.81, p < 0.001, Fig. 3D). Significant differences in microbial community composition between the three depth clusters were also observed (PERMANOVA: methanogens – F = 10.57, R2 = 0.64, p < 0.001; MOB – F = 13.32, R2 = 0.69, p < 0.001; Cyanobacteria – F = 6.57, R2 = 0.52, p < 0.001; Chloroplasts – F = 8.4, R2 = 0.58, p < 0.001; Fig. S4 in the Supplement). Stratification was especially pronounced for cyanobacterial and chloroplast sequences, which formed three distinct depth-specific clusters corresponding to the eutrophic, carbonate, and deep eutrophic intervals (Fig. S4 in the Supplement). The methanotrophic community separated into two main groups, upper and deep eutrophic, while samples from the carbonate layer did not form a distinct cluster (Fig. S4 in the Supplement). Methanogens, however, displayed clearer depth partitioning, with methylotrophic Methanomassiliicoccales dominating in the deep eutrophic interval and hydrogenotrophic Methanobacteriales increasing toward the surface (Fig. 4). Notably, depth patterns in methanogens and methanotrophs, as well as cyanobacterial and chloroplast-related sequences, tracked the same environmental gradients, with CH4, , and CRS showing the strongest correlations and AVS, δ13Corg, and sedimentary P exhibiting secondary correlations (Figs. 3E and S3 in the Supplement).
3.3.5 Methanogenic and methanotrophic microbial communities
The relative sequence abundance of methanogens consistently accounted for more than 1 % of the microbial community across all sampled depths (Fig. 4A). Methanomassiliicoccales, which are H2 dependent methylotrophs, were the dominant methanogenic group in the deep eutrophic sediments, accounting for 1.4 % of the microbial community at a depth of 37.5 cm (Fig. 4A). In contrast, Methanomicrobiales (hydrogenotrophs) was the most abundant methanogen group in the middle carbonate interval, (reaching 2.1 % of the microbial community) while Methanobacteriales (hydrogenotrophs) sequences were most abundant in the upper eutrophic sediments (11–0 cm, reaching 1.18 % of the microbial community, Fig. 4A). Sequences affiliated with Methanosarciniales (metabolic versatile) were rare throughout the profile (< 0.01 %).
Across samples, the relative sequence abundances of Crenothrix and Methylobacter exhibited significant positive correlations with porewater and (Spearman, p < 0.05; Fig. 5). By contrast, methanogen families tracked depth-defined intervals rather than these nutrients (Fig. 4). In the deep and middle sediments below 19.5 cm, Rhizobiales-affiliated methylotrophs (e.g., Methylocystis, Methylocapsa, Methyloligellaceae) (Tamas et al., 2014; Vekeman et al., 2016) were the dominant putative methanotrophs, although they represented a modest portion of the community (max. 0.6 %) (Fig. 4B). Anaerobic methanotrophs from ANME archaeal groups could not be identified in the sedimentary profile of Lake Joux. Still, between 23 and 16 cm, Methylomirabilota NC10 bacteria capable of nitrite-dependent methane oxidation with intracellularly produced O2 under anoxic conditions (Ettwig et al., 2010) were detected at a relative abundance of 0.2 %–1.3 %.
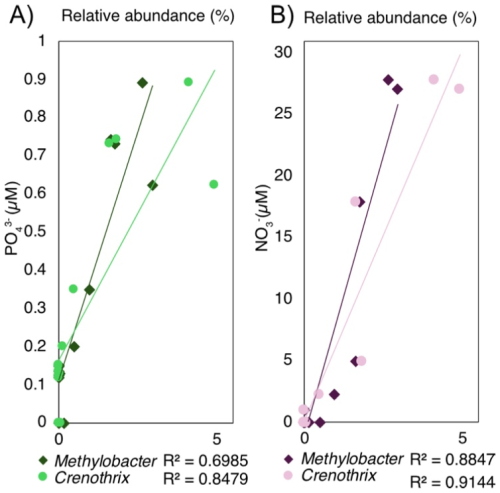
Figure 5(A) Correlation between concentration and the relative sequence abundance of Crenothrix and Methylobacter. (B) Correlation between concentration and the relative sequence abundance of Crenothrix and Methylobacter.
The 16S rRNA gene sequences of aerobic MOBs represented between 0.3 % and 8.7 % of the microbial community throughout the sediment profile and were especially numerous (> 1 %) above a depth of 19.5 cm (Fig. 3C). The most abundant methanotrophs from 19.5 cm depth to the surface were members of the order Methylococcales, with two genera prevalent near the surface: Crenothrix and Methylobacter (Fig. 4B). At the resolution available with V4-region 16S rRNA gene amplicon analyses, limited within-group diversity is detected, as four ASVs affiliated with Methylobacter, 17 ASVs with Crenothrix, and a small number of ASVs assigned to other Methylococcales genera, including a single abundant Methylomonas ASV (Fig. S1 in the Supplement), were recovered. Within the MOB community, the fraction of the Methylomonas ASV increased with depth (≈ 30 % to ≈ 70 %), whereas the fraction of the three most abundant Crenothrix ASVs remained stable. The abundance of all individual MOB ASVs decreased with depth, relative to the total community (Fig. S1 in the Supplement).
4.1 Tracing historical land use, industrialization, and eutrophication
The intensely black sediments abundant in chloroplast-related sequences and elevated TOC content with low ratios and light δ13Corg in the deep eutrophic interval (55–30 cm, Fig. 1) denote predominantly autochthonous organic matter, derived from phytoplankton blooms (Lamb et al., 2006; Morales-Williams et al., 2017). Similar patterns recorded in other Lake Joux sediment profiles (Dubois, 2016; Lavrieux et al., 2017; Magny et al., 2008) correspond to a period of intensified deforestation and settlement expansion between 1525 and 1790 CE (Dubois, 2016; Lavrieux et al., 2017; Magny et al., 2008). While no official records confirm eutrophication during this period, these anthropogenic activities likely led to increased erosion and sediment/nutrient transport, stimulating phytoplankton productivity (Fig. 1). Notably, these sediments also exhibit high relative H2S and AVS concentrations (Fig. 2B and K), which are characteristic of late sediment diagenesis under eutrophic depositional conditions (Holmer and Storkholm, 2001).
The middle carbonate layer (30–11 cm depth) reflects a shift towards more oligotrophic conditions, likely linked to the abandonment of land-intensive activities and the switch to manufacturing in the 18th century. In addition, the 1777 construction of a dike between Lake Joux and Lake Brenet lowered the lake level by 3.6 m, mobilizing limestone-rich sediments (high TIC) and terrestrial plant material with heavier δ13Corg and higher ratios (Lavrieux et al., 2017; Magny et al., 2008; Monchamp et al., 2021) (Fig. 2). The sedimentological transition that marks the beginning of this interval at 30 cm depth aligns with a shift from methylotrophic to hydrogenotrophic methanogens (Fig. 4), likely responsible for the lighter δ13CDIC values (Fig. 2F). Furthermore, the white-colored boundary of the middle carbonate layer (16–11 cm) coincides with warmer post-Little Ice Age conditions, promoting calcium carbonate precipitation and TIC enrichment (Lavrieux et al., 2017).
The upper eutrophic sedimentary interval (11–0 cm) consists of black sediments rich in TOC, lighter δ13Corg values, low ratios, and abundant chloroplast- and cyanobacteria-related sequences reflecting a well-documented 20th century eutrophication phase (Lavrieux et al., 2017; Magny et al., 2008; Monchamp et al., 2021) (Figs. 2 and 3B). Elevated nutrient levels in this interval could result from external nutrient inputs trapped in porewater or from organic matter remineralization. Porewaters are strongly reducing, reflected by elevated H2S and CRS, and the absence of O2 below 0.5 cm. Downward diffusing meets upward-diffusing CH4, and in the 7.5–0 cm horizon, CH4 concentrations fall sharply while becomes heavier and δ13CDIC lighter (Fig. 2A, B and K, M). Aerobic methanotrophy fractionates carbon, preferentially consuming 12CH4 while leaving behind heavier CH4 (enriched in 13C) and can produce lighter (relatively less 13C) co-localized DIC depending on mixing and available electron acceptors. Together with the dominance of MOB and the absence of ANME-related 16S rRNA gene amplicon sequences, the paired isotopic shifts observed indicate methanotrophy dominated by MOB as the main CH4 sink in these anoxic, nutrient-replete surface sediments.
4.2 Methylotrophic methanogens selected by past eutrophication
Changes in organic matter sources to Lake Joux over the last four centuries appear closely tied to shifts in dominant methanogenic groups within its sediments. Deep eutrophic sediments, characterized by the highest CH4 concentrations, are dominated by Methanomassiliicoccales, which are hydrogen-dependent methylotrophic methanogens, meaning that they use H2 as the electron donor and methylated one-carbon compounds (e.g., methanol, methylamines, methylated S compounds) as electron acceptors, rather than reducing CO2 (Bueno De Mesquita et al., 2023; Ellenbogen et al., 2024; Söllinger and Urich, 2019; Sun et al., 2019; Wang and Lee, 1994). The decomposition of algal and cyanobacterial biomass can release methylated sulfur compounds (including DMS and dimethylsulfoxide) and methylated amines, which have stimulated methylotrophic methanogenesis in laboratory experiments and natural environments (Bose et al., 2008; Chistoserdova, 2011; Chistoserdova et al., 2009; Huang et al., 2018; Singh et al., 2005; Tebbe et al., 2023; Whiticar, 1999; Zhou et al., 2022).
Indirect evidence for the presence of methylated sulfur compounds comes from relatively higher concentrations of H2S, AVS, and CRS at depth (Fig. 2B and K), indicating active sulfur cycling despite limited availability. Furthermore, δ34S values measured in CRS (primarily pyrite) consistently show positive isotopic signatures (7 ‰ to 10 ‰) in both the deep eutrophic and middle carbonate zones. While microbial reduction typically produces 34S-depleted sulfides (δ34S < 0 ‰) under open-system or moderately sulfate-limited conditions (Bradley et al., 2016; Canfield, 2001; Habicht and Canfield, 1997), the isotopic enrichment observed here is more consistent with either the degradation of sulfurized organic matter or methylated sulfur compounds (Phillips et al., 2022; Raven et al., 2019; Werne et al., 2004). These could simultaneously fuel methylotrophic methanogenesis and pyrite formation. This interpretation warrants confirmation through direct measurements of methylated sulfur species in future studies. Alternatively, the 34S enrichment could reflect near complete consumption of a limited pool so that 34S sulfide reflects positive values of the original sulfate (Bernasconi et al., 2017)(Fig. 2).
It is important to note that methylotroph distributions could also be influenced by competition for H2 with CO2-reducing hydrogenotrophs. In sulfate-poor anoxic sediments, H2 is typically buffered at low steady-state levels by continuous fermentative supply and rapid consumption – reflecting thermodynamic control rather than chronic scarcity (Conrad, 1999; Schütz et al., 1988; Kessler et al., 2019). Obligately methyl-reducing methanogens have very low H2 thresholds and are predicted to outcompete hydrogenotrophs for H2 when methyl groups are available. Thus, their activity is primarily considered to be limited by the availability of methylated substrates (Borrel et al., 2023; Bueno De Mesquita et al., 2023; Feldewert et al., 2020; Söllinger and Urich, 2019; Speth and Orphan, 2018). Given the dominance of Methanomassiliicoccales at depth, we infer that methylated-substrate supply rather than H2 limitation is the primary factor structuring the methanogenic community in the deep eutrophic interval. This interpretation is consistent with isotope patterns, as we have recorded comparatively heavier δ13CDIC in the deep eutrophic layer and a shift to lighter δ13CDIC above ∼ 30 cm where the relative abundance of CO2-reducing hydrogenotrophic methanogens increased (Fig. 2F).
Methylotrophic methanogenesis is typically a minor pathway in freshwater sediments because methylated substrates are scarce (Borrel et al., 2011; Bueno De Mesquita et al., 2023). However, in the deep eutrophic layer, prolonged algal biomass degradation likely generated a reservoir of recalcitrant methylated compounds (Achtnich et al., 1995; Rissanen et al., 2018), favoring methylotrophic methanogens. In contrast, hydrogenotrophic (using CO2 + H2) and acetoclastic (using acetate) methanogens primarily depend on fresh, labile organic matter, which rapidly becomes limited with burial (Achtnich et al., 1995; Meier et al., 2024; Rissanen et al., 2023; Rissanen et al., 2018). Thus, methylotrophs gain a selective advantage in these older, more refractory sediments. Above ∼ 28.5 cm, concurrent with a shift toward more terrestrial organic matter, methylotrophic methanogens decline and hydrogenotrophs progressively dominate (Fig. 4A). We interpret this pattern as consistent with a reduced supply of methylated substrates typically derived from algal organic matter although these compounds were not directly measured.
To further support the interpretation of distinct methanogenic pathways, we analyzed the , reflecting the isotopic discrimination during CH4 formation from Corg (Fig. 2O). Methanogenesis discriminates against the heavier 13C isotope (Conrad, 2005). In theory, when CH4 is produced from CO2 + H2 (hydrogenotrophy), microbes selectively withdraw 12C from the DIC pool, leaving residual DIC relatively 13C-enriched (less negative δ13CDIC); when methylotrophy dominates, CH4 carbon is drawn from methyl pools and δ13CDIC is affected less. We observed lower ε values in deeper eutrophic sediments compared to shallower zones. Although interpreting specific metabolic pathways from isotopic fractionation is challenging in mixed microbial communities, the contrast in (Fig. 2O) indicates distinct CH4-producing processes dominate at different sediment depths.
Our results support the view that eutrophication leaves a distinct imprint on methanogen stratification in sediments. In contrast to earlier studies that reported either weak vertical structuring (Meier et al., 2024) or only subtle shifts in methanogen dominance (Rissanen et al., 2023), we found clear zonation with Methanomassiliicoccales prevailing in the deepest eutrophic interval, Methanomicrobiaceae in the carbonate-rich middle section, and Methanobacteriaceae dominating the upper eutrophic sediments. Considered alongside these previous observations in other lakes, our findings question the usefulness of broad generalizations and suggest that methanogen communities are primarily shaped by habitat-specific conditions – such as lithology, organic-matter quality, and redox context – rather than exhibiting universal hydrogenotroph dominance. By comparison, a pronounced vertical structuring of methane-oxidizing bacteria appears more consistent across systems (Mayr et al., 2020; Rissanen et al., 2018; Van Grinsven et al., 2022).
4.2.1 Aerobic methanotrophs are selected by nutrient availability
Within anoxic lacustrine sediments, CH4 is typically oxidized anaerobically (Borrel et al., 2011; Martinez-Cruz et al., 2018). Interestingly, in the anoxic sediments of Lake Joux, anaerobic methanotrophic archaea are not detectable. Sequences related to Candidatus Methylomirabilis – capable of intracellular O2 production to fuel methane monooxygenase activity – occur in notable relative sequence abundances but are confined to 16–23 cm depth within the middle carbonate interval, but are relatively scarce compared to their aerobic counterparts. Namely, gammaproteobacterial MOB 16S rRNA gene sequences recovered from Lake Joux sediments are highly abundant (1 %–9 %) from 19.5 cm upward, despite prevailing anoxic conditions (Figs. 4 and 5). The MOB 16S rRNA gene sequences primarily affiliate with Methylococcales, notably the genera Crenothrix and Methylobacter (Fig. 5). The dominance of Methylococcales-associated MOB in the methane oxidation zone suggests that these taxa serve as the dominant CH4 sink in these nutrient-replete but anoxic surface sediments (Figs. 2N and 5). Aerobic (and denitrifying) methanotrophy also fractionates carbon, preferentially consuming 12CH4. This depletes residual CH4 in 12C while generating lighter (more 12C) co-localized DIC depending on mixing and available electron acceptors. The paired heavier δ13C and lighter δ13CDIC in the 7.5–0 cm horizon thus support the occurrence of active methane oxidation.
How these aerobic methanotrophs meet their O2 demand below 0.4 cm – where no O2 could be detected, remains unresolved. Nanomolar O2 concentrations cannot be excluded, but diffusive supply from the sediment–water interface to higher sediment depths is implausible. While some Methylococcales respire alternative electron acceptors (e.g., nitrate, Fe(III)) at low O2 levels (Li et al., 2023; Van Grinsven et al., 2020; Yang et al., 2025), methane monooxygenase remains O2-dependent for the oxidation of CH4 to methanol. Three microbial mechanisms could generate microscale O2 at depth within the sediment (“dark O2”): methanobactin-mediated water splitting (Dershwitz et al., 2021), chlorite () dismutation by (per)chlorate-respiring bacteria (Xu and Logan, 2003), and nitric-oxide dismutation as described for NC10 bacteria (Ettwig et al., 2010).
Water lysis has been proposed as a potential mechanism for O2 generation via methanobactin-mediated metal reduction, particularly under metal-rich conditions. (Dershwitz et al., 2021). This process has been primarily associated with Alphaproteobacterial methanotrophs, which are not prevalent in Lake Joux. Chlorite dismutation, catalyzed by chlorite dismutase found in over 60 genera across 13 phyla (Barnum and Coates, 2023), could be a source of O2, as Pseudomonadota and Actinobacteria are abundant in these sediments (Fig. 3A). However, environmental levels of (per)chlorate are likely too low to support this pathway at significant levels (Lv et al., 2019; Miller et al., 2014; Wang et al., 2024).
One potential source of in-situ O2 is nitric oxide dismutation catalyzed by the nitric oxide dismutase (NOD) enzyme, which has been recently attributed to multiple bacterial lineages, including several families within the phylum Bacteroidota (Ruff et al., 2024). In Lake Joux, putatively NOD-containing Bacteroidota account for ∼ 0.54 ± 0.2 % of the microbial community in the upper eutrophic sediments, suggesting this pathway may contribute to localized O2 production. However, as NOD is not encoded by all representatives of these taxa, we can not perform further reliable abundance estimates of NOD based on the available 16S rRNA gene amplicon data. The mechanism of O2 production is nevertheless consistent with our geochemical context: porewater remained near detection limit with no subsurface maximum (Fig. S2 in the Supplement), indicating rapid NO:x turnover typical of energy-limited sediments. Independent work shows some gammaproteobacterial methanotrophs, including Crenothrix and Methylobacter, possess genes for respiratory reduction (Almog et al., 2024; He et al., 2022; Martinez-Cruz et al., 2017; Milucka et al., 2012; Schorn et al., 2024). Active respiration has only been demonstrated experimentally for Methylomonas denitrificans cultures (Kits et al., 2015) and indirectly by denitrification gene expression by MOB in Lake Zug (Schorn et al., 2024). The latter study revealed that Crenothrix and Methylobacter related microorganisms continue CH4 oxidation in hypoxic and anoxic regions of the water column by performing denitrification or fermentation-based methanotrophy (Schorn et al., 2024). In Lake Joux, these same MOB taxa dominate the highly reducing, upper eutrophic sediments (Fig. 4B).
The positive covariance of Crenothrix and Methylobacter with and is also consistent with nutrient-stimulated MOB activity and/or a shared response to favorable near-surface conditions. While the source of surprisingly high concentrations cannot be resolved here, possible mechanisms of generation include oxidation of by Mn(IV) or Fe(III) oxides. Importantly, the MOB–nutrient correlations may also reflect a shared response to favorable near-surface conditions (e.g., sustained inputs of labile organic matter or higher porosity), rather than direct nutrient control. Nevertheless, it has been experimentally demonstrated that , , and additions can directly enhance CH4 oxidation rates by MOB and, in particular, Methylobacter (Almog et al., 2024; Kits et al., 2015; Nijman et al., 2022; Xia et al., 2021; Yang et al., 2025). Taken together, these observations suggest that nutrient availability may play a direct role in shaping the structure and activity of MOB communities.
Our results show that historical eutrophication left a lasting sedimentary legacy that structures contemporary methane-cycling microbial communities, selecting methylotrophic methanogens. In upper eutrophic, anoxic sediments, the surprisingly high relative sequence abundances of MOB (up to ∼ 9 %) specifically Methylococcales-affiliated Gammaproteobacteria, co-vary with elevated and concentrations. This suggests that eutrophication can simultaneously stimulate CH4 production and enhance its oxidation by shaping microbial assemblages.
As eutrophication continues to alter freshwater systems globally, understanding nutrient- and substrate-driven shifts in CH4-cycling communities becomes increasingly important. Future studies should focus on elucidating the in-situ activity of aerobic methanotrophs and molecular mechanism of methane oxidation under anoxic conditions, as presumably aerobic MOB have been widely reported in anoxic sediments (Almog et al., 2024; Ruff et al., 2024; Schorn et al., 2024). Combining molecular, isotopic, and geochemical approaches will be essential to better constrain methane fluxes in lakes undergoing or recovering from eutrophication.
All geochemical data are included in this published article and its Supplement. The 16S rRNA gene amplicon sequencing data are publicly available in the National Center for Biotechnology Information Sequence Read Archive (SRA) under BioProject accession PRJNA1207472 and can be accessed at: https://www.ncbi.nlm.nih.gov/bioproject/?term=PRJNA1207472 (last access: 24 April 2026).
The supplement related to this article is available online at https://doi.org/10.5194/bg-23-2909-2026-supplement.
ABS contributed to conceptualization, data curation, formal analysis, funding acquisition, methodology, supervision, validation, visualization, writing original draft preparation, review, and editing. ERBB contributed to data curation, formal analysis, and manuscript editing; SK contributed to data curation, formal analysis, and manuscript editing; MEM contributed to resources and manuscript editing; JS contributed to data curation, formal analysis, methodology, validation, visualization, designing and implementing computer codes, and writing original draft preparation, review, and editing. PP contributed to conceptualization, formal analysis, funding acquisition, supervision, validation, visualization, writing original draft preparation, review, and editing. JSB contributed to conceptualization, formal analysis, funding acquisition, supervision, validation, visualization, writing original draft preparation, review, and editing.
The contact author has declared that none of the authors has any competing interests.
Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims made in the text, published maps, institutional affiliations, or any other geographical representation in this paper. The authors bear the ultimate responsibility for providing appropriate place names. Views expressed in the text are those of the authors and do not necessarily reflect the views of the publisher.
We would like to thank two anonymous reviewers whose comments helped improve the quality of the manuscript. The authors would like to extend their gratitude to Floreana Marie Miesen (UNIL) for her invaluable support with fieldwork and logistics; to Carsten Schubert (EAWAG) for providing access to laboratory facilities for methane analyses; to Giulia Ceriotti (UNIL) for analytical and logistical support with methane analyses; to Laetitia Monbaron and Micaela Faria (UNIL) for their technical assistance in the laboratory; to Jorge Spangenberg (UNIL) for his support in methodology development and analytical assistance; to Frédéric Lardet for assistance with Fig. 1; and to William Leavitt (The University of Utah) for his help during fieldwork. Special thanks are also due to the Fondation Agassiz for the individual grant awarded to the first author, Alice Bosco-Santos, which made this study possible. The amplicon sequencing section of this work has been achieved using the Life Science Compute Cluster (LiSC) of the University of Vienna.
This work was financially supported by the 2022 and 2025 research grants from the Fondation Agassiz, awarded to Alice Bosco Santos.
This paper was edited by Lishan Ran and reviewed by two anonymous referees.
Achtnich, C., Bak, F., and Conrad, R.: Competition for electron donors among nitrate reducers, ferric iron reducers, sulfate reducers, and methanogens in anoxic paddy soil, Biol. Fertil. Soils, 19, 65–72, https://doi.org/10.1007/bf00336349, 1995.
Almog, G., Rubin-Blum, M., Murrell, J. C., Vigderovich, H., Eckert, W., Larke-Mejía, N., and Sivan, O.: Survival strategies of aerobic methanotrophs to hypoxia in methanogenic lake sediments, Environ. Microbiome, 19, 44, https://doi.org/10.21203/rs.3.rs-3790875/v1, 2024.
Apprill, A., McNally, S., Parsons, R., and Weber, L.: Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton, Aquat. Microb. Ecol., 75, 129–137, https://doi.org/10.3354/ame01753, 2015.
Barnett, D. J., Arts, I. C., and Penders, J.: microViz: an R package for microbiome data visualization and statistics, Journal of Open Source Software, 6, 3201, https://doi.org/10.21105/joss.03201, 2021.
Barnum, T. P. and Coates, J. D.: Chlorine redox chemistry is widespread in microbiology, ISME J., 17, 70–83, https://doi.org/10.1038/s41396-022-01317-5, 2023.
Bastviken, D., Cole, J., Pace, M., and Tranvik, L.: Methane emissions from lakes: Dependence of lake characteristics, two regional assessments, and a global estimate, Global Biogeochem. Cy., 18, https://doi.org/10.1029/2004gb002238, 2004.
Bastviken, D., Cole, J. J., Pace, M. L., and Van de Bogert, M. C.: Fates of methane from different lake habitats: Connecting whole-lake budgets and CH4 emissions, J. Geophys. Res.-Biogeo., 113, https://doi.org/10.1029/2007jg000608, 2008.
Bastviken, D., Tranvik, L. J., Downing, J. A., Crill, P. M., and Enrich-Prast, A.: Freshwater methane emissions offset the continental carbon sink, Science, 331, 50–50, https://doi.org/10.1126/science.1196808, 2011.
Beaulieu, J. J., DelSontro, T., and Downing, J. A.: Eutrophication will increase methane emissions from lakes and impoundments during the 21st century, Nat. Commun., 10, 1375, https://doi.org/10.1038/s41467-019-09100-5, 2019.
Bernasconi, S. M., Meier, I., Wohlwend, S., Brack, P., Hochuli, P. A., Bläsi, H., Wortmann, U. G., and Ramseyer, K.: An evaporite-based high-resolution sulfur isotope record of Late Permian and Triassic seawater sulfate, Geochim. Cosmochim. Ac., 204, 331–349, https://doi.org/10.1016/j.gca.2017.01.047, 2017.
Borrel, G., Jézéquel, D., Biderre-Petit, C., Morel-Desrosiers, N., Morel, J.-P., Peyret, P., Fonty, G., and Lehours, A.-C.: Production and consumption of methane in freshwater lake ecosystems, Res. Microbiol., 162, 832–847, https://doi.org/10.1016/j.resmic.2011.06.004, 2011.
Borrel, G., Fadhlaoui, K., Ben Hania, W., Gaci, N., Pehau-Arnaudet, G., Chaudhary, P. P., Vandekerckove, P., Ballet, N., Alric, M., and O'toole, P. W.: Methanomethylophilus alvi gen. nov., sp. nov., a Novel Hydrogenotrophic Methyl-Reducing Methanogenic Archaea of the Order Methanomassiliicoccales Isolated from the Human Gut and Proposal of the Novel Family Methanomethylophilaceae fam. nov, Microorganisms, 11, 2794, https://doi.org/10.3390/microorganisms11112794, 2023.
Bose, A., Pritchett, M. A., and Metcalf, W. W.: Genetic analysis of the methanol-and methylamine-specific methyltransferase 2 genes of Methanosarcina acetivorans C2A, J. Bacteriol., 190, 4017–4026, https://doi.org/10.1128/JB.00117-08, 2008.
Bradley, A. S., Leavitt, W. D., Schmidt, M., Knoll, A. H., Girguis, P. R., and Johnston, D. T.: Patterns of sulfur isotope fractionation during microbial sulfate reduction, Geobiology, 14, 91–101, https://doi.org/10.1111/gbi.12149, 2016.
Brick, S., Niggemann, J., Reckhardt, A., Könneke, M., and Engelen, B.: Interstitial microbial communities of coastal sediments are dominated by Nanoarchaeota, Front. Microbiol., 16, 1532193, https://doi.org/10.3389/fmicb.2025.1532193, 2025.
Bueno de Mesquita, C. P., Wu, D., and Tringe, S. G.: Methyl-based methanogenesis: an ecological and genomic review, Microbiol. Mol. Biol. R., 87, e00024-00022, https://doi.org/10.1128/mmbr.00024-22, 2023.
Callahan, B. J., Sankaran, K., Fukuyama, J. A., McMurdie, P. J., and Holmes, S. P.: Bioconductor workflow for microbiome data analysis: from raw reads to community analyses, F1000Research, 5, 1492, https://doi.org/10.12688/f1000research.8986.2, 2016a.
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., and Holmes, S. P.: DADA2: High-resolution sample inference from Illumina amplicon data, Nat. Methods, 13, 581–583, https://doi.org/10.1038/nmeth.3869, 2016b.
Canfield, D. E.: Isotope fractionation by natural populations of sulfate-reducing bacteria, Geochim. Cosmochim. Ac., 65, 1117–1124, https://doi.org/10.1016/s0016-7037(00)00584-6, 2001.
Chen, M., Conroy, J. L., Sanford, R. A., Wyman-Feravich, D. A., Chee-Sanford, J. C., and Connor, L. M.: Tropical lacustrine sediment microbial community response to an extreme El Niño event, Sci. Rep., 13, 6868, https://doi.org/10.1038/s41598-023-33280-2, 2023.
Chistoserdova, L.: Modularity of methylotrophy, revisited, Environ. Microbiol., 13, 2603–2622, https://doi.org/10.1111/j.1462-2920.2011.02464.x, 2011.
Chistoserdova, L., Kalyuzhnaya, M. G., and Lidstrom, M. E.: The expanding world of methylotrophic metabolism, Annu. Rev. Microbiol., 63, 477–499, https://doi.org/10.1146/annurev.micro.091208.073600, 2009.
Cline, J. D.: Spectrophotometric Determination of Hydrogen Sulfide in Natural Waters, Limnol. Oceanogr., 14, 454–458, https://doi.org/10.4319/lo.1969.14.3.0454, 1969.
Conrad, R.: Contribution of hydrogen to methane production and control of hydrogen concentrations in methanogenic soils and sediments, FEMS Microbiol. Ecol., 28, 193–202, https://doi.org/10.1111/j.1574-6941.1999.tb00575.x, 1999.
Conrad, R.: Quantification of methanogenic pathways using stable carbon isotopic signatures: a review and a proposal, Org. Geochem., 36, 739–752, https://doi.org/10.1016/j.orggeochem.2004.09.006, 2005.
Conrad, R.: Importance of hydrogenotrophic, acetoclastic and methylotrophic methanogenesis for methane production in terrestrial, aquatic and other anoxic environments: a mini review, Pedosphere, 30, 25–39, https://doi.org/10.1016/s1002-0160(18)60052-9, 2020.
Dean, J. F., Middelburg, J. J., Röckmann, T., Aerts, R., Blauw, L. G., Egger, M., Jetten, M. S., de Jong, A. E., Meisel, O. H., and Rasigraf, O.: Methane feedbacks to the global climate system in a warmer world, Rev. Geophys., 56, 207–250, https://doi.org/10.1002/2017rg000559, 2018.
Dershwitz, P., Bandow, N. L., Yang, J., Semrau, J. D., McEllistrem, M. T., Heinze, R. A., Fonseca, M., Ledesma, J. C., Jennett, J. R., DiSpirito, A. M., Athwal, N. S., Hargrove, M. S., Bobik, T. A., Zischka, H., and DiSpirito, A. A.: Oxygen generation via water splitting by a novel biogenic metal ion-binding compound, Appl. Environ. Microbiol., 87, e00286-00221, https://doi.org/10.1128/AEM.00286-21, 2021.
Deutzmann, J. S. and Schink, B.: Anaerobic oxidation of methane in sediments of Lake Constance, an oligotrophic freshwater lake, Appl. Environ. Microbiol., 77, 4429–4436, https://doi.org/10.1128/AEM.00340-11, 2011.
Dubois, N.: Traces of history in the sediments of Lake Joux, Swiss Federal Institute of Aquatic Science and Technology, https://www.eawag.ch/en/info/portal/news/news-detail/spuren-der-geschichte-im-lac-de-joux/ (last access: 24 April 2026) 2016.
Ellenbogen, J. B., Borton, M. A., McGivern, B. B., Cronin, D. R., Hoyt, D. W., Freire-Zapata, V., McCalley, C. K., Varner, R. K., Crill, P. M., and Wehr, R. A.: Methylotrophy in the Mire: direct and indirect routes for methane production in thawing permafrost, Msystems, 9, e00698-00623, https://doi.org/10.1128/msystems.00698-23, 2024.
Ettwig, K. F., Butler, M. K., Le Paslier, D., Pelletier, E., Mangenot, S., Kuypers, M. M., Schreiber, F., Dutilh, B. E., Zedelius, J., and de Beer, D.: Nitrite-driven anaerobic methane oxidation by oxygenic bacteria, Nature, 464, 543–548, https://doi.org/10.1038/nature08883, 2010.
Feldewert, C., Lang, K., and Brune, A.: The hydrogen threshold of obligately methyl-reducing methanogens, FEMS Microbiol. Lett., 367, fnaa137, https://doi.org/10.1093/femsle/fnaa137, 2020.
Fiskal, A., Deng, L., Michel, A., Eickenbusch, P., Han, X., Lagostina, L., Zhu, R., Sander, M., Schroth, M. H., Bernasconi, S. M., Dubois, N., and Lever, M. A.: Effects of eutrophication on sedimentary organic carbon cycling in five temperate lakes, Biogeosciences, 16, 3725–3746, https://doi.org/10.5194/bg-16-3725-2019, 2019.
Garcia, J.-L., Patel, B. K., and Ollivier, B.: Taxonomic, phylogenetic, and ecological diversity of methanogenic Archaea, Anaerobe, 6, 205–226, https://doi.org/10.1006/anae.2000.0345, 2000.
Habicht, K. S. and Canfield, D. E.: Sulfur isotope fractionation during bacterial sulfate reduction in organic-rich sediments, Geochim. Cosmochim. Ac., 61, 5351–5361, https://doi.org/10.1016/s0016-7037(97)00311-6, 1997.
Han, X.: Influence of eutrophication on microbial community structure, organic carbon sources, and organic carbon degradation in lake sediments through time, ETH Zurich, 2020.
Hanson, R. S. and Hanson, T. E.: Methanotrophic bacteria, Microbiol. Rev., 60, 439–471, https://doi.org/10.1128/mr.60.2.439-471.1996, 1996.
He, R., Wang, J., Pohlman, J. W., Jia, Z., Chu, Y.-X., Wooller, M. J., and Leigh, M. B.: Metabolic flexibility of aerobic methanotrophs under anoxic conditions in Arctic lake sediments, ISME J., 16, 78–90, https://doi.org/10.1038/s41396-021-01049-y, 2022.
Ho, A., Kerckhof, F. M., Luke, C., Reim, A., Krause, S., Boon, N., and Bodelier, P. L.: Conceptualizing functional traits and ecological characteristics of methane-oxidizing bacteria as life strategies, Env. Microbiol. Rep., 5, 335–345, https://doi.org/10.1111/j.1758-2229.2012.00370.x, 2013.
Holmer, M. and Storkholm, P.: Sulphate reduction and sulphur cycling in lake sediments: a review, Freshwater Biol., 46, 431–451, https://doi.org/10.1046/j.1365-2427.2001.00687.x, 2001.
Hoops, S. L. and Knights, D.: LMdist: Local Manifold distance accurately measures beta diversity in ecological gradients, Bioinformatics, 39, btad727, https://doi.org/10.1093/bioinformatics/btad727, 2023.
Huang, H., Xu, X., Shi, C., Liu, X., and Wang, G.: Response of taste and odor compounds to elevated cyanobacteria biomass and temperature, B. Environ. Contam. Tox., 101, 272–278, https://doi.org/10.1007/s00128-018-2386-5, 2018.
Huang, R., Soneson, C., Ernst, F. G., Rue-Albrecht, K. C., Yu, G., Hicks, S. C., and Robinson, M. D.: TreeSummarizedExperiment: a S4 class for data with hierarchical structure, F1000Research, 9, 1246, https://doi.org/10.12688/f1000research.26669.1, 2021.
Jarett, J. K., Nayfach, S., Podar, M., Inskeep, W., Ivanova, N. N., Munson-McGee, J., Schulz, F., Young, M., Jay, Z. J., and Beam, J. P.: Single-cell genomics of co-sorted Nanoarchaeota suggests novel putative host associations and diversification of proteins involved in symbiosis, Microbiome, 6, 1–14, https://doi.org/10.1186/s40168-018-0539-8, 2018.
Jørgensen, B. B., Weber, A., and Zopfi, J.: Sulfate reduction and anaerobic methane oxidation in Black Sea sediments, Deep-Sea Res. Pt. I, 48, 2097–2120, https://doi.org/10.1016/s0967-0637(01)00007-3, 2001.
Kessler, A. J., Chen, Y. J., Waite, D. W., Hutchinson, T., Koh, S., Popa, M. E., Beardall, J., Hugenholtz, P., Cook, P. L. M., and Greening, C.: Bacterial fermentation and respiration processes are uncoupled in anoxic permeable sediments, Nat. Microbiol., 4, 1014–1023, https://doi.org/10.1038/s41564-019-0391-z, 2019.
Khatun, S., Berg, J. S., Jézéquel, D., Moiron, M., Escoffier, N., Schubert, C. J., Bouffard, D., and Perga, M. E.: Long-range transport of littoral methane explains the metalimnetic methane peak in a large lake, Limnol. Oceanogr., 69, 2095–2108, https://doi.org/10.1002/lno.12652, 2024.
Kits, K. D., Klotz, M. G., and Stein, L. Y.: Methane oxidation coupled to nitrate reduction under hypoxia by the Gammaproteobacterium Methylomonas denitrificans, sp. nov. type strain FJG1, Environ. Microbiol., 17, 3219–3232, https://doi.org/10.1111/1462-2920.12772, 2015.
Knief, C.: Diversity and habitat preferences of cultivated and uncultivated aerobic methanotrophic bacteria evaluated based on pmoA as molecular marker, Front. Microbiol., 6, 1346, https://doi.org/10.3389/fmicb.2015.01346, 2015.
Knittel, K. and Boetius, A.: Anaerobic oxidation of methane: progress with an unknown process, Annu. Rev. Microbiol., 63, 311–334, https://doi.org/10.1146/annurev.micro.61.080706.093130, 2009.
Lamb, A. L., Wilson, G. P., and Leng, M. J.: A review of coastal palaeoclimate and relative sea-level reconstructions using δ13C and ratios in organic material, Earth-Sci. Rev., 75, 29–57, https://doi.org/10.1016/j.earscirev.2005.10.003, 2006.
Lavrieux, M., Schubert, C. J., Hofstetter, T., Eglinton, T. I., Hajdas, I., Wacker, L., and Dubois, N.: From medieval land clearing to industrial development: 800 years of human-impact history in the Joux Valley (Swiss Jura), Holocene, 27, 1443–1454, https://doi.org/10.1177/0959683617693892, 2017.
Li, B., Tao, Y., Mao, Z., Gu, Q., Han, Y., Hu, B., Wang, H., Lai, A., Xing, P., and Wu, Q. L.: Iron oxides act as an alternative electron acceptor for aerobic methanotrophs in anoxic lake sediments, Water Res., 234, 119833, https://doi.org/10.1016/j.watres.2023.119833, 2023.
Lods-Crozet, B., Reymond, O., and Strawczynski, A.: Evaluation de la qualité chimique et biologique du lac de Joux (Jura Suisse) entre 1985 et 2004, Bull. Soc. Ne. Sci. Nat., 129, 29–47, 2006.
Lv, P.-L., Shi, L.-D., Wang, Z., Rittmann, B., and Zhao, H.-P.: Methane oxidation coupled to perchlorate reduction in a membrane biofilm batch reactor, Sci. Total Environ., 667, 9–15, https://doi.org/10.1016/j.scitotenv.2019.02.330, 2019.
Magny, M., Gauthier, E., Vannière, B., and Peyron, O.: Palaeohydrological changes and human-impact history over the last millennium recorded at Lake Joux in the Jura Mountains, Switzerland, Holocene, 18, 255–265, https://doi.org/10.1177/0959683607086763, 2008.
Martinez-Cruz, K., Leewis, M. C., Herriott, I. C., Sepulveda-Jauregui, A., Anthony, K. W., Thalasso, F., and Leigh, M. B.: Anaerobic oxidation of methane by aerobic methanotrophs in sub-Arctic lake sediments, Sci. Total Environ., 607, 23–31, https://doi.org/10.1016/j.scitotenv.2017.06.187, 2017.
Martinez-Cruz, K., Sepulveda-Jauregui, A., Casper, P., Anthony, K. W., Smemo, K. A., and Thalasso, F.: Ubiquitous and significant anaerobic oxidation of methane in freshwater lake sediments, Water Res., 144, 332–340, https://doi.org/10.1016/j.watres.2018.07.053, 2018.
Mayr, M. J., Zimmermann, M., Guggenheim, C., Brand, A., and Bürgmann, H.: Niche partitioning of methane-oxidizing bacteria along the oxygen–methane counter gradient of stratified lakes, ISME J., 14, 274–287, https://doi.org/10.1038/s41396-019-0515-8, 2020.
McLaren, M. R. and Callahan, B. J.: Silva 138.1 prokaryotic SSU taxonomic training data formatted for DADA2, Zenodo, https://doi.org/10.5281/zenodo.4587955, 2021.
McMurdie, P. J. and Holmes, S.: phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data, PLOS ONE, 8, e61217, https://doi.org/10.1371/journal.pone.0061217, 2013.
Meier, D., van Grinsven, S., Michel, A., Eickenbusch, P., Glombitza, C., Han, X., Fiskal, A., Bernasconi, S., Schubert, C. J., and Lever, M. A.: Hydrogen-independent CO2 reduction dominates methanogenesis in five temperate lakes that differ in trophic states, ISME Communications, 4, ycae089, https://doi.org/10.1093/ismeco/ycae089, 2024.
Meyers, P. A.: Preservation of elemental and isotopic source identification of sedimentary organic matter, Chem. Geol., 114, 289–302, https://doi.org/10.1016/0009-2541(94)90059-0, 1994.
Miller, L. G., Baesman, S. M., Carlström, C. I., Coates, J. D., and Oremland, R. S.: Methane oxidation linked to chlorite dismutation, Front. Microbiol., 5, 275, https://doi.org/10.3389/fmicb.2014.00275, 2014.
Milucka, J., Ferdelman, T. G., Polerecky, L., Franzke, D., Wegener, G., Schmid, M., Lieberwirth, I., Wagner, M., Widdel, F., and Kuypers, M. M.: Zero-valent sulphur is a key intermediate in marine methane oxidation, Nature, 491, 541–546, https://doi.org/10.1038/nature11656, 2012.
Mitchell, E., van der Knaap, W. O., van Leeuwen, J. F., Buttler, A., Warner, B. G., and Gobat, J.-M.: The palaeoecological history of the Praz-Rodet bog (Swiss Jura) based on pollen, plant macrofossils and testate amoebae (Protozoa), Holocene, 11, 65–80, https://doi.org/10.1191/095968301671777798, 2001.
Monchamp, M.-È., Bruel, R., Frossard, V., McGowan, S., Lavrieux, M., Muschick, M., Perga, M.-É., and Dubois, N.: Paleoecological evidence for a multi-trophic regime shift in a perialpine lake (Lake Joux, Switzerland), Anthropocene, 35, 100301, https://doi.org/10.1016/j.ancene.2021.100301, 2021.
Morales-Williams, A. M., Wanamaker Jr., A. D., and Downing, J. A.: Cyanobacterial carbon concentrating mechanisms facilitate sustained CO2 depletion in eutrophic lakes, Biogeosciences, 14, 2865–2875, https://doi.org/10.5194/bg-14-2865-2017, 2017.
Nijman, T. P., Amado, A. M., Bodelier, P. L., and Veraart, A. J.: Relief of phosphate limitation stimulates methane oxidation, Frontiers in Environmental Science, 10, 804512, https://doi.org/10.3389/fenvs.2022.804512, 2022.
Oswald, K., Milucka, J., Brand, A., Hach, P., Littmann, S., Wehrli, B., Kuypers, M. M., and Schubert, C. J.: Aerobic gammaproteobacterial methanotrophs mitigate methane emissions from oxic and anoxic lake waters, Limnol. Oceanogr., 61, S101-S118, https://doi.org/10.1002/lno.10312, 2016.
Parada, A. E., Needham, D. M., and Fuhrman, J. A.: Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples, Environ. Microbiol., 18, 1403–1414, https://doi.org/10.1111/1462-2920.13023, 2016.
Penger, J., Conrad, R., and Blaser, M.: Stable carbon isotope fractionation by methylotrophic methanogenic archaea, Appl. Environ. Microbiol., 78, 7596–7602, https://doi.org/10.1128/AEM.01773-12, 2012.
Phillips, A. A., White, M. E., Seidel, M., Wu, F., Pavia, F. F., Kemeny, P. C., Ma, A. C., Aluwihare, L. I., Dittmar, T., and Sessions, A. L.: Novel sulfur isotope analyses constrain sulfurized porewater fluxes as a minor component of marine dissolved organic matter, P. Natl. Acad. Sci. USA, 119, e2209152119, https://doi.org/10.1073/pnas.2209152119, 2022.
Piguet, A. (Eds.): Le territoire et la commune du Lieu jusqu’en 1536 : Le Sentier, Imprimerie R. Dupuis, Switzerland, 171 pp., 1946.
Pjevac, P., Hausmann, B., Schwarz, J., Kohl, G., Herbold, C. W., Loy, A., and Berry, D.: An economical and flexible dual barcoding, two-step PCR approach for highly multiplexed amplicon sequencing, Front. Microbiol., 12, 669776, https://doi.org/10.3389/fmicb.2021.669776, 2021.
Quast, C., Pruesse, E., Yilmaz, P., Gerken, J., Schweer, T., Yarza, P., Peplies, J., and Glöckner, F. O.: The SILVA ribosomal RNA gene database project: improved data processing and web-based tools, Nucleic Acid. Res., 41, D590-D596, https://doi.org/10.1093/nar/gks1219, 2012.
Raven, M., Fike, D., Gomes, M., and Webb, S.: Chemical and isotopic evidence for organic matter sulfurization in redox gradients around mangrove roots, Front. Earth Sci., 7, 98, https://doi.org/10.3389/feart.2019.00098, 2019.
Reis, P. C., Thottathil, S. D., Ruiz-González, C., and Prairie, Y. T.: Niche separation within aerobic methanotrophic bacteria across lakes and its link to methane oxidation rates, Environ. Microbiol., 22, 738–751, https://doi.org/10.1111/1462-2920.14877, 2020.
Reis, P. C., Tsuji, J. M., Weiblen, C., Schiff, S. L., Scott, M., Stein, L. Y., and Neufeld, J. D.: Enigmatic persistence of aerobic methanotrophs in oxygen-limiting freshwater habitats, ISME J., 18, wrae041, https://doi.org/10.1093/ismejo/wrae041, 2024.
Rissanen, A. J., Saarenheimo, J., Tiirola, M., Peura, S., Aalto, S. L., Karvinen, A., and Nykänen, H.: Gammaproteobacterial methanotrophs dominate methanotrophy in aerobic and anaerobic layers of boreal lake waters, Aquat. Microb. Ecol., 81, 257–276, https://doi.org/10.3354/ame01874, 2018.
Rissanen, A. J., Jilbert, T., Simojoki, A., Mangayil, R., Aalto, S. L., Khanongnuch, R., Peura, S., and Jäntti, H.: Organic matter lability modifies the vertical structure of methane-related microbial communities in lake sediments, Microbiology Spectrum, 11, e01955-01923, https://doi.org/10.1128/spectrum.01955-23, 2023.
Ruff, S. E., Schwab, L., Vidal, E., Hemingway, J. D., Kraft, B., and Murali, R.: Widespread occurrence of dissolved oxygen anomalies, aerobic microbes, and oxygen-producing metabolic pathways in apparently anoxic environments, FEMS Microbiol. Ecol., 100, fiae132, https://doi.org/10.1093/femsec/fiae132, 2024.
Sanches, L. F., Guenet, B., Marinho, C. C., Barros, N., and de Assis Esteves, F.: Global regulation of methane emission from natural lakes, Sci. Rep., 9, 255, https://doi.org/10.1038/s41598-018-36519-5, 2019.
Saunois, M., Stavert, A. R., Poulter, B., Bousquet, P., Canadell, J. G., Jackson, R. B., Raymond, P. A., Dlugokencky, E. J., Houweling, S., Patra, P. K., Ciais, P., Arora, V. K., Bastviken, D., Bergamaschi, P., Blake, D. R., Brailsford, G., Bruhwiler, L., Carlson, K. M., Carrol, M., Castaldi, S., Chandra, N., Crevoisier, C., Crill, P. M., Covey, K., Curry, C. L., Etiope, G., Frankenberg, C., Gedney, N., Hegglin, M. I., Höglund-Isaksson, L., Hugelius, G., Ishizawa, M., Ito, A., Janssens-Maenhout, G., Jensen, K. M., Joos, F., Kleinen, T., Krummel, P. B., Langenfelds, R. L., Laruelle, G. G., Liu, L., Machida, T., Maksyutov, S., McDonald, K. C., McNorton, J., Miller, P. A., Melton, J. R., Morino, I., Müller, J., Murguia-Flores, F., Naik, V., Niwa, Y., Noce, S., O'Doherty, S., Parker, R. J., Peng, C., Peng, S., Peters, G. P., Prigent, C., Prinn, R., Ramonet, M., Regnier, P., Riley, W. J., Rosentreter, J. A., Segers, A., Simpson, I. J., Shi, H., Smith, S. J., Steele, L. P., Thornton, B. F., Tian, H., Tohjima, Y., Tubiello, F. N., Tsuruta, A., Viovy, N., Voulgarakis, A., Weber, T. S., van Weele, M., van der Werf, G. R., Weiss, R. F., Worthy, D., Wunch, D., Yin, Y., Yoshida, Y., Zhang, W., Zhang, Z., Zhao, Y., Zheng, B., Zhu, Q., Zhu, Q., and Zhuang, Q.: The Global Methane Budget 2000–2017, Earth Syst. Sci. Data, 12, 1561–1623, https://doi.org/10.5194/essd-12-1561-2020, 2020.
Schorn, S., Graf, J. S., Littmann, S., Hach, P. F., Lavik, G., Speth, D. R., Schubert, C. J., Kuypers, M. M., and Milucka, J.: Persistent activity of aerobic methane-oxidizing bacteria in anoxic lake waters due to metabolic versatility, Nat. Commun., 15, 5293, https://doi.org/10.1038/s41467-024-49602-5, 2024.
Schütz, H., Conrad, R., Goodwin, S., and Seiler, W.: Emission of hydrogen from deep and shallow freshwater environments, Biogeochemistry, 5, 295–311, https://doi.org/10.1007/bf02180069, 1988.
Singh, N., Kendall, M. M., Liu, Y., and Boone, D. R.: Isolation and characterization of methylotrophic methanogens from anoxic marine sediments in Skan Bay, Alaska: description of Methanococcoides alaskense sp. nov., and emended description of Methanosarcina baltica, Int. J. Syst. Evol. Micr., 55, 2531–2538, https://doi.org/10.1099/ijs.0.63886-0, 2005.
Söllinger, A. and Urich, T.: Methylotrophic methanogens everywhere—physiology and ecology of novel players in global methane cycling, Biochem. Soc. T., 47, 1895–1907, https://doi.org/10.1042/bst20180565, 2019.
Spangenberg, J. E. and Bosco-Santos, A.: Sulfur isotope analyses using 3× elemental analysis/isotope ratio mass spectrometry: Saving helium and energy while reducing analytical time and costs, Rapid Commun. Mass Sp., 38, e9866, https://doi.org/10.1002/rcm.9866, 2024.
Speth, D. R. and Orphan, V. J.: Metabolic marker gene mining provides insight in global mcrA diversity and, coupled with targeted genome reconstruction, sheds further light on metabolic potential of the Methanomassiliicoccales, PeerJ, 6, e5614, https://doi.org/10.7717/peerj.5614, 2018.
Sun, J., Mausz, M. A., Chen, Y., and Giovannoni, S. J.: Microbial trimethylamine metabolism in marine environments, Environ. Microbiol., 21, 513–520, https://doi.org/10.1111/1462-2920.14461, 2019.
Tamas, I., Smirnova, A. V., He, Z., and Dunfield, P. F.: The (d)evolution of methanotrophy in the Beijerinckiaceae—a comparative genomics analysis, ISME J., 8, 369–382, https://doi.org/10.1038/ismej.2013.145, 2014.
Tebbe, D. A., Gruender, C., Dlugosch, L., Lõhmus, K., Rolfes, S., Könneke, M., Chen, Y., Engelen, B., and Schäfer, H.: Microbial drivers of DMSO reduction and DMS-dependent methanogenesis in saltmarsh sediments, ISME J., 17, 2340–2351, https://doi.org/10.1038/s41396-023-01539-1, 2023.
Tranvik, L. J., Downing, J. A., Cotner, J. B., Loiselle, S. A., Striegl, R. G., Ballatore, T. J., Dillon, P., Finlay, K., Fortino, K., and Knoll, L. B.: Lakes and reservoirs as regulators of carbon cycling and climate, Limnol. Oceanogr., 54, 2298–2314, https://doi.org/10.4319/lo.2009.54.6_part_2.2298, 2009.
Tsola, S. L., Zhu, Y., Ghurnee, O., Economou, C. K., Trimmer, M., and Eyice, Ö.: Diversity of dimethylsulfide-degrading methanogens and sulfate-reducing bacteria in anoxic sediments along the Medway Estuary, UK, Environ. Microbiol., 23, 4434–4449, https://doi.org/10.1111/1462-2920.15637, 2021.
van Grinsven, S., Sinninghe Damsté, J. S., Abdala Asbun, A., Engelmann, J. C., Harrison, J., and Villanueva, L.: Methane oxidation in anoxic lake water stimulated by nitrate and sulfate addition, Environ. Microbiol., 22, 766–782, https://doi.org/10.1111/1462-2920.14886, 2020.
van Grinsven, S., Meier, D. V., Michel, A., Han, X., Schubert, C. J., and Lever, M. A.: Redox zone and trophic state as drivers of methane-oxidizing bacterial abundance and community structure in lake sediments, Frontiers in Environmental Science, 10, 857358, https://doi.org/10.3389/fenvs.2022.857358, 2022.
Vekeman, B., Kerckhof, F. M., Cremers, G., De Vos, P., Vandamme, P., Boon, N., Op den Camp, H. J., and Heylen, K.: New Methyloceanibacter diversity from North Sea sediments includes methanotroph containing solely the soluble methane monooxygenase, Environ. Microbiol., 18, 4523–4536, https://doi.org/10.1111/1462-2920.13485, 2016.
Veraart, A. J., Steenbergh, A. K., Ho, A., Kim, S. Y., and Bodelier, P. L.: Beyond nitrogen: the importance of phosphorus for CH4 oxidation in soils and sediments, Geoderma, 259, 337–346, https://doi.org/10.1016/j.geoderma.2015.03.025, 2015.
Wang, X.-C. and Lee, C.: Sources and distribution of aliphatic amines in salt marsh sediment, Org. Geochem., 22, 1005–1021, https://doi.org/10.1016/0146-6380(94)90034-5, 1994.
Wang, Y., Liu, X., Wu, M., and Guo, J.: Methane-Driven Perchlorate Reduction by a Microbial Consortium, Environ. Sci. Technol., 58, 13370–13379, https://doi.org/10.1021/acs.est.4c04439, 2024.
Wegener, G. and Boetius, A.: An experimental study on short-term changes in the anaerobic oxidation of methane in response to varying methane and sulfate fluxes, Biogeosciences, 6, 867–876, https://doi.org/10.5194/bg-6-867-2009, 2009.
Wegener, G., Krukenberg, V., Riedel, D., Tegetmeyer, H. E., and Boetius, A.: Intercellular wiring enables electron transfer between methanotrophic archaea and bacteria, Nature, 526, 587–590, https://doi.org/10.1038/nature15733, 2015.
Wei, H., Wang, M., Ya, M., and Xu, C.: The denitrifying anaerobic methane oxidation process and microorganisms in the environments: a review, Frontiers in Marine Science, 9, 1038400, https://doi.org/10.3389/fmars.2022.1038400, 2022.
Wei, T. and Simko, V.: R package “corrplot”: Visualization of a Correlation Matrix (Version 0.95), https://github.com/taiyun/corrplot (last access: 20 March 2025), 2024.
Werne, J. P., Hollander, D. J., Lyons, T. W., and Damsté, J. S. S.: Organic sulfur biogeochemistry: recent advances and future research directions, in: Sulfur Biogeochemistry – Past and Present, edited by: Amend, J. P., Edwards, K. J., and Lyons, T. W., Geological Society of America, 135–150, https://doi.org/10.1130/0-8137-2379-5.135, 2004.
Whiticar, M. J.: Carbon and hydrogen isotope systematics of bacterial formation and oxidation of methane, Chem. Geol., 161, 291–314, https://doi.org/10.1016/s0009-2541(99)00092-3, 1999.
Xia, F., Jiang, Q.-Y., Zhu, T., Zou, B., Liu, H., and Quan, Z.-X.: Ammonium promoting methane oxidation by stimulating the Type Ia methane-oxidizing bacteria in tidal flat sediments of the Yangtze River estuary, Sci. Total Environ., 793, 148470, https://doi.org/10.1016/j.scitotenv.2021.148470, 2021.
Xie, Z., Li, W., Yang, K., Wang, X., Xiong, S., and Zhang, X.: Bacterial and Archaeal Communities in Erhai Lake Sediments: Abundance and Metabolic Insight into a Plateau Lake at the Edge of Eutrophication, Microorganisms, 12, 1617, https://doi.org/10.3390/microorganisms12081617, 2024.
Xu, J. and Logan, B. E.: Measurement of chlorite dismutase activities in perchlorate respiring bacteria, J. Microbiol. Meth., 54, 239–247, https://doi.org/10.1016/s0167-7012(03)00058-7, 2003.
Yan, X., Xu, X., Wang, M., Wang, G., Wu, S., Li, Z., Sun, H., Shi, A., and Yang, Y.: Climate warming and cyanobacteria blooms: Looks at their relationships from a new perspective, Water Res., 125, 449–457, https://doi.org/10.1016/j.watres.2017.09.008, 2017.
Yang, R., Peng, C., Mo, Y., Kleindienst, S., Li, S., Wang, J., Kappler, A., Wang, Z., and Lu, L.: Electron acceptors modulate methane oxidation and active methanotrophic communities in anoxic urban wetland sediments, Appl. Environ. Microbiol., 91, e00386-00325, https://doi.org/10.1128/aem.00386-25, 2025.
Yang, Y., Chen, J., Tong, T., Li, B., He, T., Liu, Y., and Xie, S.: Eutrophication influences methanotrophic activity, abundance and community structure in freshwater lakes, Sci. Total Environ., 662, 863–872, https://doi.org/10.1016/j.scitotenv.2019.01.307, 2019.
Yang, Y., Chen, J., Tong, T., Xie, S., and Liu, Y.: Influences of eutrophication on methanogenesis pathways and methanogenic microbial community structures in freshwater lakes, Environ. Pollut., 260, 114106, https://doi.org/10.1016/j.envpol.2020.114106, 2020.
Yang, Y., Chen, J., Chen, X., Jiang, Q., Liu, Y., and Xie, S.: Cyanobacterial bloom induces structural and functional succession of microbial communities in eutrophic lake sediments, Environ. Pollut., 284, 117157, https://doi.org/10.1016/j.envpol.2021.117157, 2021.
Yvon-Durocher, G., Allen, A. P., Bastviken, D., Conrad, R., Gudasz, C., St-Pierre, A., Thanh-Duc, N., and Del Giorgio, P. A.: Methane fluxes show consistent temperature dependence across microbial to ecosystem scales, Nature, 507, 488–491, https://doi.org/10.1038/nature13164, 2014.
Zhao, Y., Liu, Y., Cao, S., Hao, Q., Liu, C., and Li, Y.: Anaerobic oxidation of methane driven by different electron acceptors: A review, Sci. Total Environ., 946, 174287, https://doi.org/10.1016/j.scitotenv.2024.174287, 2024.
Zhou, C., Peng, Y., Yu, M., Deng, Y., Chen, L., Zhang, L., Xu, X., Zhang, S., Yan, Y., and Wang, G.: Severe cyanobacteria accumulation potentially induces methylotrophic methane producing pathway in eutrophic lakes, Environ. Pollut., 292, 118443, https://doi.org/10.1016/j.envpol.2021.118443, 2022.
Zhu, Y., Chen, X., Yang, Y., and Xie, S.: Impacts of cyanobacterial biomass and nitrate nitrogen on methanogens in eutrophic lakes, Sci. Total Environ., 848, 157570, https://doi.org/10.1016/j.scitotenv.2022.157570, 2022.




