the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Bacterial contribution to nitrogen processing in the atmosphere
Frédéric Mathonat
François Enault
Raphaëlle Péguilhan
Muriel Joly
Mariline Théveniot
Jean-Luc Baray
Barbara Ervens
Pierre Amato
This study investigates potential microbial interactions with nitrogen compounds in the atmosphere, with a focus on inorganic forms (mainly NH, NO, and N2). The reanalysis of metagenomes and metatranscriptomes from cloud-free and cloudy air masses collected at the mountain site of puy de Dôme (1465 m a.s.l., France) indicates equivalent representation of genes involved in organic and inorganic nitrogen utilization processes. Glutamate metabolism and denitrification (in particular nitrite reduction) contributed most (70 %) of the microbial sequences of genes and transcripts linked to nitrogen utilization pathways. Other prevalent processes included assimilatory and dissimilatory nitrate reduction, and nitrogen fixation, with the latter being overexpressed in particular during clear atmospheric conditions. The screening of bacteria isolates revealed that 15 % of them carry the biomarker gene for biological N2 fixation (nifH). In addition, laboratory incubations of rainwater points towards the processing of NH. The decay rate of NH concentration correlated positively with the relative abundance of Sphingomonadales, and negatively with that of Burkholderiales. The latter may rather obtain nitrogen from N2 and organic forms. Overall, these results demonstrate multiple potential microbiological roles in the processing of inorganic nitrogen in the atmosphere, in relation with atmospheric conditions and microbial diversity. This opens up new perspectives in our understanding of biogeochemical cycles and chemical processing in the atmosphere, as well as microbial functioning in this major part of the Earth system.
Graphical Abstract. Representation of inorganic nitrogen cycle mediated by microorganisms in the atmosphere. Colors correspond to different steps in inorganic nitrogen cycle processes by atmospheric microorganisms The grey cloud indicates the cloud conditions, the sun indicates the clear atmosphere condition and the empty set indicates that this step does not occur in the atmosphere
- Article
(3242 KB) - Full-text XML
-
Supplement
(1323 KB) - BibTeX
- EndNote
The atmosphere carries diverse living microorganisms and plays an important role in the dispersal of biodiversity and genetic material across ecosystems (Fröhlich-Nowoisky et al., 2016). Biological indicators of viability and metabolic activity such as ATP and RNA could be identified in aerosol particles and cloud water, and regularly associated with bacteria taxa such as Alphaproteobacteria (Rhodospirillales, Sphingomonadales, and Rhizobiales) and Gammaproteobacteria (Pseudomonadales) (Amato et al., 2017; Klein et al., 2016; Šantl-Temkiv et al., 2018). As in other environments, their nutritive requirements include the uptake of carbon (C), nitrogen (N), phosphorous (P) and other elements from dissolved or gaseous inorganic and organic compounds, with potential impacts on biogeochemical cycles. However, the microbial processing of chemical compounds in the atmosphere is still very poorly studied.
A large variety of bioavailable organic and inorganic chemicals, such as carboxylic acids, aldehydes, sugars, are present in the atmospheric system as gas, particles, or dissolved in water (clouds, precipitation), and these are potential nutrients for microbial cells (Jaber et al., 2021; Renard et al., 2022). So far, the few experimental and model investigations of carbon utilization by microorganisms in clouds (Nuñez López et al., 2024; Vaïtilingom et al., 2013) led to the estimation that microbes may biodegrade up to 8–11 Tg yr−1 water-soluble organic carbon (WSOC) on a global scale (Ervens and Amato, 2020). Although some studies suggest interactions between airborne bacteria and nitrogen (Hill et al., 2007; Jaber et al., 2021; Péguilhan et al., 2025), they have not been quantified so far.
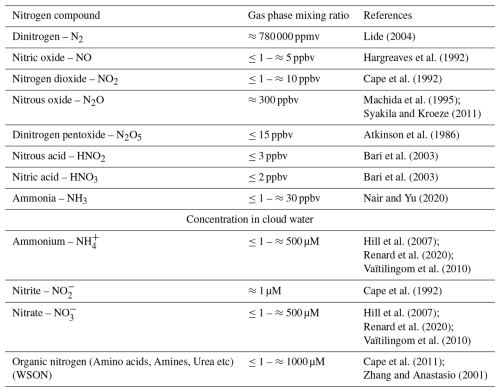
Nitrogen is an essential element for the biosphere. It is abundant in the atmosphere, which represents the planet's largest reservoir of inorganic nitrogen, primarily in the form of dinitrogen (N2), with 3.7 × 109 Tg (Sorai et al., 2007). Many other inorganic nitrogen compounds (e.g., NH, NO, NO, NOx), potentially bioavailable, are also present in the gas and aqueous phases (Table 1). They originate from natural sources (ocean, soils, biomass burning, etc.), anthropogenic activities (agriculture, industry), or are formed in the atmosphere from the processing of more complex molecules (Almaraz et al., 2018; Fowler et al., 2013).
Table 1Most common nitrogen compounds reported in the atmosphere and their concentration (WSON = Water Soluble Organic Nitrogen).
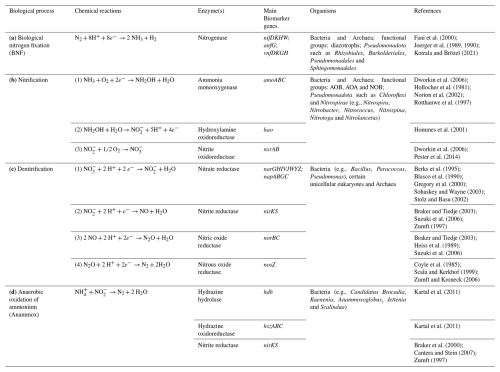
Due to their composition, microbial cells directly contribute to the nitrogen pool, with molecules such as proteins, DNA, pigments, and they can release N-containing organic compounds like glycoproteins, peptides and amino-acids (Decho and Lopez, 1993; Leck and Bigg, 2005). On the other hand, extracellular organic (amino acids, peptides, etc) and inorganic (NH, NO etc) compounds can be utilized as sources of nitrogen by living cells, detoxified, such as dinitrogen trioxide (N2O3) and peroxynitrite (ONOO−) that cause cellular damages (Espey et al., 2000; Maes et al., 2011), and also in some cases (for instance nitrates under low-oxygen conditions), used as electron acceptors for the generation of biochemical energy (Bernhard, 2010; Galimand et al., 1991). Regarding inorganic nitrogen, microbially-mediated transformation include N2 fixation, nitrification, denitrification, anaerobic oxidation of ammonium, ammonification and assimilation/dissimilation of nitrate. These processes have been well characterized in environments such as soil and freshwater and specific enzymes and genes could be identified (Bernhard, 2010) (Table 2). In the atmosphere, the reported microbial nitrogen-related processes include nitrosative stress, through atmospheric metagenomes and metatranscriptomes, and in cloud water, the processing of dissolved amino acids (Amato et al., 2019; Jaber et al., 2021; Péguilhan et al., 2025).
In the present study, a range of approaches, from the targeted reanalysis of sequence data to the observation of natural samples, were used in order to provide a large picture of the possible interactions between bacteria and inorganic nitrogen in the atmosphere. This work provides new insights into the functioning of microorganisms in the atmospheric system and their potential contribution to the nitrogen cycle.
To study the possible interactions occurring between inorganic nitrogen and atmospheric microorganisms, complementary approaches were combined: the prospection of sequences affiliated with biomarkers of the microbial nitrogen cycle in atmospheric metagenomes and metatranscriptomes (Sect. 2.1); the screening of bacteria isolates from clouds for biomarkers of nitrogen fixation (Sect. 2.2); the analysis of bacterial diversity and dissolved nitrogen ions during rainwater incubations (Sect. 2.3).
2.1 Reanalysis of metagenomes and metatranscriptomes from clouds and clear atmosphere
A set of atmospheric metagenomes (MG) and metatranscriptomes (MT) reported previously (ENA project PRJEB54740; Péguilhan et al., 2025) was reprocessed and reanalyzed specifically for genes and transcripts related to the processing of nitrogen compounds at the sequence level. Briefly, six aerosol (clear atmosphere) and nine cloud samples were originally collected in 2019–2020, from the instrumented atmospheric station of puy de Dôme Mountain summit (1465 m a.s.l., France) (Baray et al., 2020). For both atmospheric conditions, multiple high-flow rate impingers were deployed in parallel, each running at an air flow rate of 2 m3 h−1 (Šantl-Temkiv et al., 2017), and the samples were accumulated for 2 to 6 h directly into a nucleic acid preservation buffer, at 0.5X concentration for clear atmosphere and 1X for clouds in order to compensate for concentration/dilution due to evaporation/water accumulation, so as to reduce potential methodological biases. Clear-atmosphere samples were collected at relative humidity (RH) ranging from 41 % to 78 % (55 % on average) while clouds were characterized by RH = 100 % and liquid water content >0 g m−3.
The DNA and RNA extracts obtained from each sample were sequenced by Illumina HiSeq (paired end reads of 150 bp). In our study, the reads of MG and MT were trimmed (quality > 30, length > 145 bp), and 20 million randomly selected of them were aligned against the protein sequences of the NCycDB database, using MMseqs2 (e-value threshold of 10−7). NCycDB was designed for metagenomic profiling of N-cycling genes (Tu et al., 2019). This contains 68 gene (sub)families, grouped into 8 main functions: nitrification, denitrification, assimilatory nitrate reduction, dissimilatory nitrate reduction, nitrogen fixation, annamox, organic degradation and synthesis, and “other”. For each sample, the number of reads similar to one of the 68 gene families (Tu et al., 2019) was normalized by the total number of reads in the dataset and by the average size (base-pairs) of orthologs inside each family. Values are presented in parts per million base pairs (ppmbp). In order to obtain taxonomic affiliations, the matching sequences (best hits) were compared with the generalist UniRef100 protein database (January 2025) (Suzek et al., 2015).
The ratio between the relative abundance of transcripts in MTs with respect to that of their corresponding genes in MGs (RNA:DNA ratio) is often considered a proxy for microbial activity (e.g., Baldrian et al., 2012). The proportion of reads identified as being involved in each N-cycling function (i.e., nitrification, denitrification etc) in both MG and MT were summed up, and the corresponding “RNA:DNA ratio” of a given function, i.e., the number of related transcripts in an MT with respect to that of genes in the corresponding MG, was calculated for each sample.
2.2 Screening for nifH in bacteria strains isolated from clouds
The presence of the biomarker gene for nitrogen fixation, nifH, was tested in 34 bacterial strains, previously isolated from clouds (Amato et al., 2007; Renard et al., 2016; Vaïtilingom et al., 2012; Vinatier et al., 2016) (Supplement Table S1). These were selected to include known nitrogen-fixing taxa such as Pseudomonadales, Sphingomonadales, Burkholderiales, Rhizobiales, Mycobacteriales, Rhodospirillales, Hyphomicrobiales, Rhodobacterales. Bacteria cells from cultures stored at −80 °C in 10 % glycerol were re-cultured on R2A medium at 17 °C. DNA was then extracted from colonies (QIAamp DNA Mini Kit; QIAGEN; ref 51304) following the manufacturer's protocol. A polymerase chain reaction targeting the gene nifH was performed from DNA extracts (∼ 20 ng), using the primers polF: 5'-TGCGAYCCSAARGCBGACTC-3' and polR: 5'- ATSGCCATCATYTCRCCGGA-3' from Poly et al. (2001). Amplification was carried out at final concentrations of 200 µM of each dNTP (Qiagen), 0.2 µM of each primer and 0.04 U of Platinum II Taq Hot-Start DNA Polymerase (Invitrogen; ref 14966005), in Platinum II PCR buffer (1X) and Platinum GC enhancer (1X) provided with the Taq, in a total final volume of 25 µL. The PCR was performed with the following thermocycling parameters: 7 min of initial DNA polymerase activation and DNA denaturation at 95 °C, followed by 30 cycles of denaturation (95 °C, 1 min), hybridization (55 °C, 1 min), elongation (72 °C, 2 min), and a final elongation step of 7 min at 72 °C. The length of the amplicons (∼ 450 bp) was controlled by electrophoresis on 2 % agarose gel.
2.3 Analysis of rainwater samples
2.3.1 Sample collection, processing and storage
Rain samples were collected in 2023 at the Theix INRAe meteorological station, France (GPS point: 45°43′22′′ N, 3°01′09′′ E), a meadow at an altitude of 881 m a.s.l. located nearby (∼ 10 km) puy de Dôme observatory, where the samples for MG and MT were obtained (Sect. 2.1). The samples were collected over 24 h periods using an automated refrigerated (4 °C) precipitation collector (Eigenbrodt NSA 181/KHS; Königsmoor, Germany), with a collection diameter of 47.4 cm (area of 1764 cm2), into autoclaved one-liter glass bottles (Pouzet et al., 2017). The samples maintained at 4 °C in the sampler were processed within two days. Due to practical limitations, only samples with volumes >30 mL were analyzed for chemical and biological variables (see below). In addition, samples > 100 mL were distributed into sterile Erlenmeyer flasks as triplicates and incubated for 5 d under laboratory conditions (17 °C, 130 rpm shaking, dark conditions), in order to evaluate temporal changes in the microbial community and inorganic nitrogen concentrations.
2.3.2 Meteorological data and air mass back-trajectory analyses
Meteorological variables (temperature, wind, humidity) (Supplement) were monitored by the meteorological station at the Theix INRAe. Boundary layer height (BLH) and the geographical origin of the air masses (cardinal points, terrestrial or oceanic, altitude) were obtained from 72 h backward trajectories computed by the Computing Atmospheric Trajectory (CAT) tool as in Péguilhan et al. (2021).
2.3.3 Chemical analyses
The pH was measured immediately after sampling in a subsample of ∼ 1.5 mL. For quantifying major inorganic ions, a volume of 5 mL was transferred into a 15 mL sterile polypropylene vial and stored at −25 °C until analysis. The main dissolved inorganic cations and anions (Na+, NH, K+, Mg2+, Ca2+, Cl−, NO and SO) were examined by ion chromatography using a Dionex ICS1500 (column CS16) for cations as in Deguillaume et al. (2014), and a Dionex DX320 (column AS11) for anions as in Péguilhan et al. (2021).
In incubation experiments, ammonium and nitrate concentration decay rates (Δ, µmol h−1 cell−1) were calculated as:
Where (µmol L−1 h−1) corresponds to the change in ammonium or nitrate concentration during the incubation time of 120 h and xbact corresponds to the initial number concentration of microorganisms in the sample, in cells L−1.
2.3.4 Total cell counts
For the total microbial cell quantification by flow cytometry (FCM), rain samples were prepared as in Amato et al. (2017). Briefly, for each sample, triplicate volumes of 450 µL were mixed with 50 µL of 5 % glutaraldehyde (0.5 % final concentration; Sigma-Aldrich G7651, St-Louis, MO, USA), gently vortexed and stored at 4 °C until analysis within 3 weeks. Total cell concentration was quantified using a LSR Fortessa flow cytometer (Becton Dickinson, Franklin Lakes, NJ). Just before analysis, samples were mixed with 500 µL of 0.22 µm porosity filtered (47 mm diameter; ClearLine 0421A00023) Tris-EDTA pH 8.0 (40 mM Tris-Base, 1 mM EDTA, acetic acid to pH 8.0) and stained with SYBRGreen I (Molecular Probes Inc., Eugene, OR, USA) from a 100X solution, and incubated for 15 min in the dark. Cell counts were performed at excitation and emission wavelengths of λexc = 488 nm and λem = 530 nm, respectively, at a flow rate of ∼ 90 µL min−1 further determined by weighting.
For each rainwater incubation experiment, a microbial growth rate (μ) in gen h−1 was calculated as:
Where N and N0 are the number concentrations of cells at the end and at the beginning of the incubation period, respectively, in cells mL−1; dt corresponds to the incubation time (120 h).
The generation time (Θ), in h gen−1 is then:
2.3.5 DNA extraction
For DNA extraction, the remaining volume of rainwater (30 to 200 mL, depending on the sample) was extracted from mixed cellulose esters (MCE)-filtered samples (0.22 µm porosity, 47 mm diameter; ClearLine 0421A00023), using Macherey-Nagel NucleoMag® DNA/RNA Water Kit (744220.1). Each filter was stored in 5 mL Type A NucleoSpin Bead Tubes (Macherey-Nagel, 740799.50), added with 1.2 mL of lysis buffer MWA1, as in Rossi et al. (2023). For DNA extraction, bead-beating lysates (10 min at maximum vortex speed on a Genie2 vortex) were processed following a protocol adapted for 47 mm filter membranes (manufacturer's protocol: https://www.mn-net.com/media/pdf/ce/b5/38/Instruction-NucleoMag-DNA-RNA-Water.pdf, last access: February 2025). DNA was finally eluted into 50 µL RNase-free H2O. The quality ( and ratios) and concentration (ng µL−1) of the extracts were assessed using a BioSpec-nano (SHIMADZU BIOTECH corporation). The extraction products were stored at −25 °C until DNA gene quantification (q-PCR) and 16S rRNA gene amplification and sequencing.
2.3.6 Gene quantification, reaction mixture and amplification conditions for q-PCR
In order to quantify the ammonium-to-nitrate transformation gene (amoA), homemade standard assays were performed using plasmids (pEX-A128; Eurofins Genomics), in which sequences of amoA (Nitrosomonas europaea N accession L08050) (McTavish et al., 1993) and 16S rRNA gene (Pseudomonas syringae isolate PDD-32b-74 accession HQ256872.1) were ligated. Plasmids containing the genes of interest were linearized with the restriction enzyme EcoRI FastDigest (ThermoFischer; ref FD0274) during 15 min at 37 °C and the enzyme was inactivated during 5 min at 80 °C. The concentration of the plasmid DNA solutions was measured spectrophotometrically, and converted into gene copies µL−1, using the following formula from Whelan et al. (2003):
Where [DNA] is the concentration of recombinant plasmids in ng µL−1, NA is Avogadro's constant (6.023 × 1023 molecules mol−1), n is the length of the gene sequence in base pairs and mw is the average molecular mass of a base pair (660 g mol−1).
Plasmids including the target gene regions were provided by a subcontracted company (Eurofins Genomics, Lille, France), at a concentration of 19.68 ng µL−1 for 16S rRNA and 11.38 g µL−1 for amoA. The plasmid (vector + gene insert) for 16S rRNA and amoA genes have a total length of 2799 bp (2450 + 349 bp, respectively) and 2982 bp (2450 bp + 532 bp, respectively). From Eq. (4), the plasmid stock solutions for 16S rRNA and amoA genes were therefore at concentrations of 6.42 × 109 copies µL−1, and 3.48 × 109 copies µL−1, respectively. Standards for quantification were obtained by decimal dilutions of the stock solutions, considering concentrations between 2 and 108 copies µL−1. Limits of quantification were determined using the linearity of the standard curve for Ct between 10–30/5–20 [amoA/16S].
The q-PCR analyses were performed as technical triplicates, in 96-well reaction plates along with a minimum of two no-template controls and standard samples consisting of 2–106 amoA copies or 102–108 16S rRNA gene copies per reaction. Each 20 µL qPCR reaction volume contained 10 µL of IQ™ SYBR® Green Supermix assays (BIO-RAD; ref 1708882), 300 nM of each primer [amoA-1F: 5'- GGGGTTTCTACTGGTGGT -3' and amoA-2R: 5'- CCCCTCKGSAAAGCCTTCTTC -3' for amoA (Rotthauwe et al., 1997); EUBf: 5'-GGTAGTCYAYGCMSTAAACG-3' and EUBr: 5'-GACARCCATGCASCACCTG-3' for the 16S rRNA gene (Bach et al., 2002)], 8 µL or 5 µL (at a concentration of 15–50 ng µL−1) of DNA extract as the template for amoA and 16S rRNA genes, respectively, and PCR grade water (20 µL final). The q-PCR was performed with the following thermocycling parameters: 3 min of initial DNA polymerase activation and DNA denaturation at 95 °C followed by 35 cycles of denaturation (95 °C, 15 s), hybridization and elongation (60 °C for amoA or 62 °C for 16S, 1 min). The specificity was finally assessed by following up melting curves at temperatures from 60 to 95 °C, with a heating rate of 0.1 °C s−1. The standard curves had a linearity r2 > 0.99 and an efficiency (E) between 80 % and 120 %. The maximum standard deviation for each triplicate of q-PCR was set at 0.3 cycle threshold (CT). The q-PCR results were converted to gene copy numbers per mL of rainwater using the standard curves as a reference.
2.3.7 Ribosomal gene amplification and sequencing, bioinformatics and statistics
Amplification of the V4 region of the 16S sub-unit of bacterial ribosomal genes was performed from genomic DNA extracts by PCR, using the universal primers 515f (5'-GTGYCAGCMGCCGCGGTAA-3') and 806r (3'-GGACTACNVGGGTWTCTAAT-5') (Apprill et al., 2015; Parada et al., 2016) and the Platinum II Taq Hot-Start DNA Polymerase (Invitrogen; ref 14966005), following the conditions specified in Bulgarelli et al. (2012). Amplicons were purified using the QIAquick PCR Purification kit (QIAGEN; Hilden, Germany) and sequenced on Illumina Miseq 2 × 250 bp (GenoScreen; Lille, France).
The sequence data have been deposited in the European Nucleotide Archive (ENA) at EMBL-EBI under the accession number PRJEB91356. A total of 9 292 119 sequences were obtained following demultiplexing for the 57 samples investigated. Each sample contributed between 8350 and 50 157 16S rRNA gene reads. For taxa annotation, the FROGS pipeline [Find Rapidly OTUs with Galaxy Solution; Escudié et al., 2018] was used through the Metabarcoding.useGalaxy v4.1.0 (“Materials and Methods” in the Supplement).
2.3.8 Statistical analyses
All statistical tests were carried out using Past software (v.4.03) (Hammer et al., 2001). To compare the mean proportions of the different functional categories in metagenomic (MG) and metatranscriptomic (MT) datasets, a parametric Student's t-test or a non-parametric Mann–Whitney test was performed. RNA:DNA ratios were compared to a value of 1 using a one-sample Wilcoxon test, and the mean RNA:DNA ratios of the different functional categories between cloudy and clear-air conditions were assessed using the parametric Student's t-test or the non-parametric Mann–Whitney test.
For the rainwater incubation experiments, correlations among the different factors were evaluated using Pearson's rank correlation test. Mean values derived from the incubations (e.g., ion concentrations, bacterial abundances) were compared using the non-parametric Wilcoxon test.
3.1 Atmospheric metagenomes reanalysis
In atmospheric MG, organic and inorganic nitrogen processing genes are equivalently represented (p>0.05, Student's test), contributing a total of 4.08 ± 2.33 parts per million base pairs (ppmbp). Organic nitrogen processes are dominated by glutamate and urea metabolisms (2.01 ± 0.53 parts per million) (Fig. 1A; Supplement Fig. S1 and Supplement Table S2). Regarding inorganic processes, together, denitrification, assimilatory and dissimilatory nitrate reduction, and biological nitrogen fixation contribute ∼ 99 % of the N-related reads, with respective contributions of 46 % ± 15 % (0.96 ± 0.36 ppmbp), 25 % ± 3 % (0.51 ± 0.09 ppmbp), 19 % ± 10 % (0.40 ± 0.24 ppmbp) and 9 % ± 4 % (0.18 ± 0.11 ppmbp). Nitrification and anammox contribute < 1 % (Fig. 1A; Supplement Fig. S1 and Supplement Table S2). Within denitrification, nitrite reduction prevails (0.71 ± 0.25 ppmbp), with two genes (nirK and nirS) contributing 75 % ± 26 % of the denitrification genes, and 35 % ± 12 % of the genes involved in inorganic nitrogen transformations. Nitrate reduction genes (napABC, narGHIJVWYZ) evenly contribute between 0 % and 40 % of the denitrification genes (Supplement Fig. S2). Within biological nitrogen fixation, the two most represented biomarker genes are nifH and nifW, whereas nifD and nifK are underrepresented (Supplement Fig. S2).
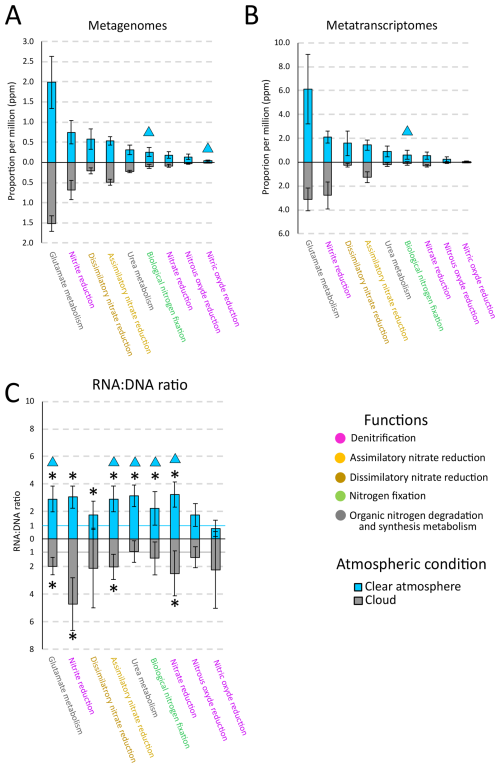
Figure 1Proportions (per million base pairs) of nitrogen processing genes (with a threshold value greater than 0.1 ppmbp) in (A) atmospheric metagenomes, (B) metatranscriptomes, and (C) corresponding RNA:DNA ratios, in clouds (grey) or clear atmosphere (blue) at the puy de Dôme mountain station. For (C), the blue line corresponding to the value 1 for clear atmosphere condition and the grey line corresponding to the value 1 for cloud condition. Triangles represent significant differences between clear atmosphere and cloud condition (Student or Mann-Whitney test). Asterisks (*) indicate that the RNA:DNA ratio is different from 1 (Student or Wilcoxon test).
Overall, nitrogen processing genes are more represented in MG during clear atmospheric conditions (4.79 ± 1.66 ppmbp) than in clouds (3.36 ± 0.65 ppmbp) (p<0.05, Student's test). Inorganic nitrogen processing genes contribute 2.50 ± 0.90 and 1.63 ± 0.47 ppmbp in clear atmosphere and in clouds, respectively, of which biological nitrogen fixation genes are 0.26 ± 0.11 ppmbp (i.e., 10 ± 4 % of the inorganic N-related reads) and 0.10 ± 0.05 ppmbp (i.e., 6 ± 3 %) (p<0.05, Student's test), respectively (Fig. 1A; Supplement Fig. S1 and Supplement Table S2).
3.2 Active microbial processes in the atmosphere and difference between clouds and clear atmosphere
Functional expression was evaluated considering the relative representation of transcripts (mRNA) in MT (Fig. 1B) with respect to their corresponding genes in MG (RNA:DNA ratio, with higher values indicating higher expression levels). While ribosomal RNAs can persist up to several days at low temperatures (Schostag et al., 2020), the average half-life of an mRNA in a bacteria cell (Mycobacterium tuberculosis) at 37 °C is between 2 and 5 min, which is much shorter than the average duration of atmospheric transport (∼ 3–4 d; Burrows et al., 2009). Some studies have shown that under “stressful” conditions or during dormancy/inactive states, such as caused by a shift in temperature, the half-life may increase by a factor of 2–3, but it still remains on the order of only a few tens of minutes (Rustad et al., 2013). Most of the transcripts identified in cloud and aerosol samples were therefore likely produced by the cells while airborne.
On average, nitrogen processing genes are more represented in MT than they are in the corresponding MG with an RNA:DNA ratio > 1 (p<0.05, One-sample Wilcoxon test). Nitrate reduction, nitrate assimilation, biological nitrogen fixation, as well as glutamate and urea metabolisms all have an RNA:DNA ratio > 1 (p<0.05, One-sample Wilcoxon test), and this is higher in clear atmospheric conditions than in clouds (p<0.05, Mann-Whitney test) (Fig. 1C).
Functions with RNA:DNA ratios that are significantly greater than unity (p<0.05, One-sample Wilcoxon test) in clear atmosphere include nitrate reduction (3.23 ± 0.92), nitrite reduction (3.04 ± 0.81), nitrate assimilation (2.90 ± 0.94), dissimilatory nitrate reduction (1.73 ± 1.01), biological nitrogen fixation (2.22 ± 1.21), and genes associated with organic nitrogen metabolism, i.e., glutamate (2.90 ± 0.92) and urea metabolisms (3.15 ± 0.80). In clouds, functions with a ratio significantly > 1 include nitrate reduction (2.49 ± 1.62), nitrite reduction (4.71 ± 1.92), nitrate assimilation (2.01 ± 0.91), and glutamate metabolism genes (1.98 ± 0.64).
Functions with similar representation in MT as in MG (RNA:DNA ratio ∼ 1) (p>0.05, One-sample Wilcoxon test) include, in clear atmosphere conditions, nitric oxide reduction (0.76 ± 0.58) and nitrous oxide reduction (1.77 ± 0.83). In cloud conditions, the same applies to nitric oxide reduction (2.25 ± 2.77), nitrous oxide reduction (1.33 ± 0.76), dissimilatory nitrate reduction (2.10 ± 2.89), biological nitrogen fixation (1.40 ± 1.19), and urea metabolism (0.91 ± 0.78) (Fig. 1C).
Under clear-sky conditions, the atmospheric samples used to generate the data were collected at relative humidity (RH) values ranging from 41 % to 78 %, with a mean of 55 %; no relationship between the expression of biological functions and RH could be detected (see Péguilhan et al., 2025 for further details). Nevertheless, RH is known to impact the viability of model airborne bacteria, with often higher survival at extremely low or high RH levels (Cox and Goldberg, 1972; Wright et al., 1969), and to influence their gene expression patterns (Barnes and Wu, 2022). Larger datasets will be necessary to examine such relationships in the natural environment.
3.3 Bacteria associated with functional genes and transcripts
The bacteria associated with atmospheric nitrogen processing genes in MG and MT are affiliated with 61 distinct phyla and 331 orders (Fig. 2). Within each nitrogen function, the contributing taxa are evenly distributed at the phylum level, with no difference between clouds and clear atmospheric conditions, except for the rare taxa related to nitrification. The most represented phyla for most functions are Pseudomonadota (contributing from 5 % to 32 % of the N-associated functional sequences), Acidobacteriota, Actinomycetota, Bacteriodota, Bacillota and Planctomycetota (Fig. 2A). Some functions involve specific phyla, such as Nitrospirota and Nitrososphaerota for nitrification, and Cyanobacteriota for N2 fixation.
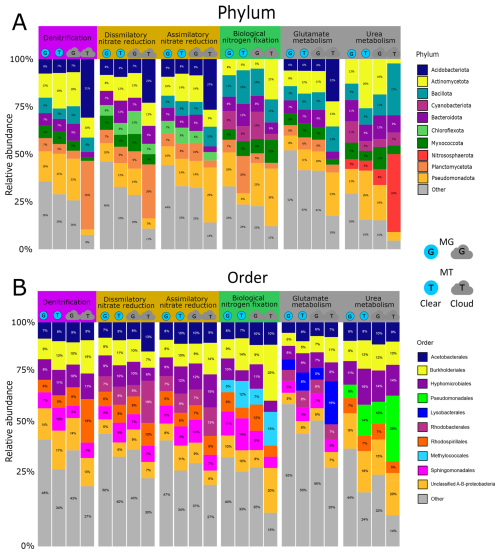
Figure 2Proportion of bacterial phyla (A) and orders (B) in clear-atmosphere and cloud metagenomes (MG) and metatranscriptomes (MT). For each active biological function linked to the nitrogen cycle (with a threshold value greater than 0.1 ppmbp in MG and MT), the eight major phyla and orders are represented by different colors. Proportions greater than or equal to 5 % are displayed directly on the figure.
At the order level, the dominant orders include Acetobacterales, Burkholderiales, Hyphomicrobiales, Sphingomonadales, and Rhodospirillales (Fig. 2B). Nitrification processes are contributed by orders such as Nitrososphaerales, Nitrosomonadales, Nitrospirales, Isosphaerales, and Cytophagales, which account for 100 % of the orders contributing transcripts in clouds. Certain bacterial orders are associated only with specific functions, such as Methylococcales for N2 fixation and Pseudomonadales for urea metabolism.
In terms of diversity, similar taxa contribute to MG and MT for a given process, with differences in the relative abundance of taxa's representation. The major phyla in MG are also those dominating in MT, notably Planctomycetota and Acidobacteriota, which, for the denitrification process, account for <5 % and 7 % of the reads in MG and 36 % and 31 % in MT, respectively, under cloud conditions. Regarding nitrification, Acidobacteriota is the only phylum represented in MT, while this contributes only 10 % of N-related sequences in MG.
3.4 Screening for potential diazotrophic microorganisms among isolates
Thirty-four (34) strains of Alphaproteobacteria and Gammaproteobacteria isolated from clouds in earlier work were tested by PCR for carrying the N2 fixation biomarker gene (nifH). These strains were selected to belong to taxa that include nitrogen-fixers, and they are among the most frequent viable bacteria in clouds (Vaïtilingom et al., 2012). In total, five strains (∼ 15 %) were positive: four Sphingomonadales and one Rhizobiales (Supplement Table S1).
3.5 Fresh rain water samples analysis
Among the 30 rain samples assessed in this study, ammonium and nitrate concentrations ranged from 3.3 ± 0.1 to 79.8 ± 0.1 µmol L−1, and from 1.0 ± 0.1 to 34.8 ± 0.1 µmol L−1, respectively (Supplement Table S3). This was linked with the geographical origin of the air masses, as attested by backward trajectories, with continental sources as major contributors (Supplement Fig. S3) (p<0.05, Pearson test, on the correlation between nitrate + ammonium concentration and the origin of air masses).
Bacterial concentration in the samples ranged from 1.94 ± 0.88 × 102 to 6.73 ± 1.31 × 104 cells mL−1 (Supplement Table S3). This was not correlated with the origin of the air masses (p>0.05, Pearson test, Supplement Table S4).
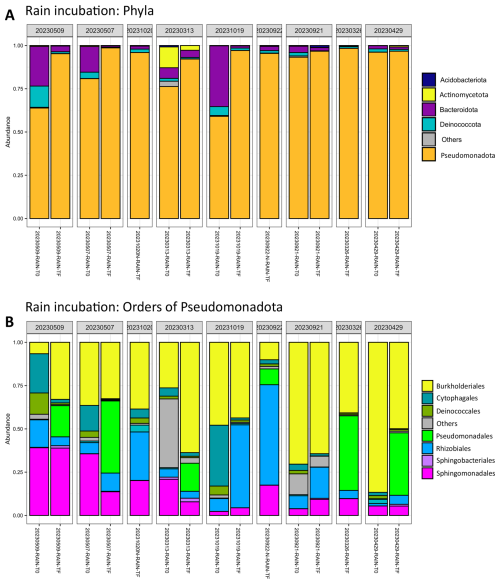
The majority of bacteria in rain samples are affiliated with Pseudomonadota (between 80 % and 90 % on average, Fig. 3). The major orders are Burkholderiales, Cytophagales, Pseudomonadales, Sphingomonadales, Sphingobacteriales and Rhizobiales (Fig. 3). There are clear differences in the distribution of these orders, in particular Sphingomonadales and Burkholderiales, depending on the sample (Fig. 3). The amoA gene, related to ammonium utilization in bacteria (ammonia monoxygenase), was below the detection limit of qPCR (less than 100.25 copies mL−1) in all rainwater samples, except 20230506_RAIN where 14 copies mL−1 could be quantified.
3.6 Incubation of rainwater samples
The bacteria biomass and diversity in the nine rainwater samples investigated changed during the five days of incubation (Fig. 4 and Table 3). This is a plausible duration for bacteria's residence in the atmosphere, estimated around 3 to 4 d (Burrows et al., 2009), but the actual time spent by cells within atmospheric droplets is expected to be much shorter. The data were interpolated over the 5 d periods in order to determine rates, assuming, as a first-order approximation, that these remained constant throughout the incubation time.
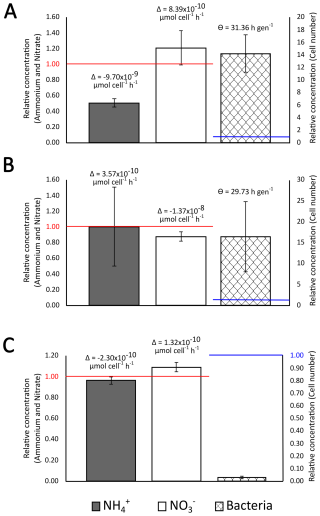
Figure 4Typical evolution of the concentration of ammonium (grey histogram) nitrate (white histogram) and microbial cell number concentration (hatched histogram) in rainwater samples after incubation for 5 d at 17 °C and 130 rpm shaking, respect to initial values, and corresponding inferred average ammonium and nitrate bioassimilation rate (Δ) and bacteria generation time (Θ), when cell number concentration increased. (A) 20230922_RAIN, (B) 20230326_RAIN and (C) 20230429_RAIN. The results for the other incubations are shown in Supplement Fig. S4.
Table 3Characteristics of rain sample incubations. T0 and TF indicate initial and final (5 d) incubation times, respectively; bdl: below detection limit; N.A.: not available.
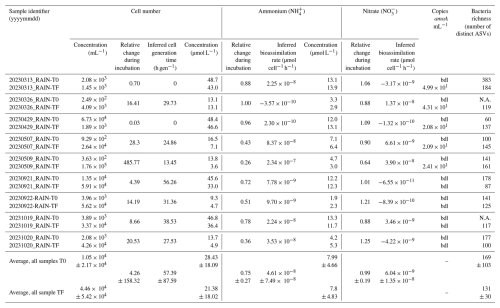
Depending on samples, at the end of incubation, the cell number concentration ranged from to 400 times that initially present. When increasing, this corresponded to generation times of 13.45 to 56.26 h gen−1.
The dominant orders initially were Burkholderiales, Cytophagales, Pseudomonadales, Sphingomonadales, Sphingobacteriales and Rhizobiales (Fig. 3). In some of the samples, their relative abundance was modified after incubation. For example, Burkholderiales' representation increased from 7 % to 30 % in the 20230509_RAIN sample, and from 26 % to 65 % in the 20230313_RAIN sample. Similar trends were observed for Rhizobiales in 20231019_RAIN, and for Pseudomonadales in 20230507_RAIN, 20230313_RAIN, and 20230429_RAIN, in these cases at the expense of the initially dominant order Burkholderiales.
The amoA gene could be quantified in the samples only upon incubation for five days (Table 3), with values ranging from 20 to 40 gene copies mL−1 of rainwater.
On average, NH concentration decreased over the nine days of incubation (Wilcoxon test; p<0.05) from 28.43 ± 18.09 to 21.38 ± 18.02 µmol L−1, i.e., between 0.26 ± 0.18 and 1.00 ± 0.50 of its initial concentration, depending on samples (Wilcoxon test; p<0.05) (Table 3). The inferred corresponding NH bioassimilation rates ranged between 2.34 × 10−7 and 2.30 × 10−10 µmol h−1 cell−1 (4.61 × 10−8 µmol h−1 cell−1 on average) i.e., this ranged over three orders of magnitude. On the contrary, NO concentration did not significantly change, with concentration of 7.99 ± 4.66 µmol L−1 in fresh samples and 7.87 ± 4.83 µmol L−1 after incubation, i.e., 0.64 ± 0.41 to 1.25 ± 0.23 its initial values (Fig. 4 and Table 3).
PCA (Fig. 5) illustrates the variability of rainwater sample composition, and its evolution during incubations. The two first components explain 52 % of the variance and allow discriminating in particular marine from continental air masses. Samples from air masses originating from marine areas (Atlantic Ocean) were enriched in Na+ and Cl− ions, whereas samples from continental air masses contained higher levels of NH, NO, and SO (p<0.05, Spearman's rank correlation). Continental air masses were also characterized by higher ambient temperatures at the sampling site, smaller water volumes, higher pH, and higher cell concentrations with respect to marine air masses (p<0.05). This is consistent with previous observations at this site (Péguilhan et al., 2021). Certain bacterial taxa could be also associated with air mass origin: the relative abundance of Sphingomonadales was significantly higher in samples from marine air masses, whereas Burkholderiales dominated in samples from continental air masses (p<0.05). Finally, the bioassimilation rates of ammonium and nitrate were positively correlated with the relative abundance of Sphingomonadales and negatively correlated with that of Burkholderiales (p<0.05), but they were independent from the initial concentrations of these ions, bacteria and amoA gene copies (p>0.05).
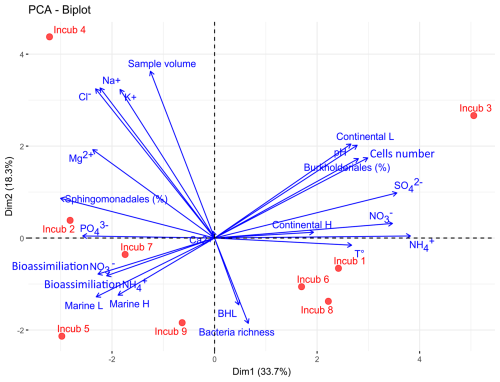
Figure 5Principal component analysis (PCA) between the meteorological and initial chemical and biological variables of rainwater samples and inorganic nitrogen bioassimilation rates during incubations (blue arrows). The red dots represent the different incubations, numbered in chronological order. BLH: Boundary Layer Height; L and H for “Marine” and “Continental” air mass origins indicate Low and High altitude, respectively.
Nitrogen compounds (NH3, NH, NO, NO, NOx, HNO3, N2O, urea, amines, proteins, etc.) are emitted to the atmosphere by a variety of natural and anthropogenic sources (Fowler et al., 2013; Sutton et al., 2013). In terrestrial ecosystems, excess reactive nitrogen disrupts the nutrient balance, reducing carbon storage (Galloway et al., 2002; Matson et al., 2002), acidifies soils and reduces biodiversity (Aber et al., 1995). The intensive use of nitrogen fertilizers in agriculture is therefore a major cause of soil pollution and degradation worldwide (Lal, 2015; Müller et al., 2018). This nitrogen is then washed into aquatic environments, causing eutrophication of fresh and coastal waters (Diaz and Rosenberg, 2008; Howarth, 2008). The nitrogen concentrations have been rising over the last few decades, amplifying the nitrogen cycle, producing more NOx, causing health issues and accelerating climate change (Cowling et al., 1998; Krey, 2014; Myhre et al., 2013). Therefore, a better understanding of the interactions between atmospheric nitrogen and microorganisms is essential.
Although some studies suggest interactions between airborne bacteria and nitrogen, including the biological degradation of amino acids in clouds and nitrosative stress in airborne microorganisms (Amato et al., 2019; Hill et al., 2007; Jaber et al., 2021; Péguilhan et al., 2025), information remains very limited. This study provides new insights into the interactions that may exist between nitrogen and microorganisms in the atmosphere.
4.1 The atmosphere expands surface ecosystems
The biological and chemical composition of the atmosphere mirrors to some extent that of emitting surfaces. In atmospheric metagenomes and metatranscriptomes, we find approximately equal proportions (∼ 50:50 %) of genes associated with inorganic and organic nitrogen transformation. This is similar to findings in other environments such as oceans, rivers, sediments and plants (Deng et al., 2024; Nie et al., 2021; Song et al., 2022; Tu et al., 2017; Wang et al., 2021). The predominant metabolic functions identified are glutamate metabolism, denitrification and nitrate assimilation/dissimilation, as in plants, rivers and oceans (Deng et al., 2024; Nie et al., 2021; Song et al., 2022). In these environments, these processes can account for up to 80 % of the genes associated with inorganic nitrogen utilization, versus ∼ 90 % in the atmosphere in our study.
Active nitrogen-related functions are maintained by bacteria in the atmosphere, which might be critical for their physiology despite much shorter residence times (a few days) than in any other environments (Burrows et al., 2009). Surprisingly, higher gene expression levels (RNA:DNA ratio) of these functions were observed in clear air than in clouds, where the presence of condensed water is rather expected to promote biological activity. Liquid water can be retained by efflorescent aerosol particless at RH < 100 % (Cruz and Pandis, 2000), which could be sufficient to sustain biological activity in clear atmosphere. In addition, it is possible that multiple dry-wet cycles occurred, in particular, before collecting the clear-air samples, which could have contributed to enhance biological activity as compared with cloud water, as observed in soil (Xiang et al., 2008).
Bacterial diversity associated with nitrogen-related processes is composed of a core of taxa, which are recurrently present and active regardless of atmospheric conditions (clear atmosphere or cloud), and also predominant in surface ecosystems (freshwater, ocean, phyllosphere) (Deng et al., 2024; Nie et al., 2021; Song et al., 2022). This is made up of six phyla: Pseudomonadota, Acidobacteriota, Actinomycetota, Bacteroidota, Bacillota, and Planctomycetota, which contrasts with other ecosystems (plants, river etc.) where nitrogen-associated processes are typically dominated by a single bacterial phylum or order (>50 %–75 % relative abundance) (Deng et al., 2024; Song et al., 2022; Tu et al., 2017). Functions carried out by more diverse assemblages of microorganisms are less sensitive to environmental changes and therefore more stable over time (Maron et al., 2018).
In contrast, certain functions and their associated taxa have very low representation in the atmosphere, such as nitrification and anammox, usually carried out by Nitrososphaerales, Nitrosomonadales and/or Nitrospirales. These occur mostly in sediments from the ocean's mesopelagic zones (Deng et al., 2024; Song et al., 2022), i.e., in areas that are not in contact with the atmosphere.
4.2 Focus on the main nitrogen elements and processes
4.2.1 Organic nitrogen content in the biomass of airborne bacteria
In total, 5 × 1019 bacteria (Whitman et al., 1998) are estimated aloft in the Earth's atmosphere, roughly contributing ∼ 500 kg of N, considering an average N content of 10−14 g per cell (Paul and Clark, 1996). According to the study by Neff et al. (2002), the annual flux of atmospheric organic nitrogen emitted as bacterial biomass would be 9.5 × 10−6 kg of N ha−2. Considering only the continents (1489 × 1010 ha), we infer that this represents ∼ 105 kg N yr−1. In another study, the total number of bacteria emitted annually into the atmosphere from surface ecosystems is estimated at ∼ 1024 cells (Burrows et al., 2009), corresponding to approximately 107 kg N yr−1 carried as bacterial biomass. In clouds, organic nitrogen compounds represent 18 % of the total molar content of nitrogen present (Hill et al., 2007). It was estimated that bacterial biomass represents less than 1 % of it (Hill et al., 2007).
Therefore, overall, the contribution of airborne bacteria to atmospheric N, between 105 and 107 kg N yr−1 according to these rough estimates, remains minor compared to anthropogenic or natural inorganic nitrogen emissions, with ∼ 70 Tg NH3 yr−1, ∼ 18 Tg N2O yr−1, and ∼ 5 Tg NO yr−1 (Bouwman et al., 2013; Duce et al., 2008; Fowler et al., 2013; Pilegaard, 2013; Sutton et al., 2013; Voss et al., 2011).
4.2.2 NH utilization and glutamate metabolism
In atmospheric MG and MT, the sequences related to glutamate metabolism prevail among N-related processes. Although the inorganic forms largely predominate, organic nitrogen therefore likely represents a major source of nitrogen and energy for airborne microorganisms (Jaber et al., 2021).
Ammonium/ammonia is one of the most abundant reactive nitrogen compounds in the atmosphere, with ammonium concentrations of up to several hundred micromolar in cloud water (Hill et al., 2007; Renard et al., 2020; Vaïtilingom et al., 2010). Emissions from surfaces amount to ∼ 70 Tg NH3 yr−1, mainly from industry and agriculture (Bouwman et al., 2013; Duce et al., 2008; Fowler et al., 2013; Sutton et al., 2013).
Based on our data, assuming that our samples are representative of the global atmosphere, we estimate a global ammonium processing by bacteria in clouds of ∼ 5.5 × 107 kg yr−1, considering the average bioassimilation rate measured in rainwater incubations (4.61 × 10−8 µmol cell−1 h−1), a total airborne biomass of 5 × 1019 bacteria (Whitman et al., 1998), and a cloud fraction of 15 % (Lelieveld and Crutzen, 1990). This represents approximately ∼ 0.08 % of NH3 emissions from terrestrial ecosystems and oceans (Bouwman et al., 2013; Duce et al., 2008; Fowler et al., 2013; Sutton et al., 2013). The positive correlation between the proportion of Sphingomonadales and ammonium decay rate in the incubations suggests a direct impact of these bacteria.
The biosynthesis of glutamate is the primary pathway for the assimilation of nitrogen and this requires NH (Merrick and Edwards, 1995). This process involves glutamine synthesis via the enzyme glutamine synthetase (GS), which catalyzes an ATP-dependent amidation of glutamate, utilizing ammonium as the nitrogen source (Walker and van der Donk, 2016). Glutamine then acts as a nitrogen donor for approximately half of the nitrogen atoms incorporated into purines and pyrimidines, whereas glutamate provides nitrogen atoms for these nucleic acid bases, as well as amino groups for all other amino acids. Therefore, the GS-Glutamate synthase biosynthesis pathway, also known as the GS-GOGAT pathway, constitutes a major route for nitrogen assimilation from ammonium (Walker and van der Donk, 2016).
4.2.3 Nitrate utilization (NO)
After glutamate/ammonium metabolism, denitrification (mainly nitrate reduction) and nitrate assimilation and dissimilation are the most represented processes in atmospheric MG and MT. Like ammonium, nitrate is also usually present at concentrations up to several hundred micromolar in cloud water (Hill et al., 2007; Renard et al., 2020; Vaïtilingom et al., 2010).This is a major target of microbial processes as well in ecosystems. Nitrate concentration decreased in ∼ 40 % of the rainwater incubation experiments. Extrapolating these observations to the global scale, this corresponds to the transformation of ∼ 2 × 107 kg yr−1 of NO in the atmosphere on a planetary scale, which is 47 % less than predicted for ammonium (Sect. 4.2.2). In all experiments, ammonium concentration decreased simultaneously to that of nitrate. Ammonium is preferentially used by microorganisms to recover nitrogen, over nitrates (McCarthy et al., 1977). The ability to respire nitrate is widely spread among bacteria that can dissimilate the produced nitrite into gaseous compounds (denitrification) for energy production, or to ammonium (dissimilatory nitrate reduction) for nitrogen assimilation (Philippot and Højberg, 1999).
Denitrification is a process that generally occurs under oxygen-limited or anoxic conditions. However, some microorganisms are capable of performing aerobic denitrification (AD). Aerobic denitrifying bacteria are predominantly Gram-negative bacteria affiliated with Pseudomonadota, with nearly 50 % of them belonging to Pseudomonas (Ji et al., 2015), a taxon frequent in the atmosphere (Vaïtilingom et al., 2012). In addition, yeasts with AD capacity were reported from surface sediments (Fang et al., 2021; Zeng et al., 2020), as well as Actinomycetes from aquatic ecosystems (Ma et al., 2022). In AD, O2 and NO compete for electrons, with O2 being thermodynamically favored. The O2 concentration regulates enzymatic activity and therefore the efficiency of denitrification, with three possible patterns observed across different microorganisms: (i) efficiency decreases with increasing dissolved oxygen (DO) until a threshold is reached (Wilson and Bouwer, 1997), (ii) efficiency is optimal only within a specific DO concentration range (Chen and Ni, 2012) and (iii) rare tolerance to high DO concentrations (Ji et al., 2014; Zhang et al., 2011). At present, there is no consensus on the mechanisms underlying AD, but several theories have been proposed (Hao et al., 2022). The first is the microenvironmental theory, where oxygen diffusion is limited in cell aggregates. The second is the enzyme theory, which attributes aerobic denitrification to the activity of specific enzymes. For example, in Thiosphaera pantotropha, two nitrate reductases coexist: M-NAR (active only in the absence of O2) and P-NAR (active even in the presence of O2). This dual capacity allows the cell to reduce both oxygen and nitrate simultaneously, making denitrification possible under aerobic conditions (Kumar and Lin, 2010; Yang et al., 2020). The third theory, which is not mutually exclusive with the enzyme theory, is the electron transfer theory. It explains AD as the result of a bottleneck in the respiratory chain that prevents all electrons from being transferred to O2. Instead, some electrons are redirected to denitrification enzymes, enabling the simultaneous use of O2 and NO as electron acceptors (Chen et al., 2006; Kong et al., 2006; Robertson and Kuenen, 1984). The efficiency of this process depends on energy demand, O2 concentration, and the presence of specific enzymes such as Nap, NAR, and NIR.
4.2.4 Biological nitrogen fixation (N2)
The third most represented function in inorganic nitrogen processes is biological nitrogen fixation that converts molecular nitrogen to ammonia. The corresponding biomarker genes (nifDHKW) represent ∼ 10 % of all genes associated with the atmospheric inorganic nitrogen cycle in MG and MT. Moreover, the screening of isolates indicates that this function is not rare in Pseudomonadota viable in the atmosphere. N2 is the most abundant inorganic nitrogen form in the atmosphere. Biological nitrogen fixation transcripts were found more represented in clear atmosphere conditions than in clouds.
Nitrogen fixation is an energy-demanding process, but it enables microorganisms capable of performing it to assimilate atmospheric nitrogen for biomass production. Microbes only activate N2 fixation when they lack access to more readily assimilable nitrogen sources, such as ammonium. This situation can occur under conditions of limited bioavailable nitrogen which is supported by indications of amino-acid starvation in metatranscriptomes (Péguilhan et al., 2025). One could envision that the atmospheric environments serve as niches for nitrogen fixers: while certain bacteria grow rapidly by using available compounds such as ammonium, diazotrophs that are typically growing more slowly can subsequently, or in parallel, develop in nitrogen-limited environments.
In some rain incubation experiments, we observed bacteria multiplication but no decrease of ammonium or nitrate concentration, so other source(s) of nitrogen, such as N2 and/or organic forms supported bacterial growth. In these cases, Burkholderiales dominated the bacteria community; this order is known to include nitrogen-fixing bacteria (Bahulikar et al., 2021; Jean et al., 2020), including anoxygenic phototrophs, i.e. photosynthetic bacteria using light to produce energy, with no production of molecular oxygen (Imhoff et al., 2019; Vergne et al., 2021). In these bacteria, phototrophy supports atmospheric nitrogen fixation (through the enzyme nitrogenase), one of the most energy-intensive reactions in living organisms (Kim and Rees, 1994). These microorganisms do not use water as the electron donor but instead exploit a variety of reduced organic (e.g., organic acids) or inorganic compounds such as Fe2+, H2, HS−, S2O, NO, and AsO (Trüper and Pfennig, 1981). While potential electron donors such as H2S and thiosulfate are relatively scarce (<0.1–1 ppbv), H2, whose mixing ratio is around 500 ppbv, along with organic compounds such as formic and acetic acids, could support anoxygenic phototrophy, and nitrogen fixation from them. A study conducted in an atmospheric simulation chamber using a cloud-isolated Methylobacterium sp. demonstrated that this auxiliary metabolism enhances bacterial fitness during atmospheric transport and may facilitate subsequent environmental colonization (Mathonat et al., 2025).
In our work, attempts to quantify N2 fixation using acetylene reduction assay (ARA) (Hardy et al., 1968) failed, likely because of the limitations due to low microbial biomass. Considering that half of the airborne bacteria (5 × 1019 cells) are Pseudomonadota, and that 15 % are diazotroph (theoretical upper-bound estimate), biological N2 fixation in the atmosphere would account for approximately 0.018 Tg N2 yr−1, assuming an average nitrogen fixation rate of 10 fmol N2 cell−1 h−1 by a marine diazotroph (Foster et al., 2022). Although a more precise quantification would be needed, this upper estimation indicates that biological nitrogen fixation in clouds likely represents an insignificant fraction of global biological nitrogen fixation compared to estimates for terrestrial (58 Tg N2 year−1 for natural ecosystems and 60 Tg N2 yr−1 for agricultural crops (Herridge et al., 2008)) or marine systems (140 Tg N2 yr−1 (Voss et al., 2013)). However, this may support microbial maintenance during atmospheric transport.
4.3 Perspectives
Atmospheric bacteria assimilate and transform more nitrogen than they contribute through their biomass, suggesting that they may act as sinks of atmospheric nitrogen. However, the accuracy of these transformation estimates at global scale could be improved. For instance, the measured bioassimilation rates were obtained from rainwater incubations at 17 °C, which do not fully represent atmospheric aerosol conditions, as the liquid bulk phase does not account for the particulate nature of aerosols, and the temperature is relatively high compared to actual atmospheric conditions. Atmospheric regions such as the cloud transition zone (i.e., the interface between clouds and the free atmosphere) exhibit humidity levels close to saturation and a broader atmospheric coverage than previously estimated (Calbó et al., 2024; Ruiz de Morales et al., 2024). This region may be particularly favorable for bacterial activity, as it is expected to experience longer atmospheric residence times and higher temperatures than those prevailing within clouds.
Water availability is another important factor to consider. While ammonium and nitrate assimilation was extrapolated to cloud environments based on rainwater data, it is important to note that even under non-cloud conditions, aerosol particles are not entirely devoid of liquid water (Ervens et al., 2025; Pandis and Seinfeld, 1989), making nitrogen assimilation potentially feasible in clear atmosphere conditions as well.
Moreover, while these estimates are scaled to the global level, atmospheric conditions and stressors, nitrogen speciation and concentrations and microbial diversity and abundance are subject to strong spatial and temporal variability. Hence, nitrogen-related biological processes are expected to be highly variable in space and time as well. Our estimates, which are based on samples collected on a single location, for limited periods of time, and derived from laboratory incubations under defined fixed conditions are therefore associated with high uncertainties. Addressing variations and heterogeneity of these processes will require more investigations. One could envision enhanced microbial nitrogen cycling activity following major nitrogen release events (e.g., fertilizer application in agriculture). Conversely, such nitrogen “overloads” might induce nitrosative stress, potentially reaching toxic levels for airborne microorganisms. This is supported by chamber studies showing that high NO and N2O concentrations can reduce bacterial culturability (Vernocchi et al., 2023).
This study shows that atmospheric microorganisms are potential contributors to the atmospheric nitrogen cycle. Using a combination of metagenomic, metatranscriptomic, and experimental approaches, we showed that atmospheric microorganisms exhibit genes associated with nitrogen cycling, including both inorganic and organic nitrogen pathways, with near-equal representation (∼ 50:50), similar to other environments such as oceans, rivers, and plant-associated microbiomes. Denitrification, nitrate assimilation/dissimilation, and glutamate metabolism emerge as the dominant functional pathways, both in terms of genetic presence and transcriptional activity, highlighting the metabolic versatility of airborne microbial communities whether in a cloud or free atmosphere conditions, noting a functional resilience of these N-processing.
Rainwater incubations showed bioassimilation of ammonium and, to a lesser extent, nitrate with an important link to the microbial diversity. The microbial community composition plays a significant role in nitrogen transformation potential. Atmospheric back-trajectories provide contextual information and help linking air mass origin (marine vs. continental) to both concentrations of nitrogen-containing ions and microbial community composition in precipitation. For example, we show that air masses of continental origin carry higher levels of ammonium and nitrate and are associated with microbial communities enriched in taxa such as Burkholderiales, thereby influencing nitrogen transformation dynamics. Overall, our findings suggest that atmospheric microbial communities, though transient and exposed to harsh conditions, maintain stable and functionally resilient nitrogen-cycling capabilities.
The sequence data of 16S rRNA gene amplicons have been deposited in the European Nucleotide Archive (ENA) at EMBL-EBI under the accession number PRJEB91356.
The supplement related to this article is available online at https://doi.org/10.5194/bg-23-2885-2026-supplement.
FM, PA, MJ designed the experiments. FM performed the experiments. FM and PA collected samples and data. FM, RP and FE conducted the bioinformatics analysis. J-LB performed the backward trajectory analysis. FM and PA wrote the paper. All authors have contributed to the editing and formatting of figures and text. PA and BE supervised the research, coordination and strategy.
At least one of the (co-)authors is a member of the editorial board of Biogeosciences. The peer-review process was guided by an independent editor, and the authors also have no other competing interests to declare.
Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims made in the text, published maps, institutional affiliations, or any other geographical representation in this paper. The authors bear the ultimate responsibility for providing appropriate place names. Views expressed in the text are those of the authors and do not necessarily reflect the views of the publisher.
We are grateful for the support and facilities provided by Guillaume Voyard at the ion chromatography platform at the Clermont-Ferrand chemistry institute (ICCF, France). We also thank Jonathan Colombet and Hermine Billard for their help with the cytometry platform at the Laboratoire Microorganismes: Genome Environnement (LMGE, France). CO-PDD is an instrumented site of the OPGC observatory and LaMP laboratory supported by the Université Clermont Auvergne (UCA), by the Centre National de la Recherche Scientifique (CNRS-INSU), and by the Centre National d'Etudes Spatiales (CNES) for collecting of atmospheric samples. We would also like to thank Julien Pottier of the INRAE UREP for his help with the data collection of the meteorological variable.
FM was supported by a PhD fellowship from the French Ministry of Education and Research. The authors also acknowledge funding from the Fédération des Recherches en Environnement of Clermont-Ferrand, ClermontAuvergne University, and CNRS.
This paper was edited by Tina Šantl-Temkiv and reviewed by Kai Finster and one anonymous referee.
Aber, J. D., Magill, A., Mcnulty, S. G., Boone, R. D., Nadelhoffer, K. J., Downs, M., and Hallett, R.: Forest biogeochemistry and primary production altered by nitrogen saturation, Water Air. Soil Pollut., 85, 1665–1670, https://doi.org/10.1007/BF00477219, 1995.
Almaraz, M., Bai, E., Wang, C., Trousdell, J., Conley, S., Faloona, I., and Houlton, B. Z.: Agriculture is a major source of NOx pollution in California, Sci. Adv., 4, eaao3477, https://doi.org/10.1126/sciadv.aao3477, 2018.
Amato, P., Parazols, M., Sancelme, M., Laj, P., Mailhot, G., and Delort, A.-M.: Microorganisms isolated from the water phase of tropospheric clouds at the Puy de Dôme: major groups and growth abilities at low temperatures, FEMS Microbiol. Ecol., 59, 242–254, https://doi.org/10.1111/j.1574-6941.2006.00199.x, 2007.
Amato, P., Joly, M., Besaury, L., Oudart, A., Taib, N., Moné, A. I., Deguillaume, L., Delort, A.-M., and Debroas, D.: Active microorganisms thrive among extremely diverse communities in cloud water, PLOS ONE, 12, e0182869, https://doi.org/10.1371/journal.pone.0182869, 2017.
Amato, P., Besaury, L., Joly, M., Penaud, B., Deguillaume, L., and Delort, A.-M.: Metatranscriptomic exploration of microbial functioning in clouds, Sci. Rep., 9, 4383, https://doi.org/10.1038/s41598-019-41032-4, 2019.
Apprill, A., Mcnally, S., Parsons, R., and Weber, L.: Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton, Aquat. Microb. Ecol., 75, 129–137, https://doi.org/10.3354/ame01753, 2015.
Atkinson, R., Winer, A. M., and Pitts, J. N.: Estimation of night-time N2O5 concentrations from ambient NO2 and NO3 radical concentrations and the role of N2O5 in night-time chemistry, Atmos. Environ., 20, 331–339, https://doi.org/10.1016/0004-6981(86)90035-1, 1986.
Bach, H.-J., Tomanova, J., Schloter, M., and Munch, J. C.: Enumeration of total bacteria and bacteria with genes for proteolytic activity in pure cultures and in environmental samples by quantitative PCR mediated amplification, J. Microbiol. Meth., 49, 235–245, https://doi.org/10.1016/S0167-7012(01)00370-0, 2002.
Bahulikar, R. A., Chaluvadi, S. R., Torres-Jerez, I., Mosali, J., Bennetzen, J. L., and Udvardi, M.: Nitrogen Fertilization Reduces Nitrogen Fixation Activity of Diverse Diazotrophs in Switchgrass Roots, Phytobiomes J., 5, 80–87, https://doi.org/10.1094/PBIOMES-09-19-0050-FI, 2021.
Baldrian, P., Kolařík, M., Štursová, M., Kopecký, J., Valášková, V., Větrovský, T., Žifčáková, L., Šnajdr, J., Rídl, J., Vlček, Č., and Voříšková, J.: Active and total microbial communities in forest soil are largely different and highly stratified during decomposition, ISME J., 6, 248–258, https://doi.org/10.1038/ismej.2011.95, 2012.
Baray, J.-L., Deguillaume, L., Colomb, A., Sellegri, K., Freney, E., Rose, C., Van Baelen, J., Pichon, J.-M., Picard, D., Fréville, P., Bouvier, L., Ribeiro, M., Amato, P., Banson, S., Bianco, A., Borbon, A., Bourcier, L., Bras, Y., Brigante, M., Cacault, P., Chauvigné, A., Charbouillot, T., Chaumerliac, N., Delort, A.-M., Delmotte, M., Dupuy, R., Farah, A., Febvre, G., Flossmann, A., Gourbeyre, C., Hervier, C., Hervo, M., Huret, N., Joly, M., Kazan, V., Lopez, M., Mailhot, G., Marinoni, A., Masson, O., Montoux, N., Parazols, M., Peyrin, F., Pointin, Y., Ramonet, M., Rocco, M., Sancelme, M., Sauvage, S., Schmidt, M., Tison, E., Vaïtilingom, M., Villani, P., Wang, M., Yver-Kwok, C., and Laj, P.: Cézeaux-Aulnat-Opme-Puy De Dôme: a multi-site for the long-term survey of the tropospheric composition and climate change, Atmos. Meas. Tech., 13, 3413–3445, https://doi.org/10.5194/amt-13-3413-2020, 2020.
Bari, A., Ferraro, V., Wilson, L. R., Luttinger, D., and Husain, L.: Measurements of gaseous HONO, HNO3, SO2, HCl, NH3, particulate sulfate and PM2.5 in New York, NY, Atmos. Environ., 37, 2825–2835, https://doi.org/10.1016/S1352-2310(03)00199-7, 2003.
Barnes, N. M. and Wu, H.: Mechanisms regulating the airborne survival of Klebsiella pneumoniae under different relative humidity and temperature levels, Indoor Air, 32, e12991, https://doi.org/10.1111/ina.12991, 2022.
Berks, B. C., Ferguson, S. J., Moir, J. W. B., and Richardson, D. J.: Enzymes and associated electron transport systems that catalyse the respiratory reduction of nitrogen oxides and oxyanions, Biochim. Biophys. Acta BBA – Bioenerg., 1232, 97–173, https://doi.org/10.1016/0005-2728(95)00092-5, 1995.
Bernhard, A.: The nitrogen cycle: processes, players, and human impact, Nat. Educ. Knowl., 3, 25, https://www.nature.com/scitable/knowledge/library/the-nitrogen-cycle-processes-players-and-human-15644632/ (last access: April 2026), 2010.
Blasco, F., Iobbi, C., Ratouchniak, J., Bonnefoy, V., and Chippaux, M.: Nitrate reductases of Escherichia coli: Sequence of the second nitrate reductase and comparison with that encoded by the narGHJI operon, Mol. Gen. Genet. MGG, 222, 104–111, https://doi.org/10.1007/BF00283030, 1990.
Bouwman, L., Goldewijk, K. K., Van Der Hoek, K. W., Beusen, A. H. W., Van Vuuren, D. P., Willems, J., Rufino, M. C., and Stehfest, E.: Exploring global changes in nitrogen and phosphorus cycles in agriculture induced by livestock production over the 1900–2050 period, P. Natl. Acad. Sci. USA, 110, 20882–20887, https://doi.org/10.1073/pnas.1012878108, 2013.
Braker, G. and Tiedje, J. M.: Nitric Oxide Reductase (norB) Genes from Pure Cultures and Environmental Samples, Appl. Environ. Microbiol., 69, 3476–3483, https://doi.org/10.1128/AEM.69.6.3476-3483.2003, 2003.
Braker, G., Zhou, J., Wu, L., Devol, A. H., and Tiedje, J. M.: Nitrite Reductase Genes (nirK andnirS) as Functional Markers To Investigate Diversity of Denitrifying Bacteria in Pacific Northwest Marine Sediment Communities, Appl. Environ. Microbiol., 66, 2096–2104, https://doi.org/10.1128/AEM.66.5.2096-2104.2000, 2000.
Bulgarelli, D., Rott, M., Schlaeppi, K., Ver Loren van Themaat, E., Ahmadinejad, N., Assenza, F., Rauf, P., Huettel, B., Reinhardt, R., Schmelzer, E., Peplies, J., Gloeckner, F. O., Amann, R., Eickhorst, T., and Schulze-Lefert, P.: Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota, Nature, 488, 91–95, https://doi.org/10.1038/nature11336, 2012.
Burrows, S. M., Butler, T., Jöckel, P., Tost, H., Kerkweg, A., Pöschl, U., and Lawrence, M. G.: Bacteria in the global atmosphere – Part 2: Modeling of emissions and transport between different ecosystems, Atmos. Chem. Phys., 9, 9281–9297, https://doi.org/10.5194/acp-9-9281-2009, 2009.
Calbó, J., González, J.-A., Jahani, B., Sola, Y., and Morales, J. R. de: How important is the transition zone between clouds and aerosol?, AIP Conf. Proc., 2988, 070005, https://doi.org/10.1063/5.0182769, 2024.
Cantera, J. J. L. and Stein, L. Y.: Molecular diversity of nitrite reductase genes (nirK) in nitrifying bacteria, Environ. Microbiol., 9, 765–776, https://doi.org/10.1111/j.1462-2920.2006.01198.x, 2007.
Cape, J. N., Hargreaves, K. J., Storeton-West, R., Fowler, D., Colville, R. N., Choularton, T. W., and Gallagher, M. W.: Nitrite in orographic cloud as an indicator of nitrous acid in rural air, Atmos. Environ., Part Gen. Top., 26, 2301–2307, https://doi.org/10.1016/0960-1686(92)90361-N, 1992.
Cape, J. N., Cornell, S. E., Jickells, T. D., and Nemitz, E.: Organic nitrogen in the atmosphere – Where does it come from? A review of sources and methods, Atmos. Res., 102, 30–48, https://doi.org/10.1016/j.atmosres.2011.07.009, 2011.
Chen, F., Xia, Q., and Ju, L.-K.: Competition between oxygen and nitrate respirations in continuous culture of Pseudomonas aeruginosa performing aerobic denitrification, Biotechnol. Bioeng., 93, 1069–1078, https://doi.org/10.1002/bit.20812, 2006.
Chen, Q. and Ni, J.: Ammonium removal by Agrobacterium sp. LAD9 capable of heterotrophic nitrification–aerobic denitrification, J. Biosci. Bioeng., 113, 619–623, https://doi.org/10.1016/j.jbiosc.2011.12.012, 2012.
Cowling, E. B., Erisman, J. W., Smeulders, S. M., Holman, S. C., and Nicholson, B. M.: Optimizing air quality management in Europe and North America: Justification for integrated management of both oxidized and reduced forms of nitrogen, Environ. Pollut., 102, 599–608, https://doi.org/10.1016/S0269-7491(98)80088-2, 1998.
Cox, C. S. and Goldberg, L. J.: Aerosol Survival of Pasteurella tularensis and the Influence of Relative Humidity, Appl. Microbiol., 23, 1–3, https://doi.org/10.1128/am.23.1.1-3.1972, 1972.
Coyle, C. L., Zumft, W. G., Kroneck, P. M. H., Körner, H., and Jakob, W.: Nitrous oxide reductase from denitrifying, Eur. J. Biochem., 153, 459–467, https://doi.org/10.1111/j.1432-1033.1985.tb09324.x, 1985.
Cruz, C. N. and Pandis, S. N.: Deliquescence and Hygroscopic Growth of Mixed Inorganic-Organic Atmospheric Aerosol, Environ. Sci. Technol., 34, 4313–4319, https://doi.org/10.1021/es9907109, 2000.
Decho, A. W. and Lopez, G. R.: Exopolymer microenvironments of microbial flora: Multiple and interactive effects on trophic relationships, Limnol. Oceanogr., 38, 1633–1645, https://doi.org/10.4319/lo.1993.38.8.1633, 1993.
Deguillaume, L., Charbouillot, T., Joly, M., Vaïtilingom, M., Parazols, M., Marinoni, A., Amato, P., Delort, A.-M., Vinatier, V., Flossmann, A., Chaumerliac, N., Pichon, J. M., Houdier, S., Laj, P., Sellegri, K., Colomb, A., Brigante, M., and Mailhot, G.: Classification of clouds sampled at the puy de Dôme (France) based on 10 yr of monitoring of their physicochemical properties, Atmos. Chem. Phys., 14, 1485–1506, https://doi.org/10.5194/acp-14-1485-2014, 2014.
Deng, D., Yang, Z., Yang, Y., Wan, W., Liu, W., and Xiong, X.: Metagenomic insights into nitrogen-cycling microbial communities and their relationships with nitrogen removal potential in the Yangtze River, Water Res., 265, 122229, https://doi.org/10.1016/j.watres.2024.122229, 2024.
Diaz, R. J. and Rosenberg, R.: Spreading Dead Zones and Consequences for Marine Ecosystems, Science, 321, 926–929, https://doi.org/10.1126/science.1156401, 2008.
Duce, R. A., LaRoche, J., Altieri, K., Arrigo, K. R., Baker, A. R., Capone, D. G., Cornell, S., Dentener, F., Galloway, J., Ganeshram, R. S., Geider, R. J., Jickells, T., Kuypers, M. M., Langlois, R., Liss, P. S., Liu, S. M., Middelburg, J. J., Moore, C. M., Nickovic, S., Oschlies, A., Pedersen, T., Prospero, J., Schlitzer, R., Seitzinger, S., Sorensen, L. L., Uematsu, M., Ulloa, O., Voss, M., Ward, B., and Zamora, L.: Impacts of Atmospheric Anthropogenic Nitrogen on the Open Ocean, Science, 320, 893–897, https://doi.org/10.1126/science.1150369, 2008.
Dworkin, M., Falkow, S., Rosenberg, E., Schleifer, K.-H., and Stackebrandt, E. (Eds.): The Prokaryotes, Springer New York, New York, NY, https://doi.org/10.1007/0-387-30742-7, 2006.
Ervens, B. and Amato, P.: The global impact of bacterial processes on carbon mass, Atmos. Chem. Phys., 20, 1777–1794, https://doi.org/10.5194/acp-20-1777-2020, 2020.
Ervens, B., Amato, P., Aregahegn, K., Joly, M., Khaled, A., Labed-Veydert, T., Mathonat, F., Nuñez López, L., Péguilhan, R., and Zhang, M.: Ideas and perspectives: Microorganisms in the air through the lenses of atmospheric chemistry and microphysics, Biogeosciences, 22, 243–256, https://doi.org/10.5194/bg-22-243-2025, 2025.
Escudié, F., Auer, L., Bernard, M., Mariadassou, M., Cauquil, L., Vidal, K., Maman, S., Hernandez-Raquet, G., Combes, S., and Pascal, G.: FROGS: Find, Rapidly, OTUs with Galaxy Solution, Bioinformatics, 34, 1287–1294, https://doi.org/10.1093/bioinformatics/btx791, 2018.
Espey, M. G., Miranda, K. M., Feelisch, M., Fukuto, J., Grisham, M. B., Vitek, M. P., and Wink, D. A.: Mechanisms of Cell Death Governed by the Balance between Nitrosative and Oxidative Stress, Ann. N. Y. Acad. Sci., 899, 209–221, https://doi.org/10.1111/j.1749-6632.2000.tb06188.x, 2000.
Fang, J., Liao, S., Zhang, S., Li, L., Tan, S., Li, W., Wang, A., and Ye, J.: Characteristics of a novel heterotrophic nitrification-aerobic denitrification yeast, Barnettozyma californica K1, Bioresour. Technol., 339, 125665, https://doi.org/10.1016/j.biortech.2021.125665, 2021.
Fani, R., Gallo, R., and Liò, P.: Molecular Evolution of Nitrogen Fixation: The Evolutionary History of the nifD, nifK, nifE, and nifN Genes, J. Mol. Evol., 51, 1–11, https://doi.org/10.1007/s002390010061, 2000.
Foster, R. A., Tienken, D., Littmann, S., Whitehouse, M. J., Kuypers, M. M. M., and White, A. E.: The rate and fate of N2 and C fixation by marine diatom-diazotroph symbioses, ISME J., 16, 477–487, https://doi.org/10.1038/s41396-021-01086-7, 2022.
Fowler, D., Coyle, M., Skiba, U., Sutton, M. A., Cape, J. N., Reis, S., Sheppard, L. J., Jenkins, A., Grizzetti, B., Galloway, J. N., Vitousek, P., Leach, A., Bouwman, A. F., Butterbach-Bahl, K., Dentener, F., Stevenson, D., Amann, M., and Voss, M.: The global nitrogen cycle in the twenty-first century, Philos. Trans. R. Soc. B Biol. Sci., https://doi.org/10.1098/rstb.2013.0164, 2013.
Fröhlich-Nowoisky, J., Kampf, C. J., Weber, B., Huffman, J. A., Pöhlker, C., Andreae, M. O., Lang-Yona, N., Burrows, S. M., Gunthe, S. S., Elbert, W., Su, H., Hoor, P., Thines, E., Hoffmann, T., Després, V. R., and Pöschl, U.: Bioaerosols in the Earth system: Climate, health, and ecosystem interactions, Atmos. Res., 182, 346–376, https://doi.org/10.1016/j.atmosres.2016.07.018, 2016.
Galimand, M., Gamper, M., Zimmermann, A., and Haas, D.: Positive FNR-like control of anaerobic arginine degradation and nitrate respiration in Pseudomonas aeruginosa, J. Bacteriol., 173, 1598–1606, https://doi.org/10.1128/jb.173.5.1598-1606.1991, 1991.
Galloway, J. N., Cowling, E. B., Seitzinger, S. P., and Socolow, R. H.: Reactive Nitrogen: Too Much of a Good Thing?, Ambio, 31, 60–63, 2002.
Gregory, L. G., Karakas-Sen, A., Richardson, D. J., and Spiro, S.: Detection of genes for membrane-bound nitrate reductase in nitrate-respiring bacteria and in community DNA, FEMS Microbiol. Lett., 183, 275–279, https://doi.org/10.1111/j.1574-6968.2000.tb08971.x, 2000.
Hammer, O., Harper, D. A. T., and Ryan, P. D.: PAST: Paleontological Statistics Software Package for Education and Data Analysis, https://paleo.carleton.ca/2001_1/past/past.pdf (last access: January 2026), 2001.
Hao, Z.-L., Ali, A., Ren, Y., Su, J.-F., and Wang, Z.: A mechanistic review on aerobic denitrification for nitrogen removal in water treatment, Sci. Total Environ., 847, 157452, https://doi.org/10.1016/j.scitotenv.2022.157452, 2022.
Hardy, R. W. F., Holsten, R. D., Jackson, E. K., and Burns, R. C.: The Acetylene-Ethylene Assay for N2 Fixation: Laboratory and Field Evaluation 1, Plant Physiol., 43, 1185–1207, https://doi.org/10.1104/pp.43.8.1185, 1968.
Hargreaves, K. J., Fowler, D., Storeton-West, R. L., and Duyzer, J. H.: The exchange of nitric oxide, nitrogen dioxide and ozone between pasture and the atmosphere, Environ. Pollut., 75, 53–59, https://doi.org/10.1016/0269-7491(92)90056-G, 1992.
Heiss, B., Frunzke, K., and Zumft, W. G.: Formation of the N-N bond from nitric oxide by a membrane-bound cytochrome bc complex of nitrate-respiring (denitrifying) Pseudomonas stutzeri, J. Bacteriol., 171, 3288–3297, https://doi.org/10.1128/jb.171.6.3288-3297.1989, 1989.
Herridge, D. F., Peoples, M. B., and Boddey, R. M.: Global inputs of biological nitrogen fixation in agricultural systems, Plant Soil, 311, 1–18, https://doi.org/10.1007/s11104-008-9668-3, 2008.
Hill, K. A., Shepson, P. B., Galbavy, E. S., Anastasio, C., Kourtev, P. S., Konopka, A., and Stirm, B. H.: Processing of atmospheric nitrogen by clouds above a forest environment, J. Geophys. Res. Atmospheres, 112, https://doi.org/10.1029/2006JD008002, 2007.
Hollocher, T. C., Tate, M. E., and Nicholas, D. J.: Oxidation of ammonia by Nitrosomonas europaea, in: Definite 18O-tracer evidence that hydroxylamine formation involves a monooxygenase, J. Biol. Chem., 256, 10834–10836, https://doi.org/10.1016/S0021-9258(19)68518-2, 1981.
Hommes, N. G., Sayavedra-Soto, L. A., and Arp, D. J.: Transcript Analysis of Multiple Copies ofamo (Encoding Ammonia Monooxygenase) and hao(Encoding Hydroxylamine Oxidoreductase) in Nitrosomonas europaea, J. Bacteriol., 183, 1096–1100, https://doi.org/10.1128/jb.183.3.1096-1100.2001, 2001.
Howarth, R. W.: Coastal nitrogen pollution: A review of sources and trends globally and regionally, Harmful Algae, 8, 14–20, https://doi.org/10.1016/j.hal.2008.08.015, 2008.
Imhoff, J. F., Rahn, T., Künzel, S., and Neulinger, S. C.: Phylogeny of Anoxygenic Photosynthesis Based on Sequences of Photosynthetic Reaction Center Proteins and a Key Enzyme in Bacteriochlorophyll Biosynthesis, the Chlorophyllide Reductase, Microorganisms, 7, 576, https://doi.org/10.3390/microorganisms7110576, 2019.
Jaber, S., Joly, M., Brissy, M., Leremboure, M., Khaled, A., Ervens, B., and Delort, A.-M.: Biotic and abiotic transformation of amino acids in cloud water: experimental studies and atmospheric implications, Biogeosciences, 18, 1067–1080, https://doi.org/10.5194/bg-18-1067-2021, 2021.
Jean, M., Holland-Moritz, H., Melvin, A. M., Johnstone, J. F., and Mack, M. C.: Experimental assessment of tree canopy and leaf litter controls on the microbiome and nitrogen fixation rates of two boreal mosses, New Phytol., 227, 1335–1349, https://doi.org/10.1111/nph.16611, 2020.
Ji, B., Wang, H., and Yang, K.: Tolerance of an aerobic denitrifier (Pseudomonas stutzeri) to high O2 concentrations, Biotechnol. Lett., 36, 719–722, https://doi.org/10.1007/s10529-013-1417-x, 2014.
Ji, B., Yang, K., Zhu, L., Jiang, Y., Wang, H., Zhou, J., and Zhang, H.: Aerobic denitrification: A review of important advances of the last 30 years, Biotechnol. Bioprocess Eng., 20, 643–651, https://doi.org/10.1007/s12257-015-0009-0, 2015.
Joerger, R. D., Jacobson, M. R., Premakumar, R., Wolfinger, E. D., and Bishop, P. E.: Nucleotide sequence and mutational analysis of the structural genes (anfHDGK) for the second alternative nitrogenase from Azotobacter vinelandii, J. Bacteriol., 171, 1075–1086, https://doi.org/10.1128/jb.171.2.1075-1086.1989, 1989.
Joerger, R. D., Loveless, T. M., Pau, R. N., Mitchenall, L. A., Simon, B. H., and Bishop, P. E.: Nucleotide sequences and mutational analysis of the structural genes for nitrogenase 2 of Azotobacter vinelandii, J. Bacteriol., 172, 3400–3408, https://doi.org/10.1128/jb.172.6.3400-3408.1990, 1990.
Kartal, B., Maalcke, W. J., de Almeida, N. M., Cirpus, I., Gloerich, J., Geerts, W., Op den Camp, H. J. M., Harhangi, H. R., Janssen-Megens, E. M., Francoijs, K.-J., Stunnenberg, H. G., Keltjens, J. T., Jetten, M. S. M., and Strous, M.: Molecular mechanism of anaerobic ammonium oxidation, Nature, 479, 127–130, https://doi.org/10.1038/nature10453, 2011.
Kim, J. and Rees, D. C.: Nitrogenase and biological nitrogen fixation, Biochemistry, 33, 389–397, https://doi.org/10.1021/bi00168a001, 1994.
Klein, A. M., Bohannan, B. J. M., Jaffe, D. A., Levin, D. A., and Green, J. L.: Molecular Evidence for Metabolically Active Bacteria in the Atmosphere, Front. Microbiol., 7, https://doi.org/10.3389/fmicb.2016.00772, 2016.
Koirala, A. and Brözel, V. S.: Phylogeny of Nitrogenase Structural and Assembly Components Reveals New Insights into the Origin and Distribution of Nitrogen Fixation across Bacteria and Archaea, Microorganisms, 9, 1662, https://doi.org/10.3390/microorganisms9081662, 2021.
Kong, Q.-X., Wang, X.-W., Jin, M., Shen, Z.-Q., and Li, J.-W.: Development and application of a novel and effective screening method for aerobic denitrifying bacteria, FEMS Microbiol. Lett., 260, 150–155, https://doi.org/10.1111/j.1574-6968.2006.00306.x, 2006.
Krey, V.: Global energy-climate scenarios and models: a review, Wiley Interdiscip. Rev. Energy Environ., 3, 363–383, 2014.
Kumar, M. and Lin, J.-G.: Co-existence of anammox and denitrification for simultaneous nitrogen and carbon removal – Strategies and issues, J. Hazard. Mater., 178, 1–9, https://doi.org/10.1016/j.jhazmat.2010.01.077, 2010.
Lal, R.: Restoring Soil Quality to Mitigate Soil Degradation, Sustainability, 7, 5875–5895, https://doi.org/10.3390/su7055875, 2015.
Leck, C. and Bigg, E. K.: Source and evolution of the marine aerosol – A new perspective, Geophys. Res. Lett., 32, https://doi.org/10.1029/2005GL023651, 2005.
Lelieveld, J. and Crutzen, P. J.: Influences of cloud photochemical processes on tropospheric ozone, Nature, 343, 227–233, https://doi.org/10.1038/343227a0, 1990.
Lide, D. R.: CRC Handbook of Chemistry and Physics, 85th edn., CRC Press, 2900 pp., ISBN 13: 9780849304859, 2004.
Ma, B., Zhang, H., Ma, M., Huang, T., Guo, H., Yang, W., Huang, Y., Liu, X., and Li, H.: Nitrogen removal by two strains of aerobic denitrification actinomycetes: Denitrification capacity, carbon source metabolic ability, and raw water treatment, Bioresour. Technol., 344, 126176, https://doi.org/10.1016/j.biortech.2021.126176, 2022.
Machida, T., Nakazawa, T., Fujii, Y., Aoki, S., and Watanabe, O.: Increase in the atmospheric nitrous oxide concentration during the last 250 years, Geophys. Res. Lett., 22, 2921–2924, https://doi.org/10.1029/95GL02822, 1995.
Maes, M., Galecki, P., Chang, Y. S., and Berk, M.: A review on the oxidative and nitrosative stress (O&NS) pathways in major depression and their possible contribution to the (neuro)degenerative processes in that illness, Prog. Neuropsychopharmacol. Biol. Psychiatry, 35, 676–692, https://doi.org/10.1016/j.pnpbp.2010.05.004, 2011.
Maron, P.-A., Sarr, A., Kaisermann, A., Lévêque, J., Mathieu, O., Guigue, J., Karimi, B., Bernard, L., Dequiedt, S., Terrat, S., Chabbi, A., and Ranjard, L.: High Microbial Diversity Promotes Soil Ecosystem Functioning, Appl. Environ. Microbiol., 84, e02738-17, https://doi.org/10.1128/AEM.02738-17, 2018.
Mathonat, F., Mazzei, F., Prévot, M., Vernocchi, V., Gatta, E., Joly, M., Theveniot, M., Lehours, A.-C., Enault, F., Ervens, B., and Amato, P.: Phototrophy improves the aerial fitness in a photoheterotrophic Methylobacterium isolated from clouds, Authorea, https://doi.org/10.22541/au.174342623.31590139/v1, 2025.
Matson, P., Lohse, K. A., and Hall, S. J.: The Globalization of Nitrogen Deposition: Consequences for Terrestrial Ecosystems, Ambio, 31, 113–119, 2002.
McCarthy, J. J., Taylor, W. R., and Taft, J. L.: Nitrogenous nutrition of the plankton in the Chesapeake Bay. 1. Nutrient availability and phytoplankton preferences, Limnol. Oceanogr., 22, 996–1011, https://doi.org/10.4319/lo.1977.22.6.0996, 1977.
McTavish, H., Fuchs, J. A., and Hooper, A. B.: Sequence of the gene coding for ammonia monooxygenase in Nitrosomonas europaea, J. Bacteriol., 175, 2436–2444, https://doi.org/10.1128/jb.175.8.2436-2444.1993, 1993.
Merrick, M. J. and Edwards, R. A.: Nitrogen control in bacteria, Microbiol. Rev., 59, 604–622, https://doi.org/10.1128/mr.59.4.604-622.1995, 1995.
Müller, C., Elliott, J., Pugh, T. A. M., Ruane, A. C., Ciais, P., Balkovic, J., Deryng, D., Folberth, C., Izaurralde, R. C., Jones, C. D., Khabarov, N., Lawrence, P., Liu, W., Reddy, A. D., Schmid, E., and Wang, X.: Global patterns of crop yield stability under additional nutrient and water inputs, PLOS ONE, 13, e0198748, https://doi.org/10.1371/journal.pone.0198748, 2018.
Myhre, G., Myhre, C. E. L., Samset, B. H., and Storelvmo, T.: Aerosols and their relation to global climate and climate sensitivity, Nat. Educ. Knowl., 4, 7, https://www.nature.com/scitable/knowledge/library/aerosols-and-their-relation-to-global-climate-102215345/ (last access: April 2026), 2013.
Nair, A. A. and Yu, F.: Quantification of Atmospheric Ammonia Concentrations: A Review of Its Measurement and Modeling, Atmosphere, 11, 1092, https://doi.org/10.3390/atmos11101092, 2020.
Neff, J. C., Holland, E. A., Dentener, F. J., McDowell, W. H., and Russell, K. M.: The origin, composition and rates of organic nitrogen deposition: A missing piece of the nitrogen cycle?, Biogeochemistry, 57, 99–136, https://doi.org/10.1023/A:1015791622742, 2002.
Nie, S., Zhang, Z., Mo, S., Li, J., He, S., Kashif, M., Liang, Z., Shen, P., Yan, B., and Jiang, C.: Desulfobacterales stimulates nitrate reduction in the mangrove ecosystem of a subtropical gulf, Sci. Total Environ., 769, 144562, https://doi.org/10.1016/j.scitotenv.2020.144562, 2021.
Norton, J. M., Alzerreca, J. J., Suwa, Y., and Klotz, M. G.: Diversity of ammonia monooxygenase operon in autotrophic ammonia-oxidizing bacteria, Arch. Microbiol., 177, 139–149, https://doi.org/10.1007/s00203-001-0369-z, 2002.
Nuñez López, L., Amato, P., and Ervens, B.: Bacteria in clouds biodegrade atmospheric formic and acetic acids, Atmos. Chem. Phys., 24, 5181–5198, https://doi.org/10.5194/acp-24-5181-2024, 2024.
Pandis, S. N. and Seinfeld, J. H.: Sensitivity analysis of a chemical mechanism for aqueous-phase atmospheric chemistry, J. Geophys. Res.-Atmos., 94, 1105–1126, https://doi.org/10.1029/JD094iD01p01105, 1989.
Parada, A. E., Needham, D. M., and Fuhrman, J. A.: Every base matters: Assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples, Environ. Microbiol., 18, 1403–1414, https://doi.org/10.1111/1462-2920.13023, 2016.
Paul, E. A. and Clark, F. E.: Soil microbiology and biochemistry, Academic Press, San Diego, 340 pp., ISBN 13: 978-0-12-546807-7, 1996.
Péguilhan, R., Besaury, L., Rossi, F., Enault, F., Baray, J.-L., Deguillaume, L., and Amato, P.: Rainfalls sprinkle cloud bacterial diversity while scavenging biomass, FEMS Microbiol. Ecol., 97, fiab144, https://doi.org/10.1093/femsec/fiab144, 2021.
Péguilhan, R., Rossi, F., Joly, M., Nasr, E., Batut, B., Enault, F., Ervens, B., and Amato, P.: Clouds influence the functioning of airborne microorganisms, Biogeosciences, 22, 1257–1275, https://doi.org/10.5194/bg-22-1257-2025, 2025.
Pester, M., Maixner, F., Berry, D., Rattei, T., Koch, H., Lücker, S., Nowka, B., Richter, A., Spieck, E., Lebedeva, E., Loy, A., Wagner, M., and Daims, H.: NxrB encoding the beta subunit of nitrite oxidoreductase as functional and phylogenetic marker for nitrite-oxidizing Nitrospira, Environ. Microbiol., 16, 3055–3071, https://doi.org/10.1111/1462-2920.12300, 2014.
Philippot, L. and Højberg, O.: Dissimilatory nitrate reductases in bacteria, Biochim. Biophys. Acta BBA – Gene Struct. Expr., 1446, 1–23, https://doi.org/10.1016/S0167-4781(99)00072-X, 1999.
Pilegaard, K.: Processes regulating nitric oxide emissions from soils, Philos. Trans. R. Soc. B Biol. Sci., 368, 20130126, https://doi.org/10.1098/rstb.2013.0126, 2013.
Poly, F., Monrozier, L. J., and Bally, R.: Improvement in the RFLP procedure for studying the diversity of nifH genes in communities of nitrogen fixers in soil, Res. Microbiol., 152, 95–103, https://doi.org/10.1016/s0923-2508(00)01172-4, 2001.
Pouzet, G., Peghaire, E., Aguès, M., Baray, J.-L., Conen, F., and Amato, P.: Atmospheric Processing and Variability of Biological Ice Nucleating Particles in Precipitation at Opme, France, Atmosphere, 8, 229, https://doi.org/10.3390/atmos8110229, 2017.
Renard, P., Canet, I., Sancelme, M., Wirgot, N., Deguillaume, L., and Delort, A.-M.: Screening of cloud microorganisms isolated at the Puy de Dôme (France) station for the production of biosurfactants, Atmos. Chem. Phys., 16, 12347–12358, https://doi.org/10.5194/acp-16-12347-2016, 2016.
Renard, P., Bianco, A., Baray, J.-L., Bridoux, M., Delort, A.-M., and Deguillaume, L.: Classification of Clouds Sampled at the Puy de Dôme Station (France) Based on Chemical Measurements and Air Mass History Matrices, Atmosphere, 11, 732, https://doi.org/10.3390/atmos11070732, 2020.
Renard, P., Brissy, M., Rossi, F., Leremboure, M., Jaber, S., Baray, J.-L., Bianco, A., Delort, A.-M., and Deguillaume, L.: Free amino acid quantification in cloud water at the Puy de Dôme station (France), Atmos. Chem. Phys., 22, 2467–2486, https://doi.org/10.5194/acp-22-2467-2022, 2022.
Robertson, L. A. and Kuenen, J. G.: Aerobic denitrification – old wine in new bottles?, Antonie Van Leeuwenhoek, 50, 525–544, https://doi.org/10.1007/BF02386224, 1984.
Rossi, F., Péguilhan, R., Turgeon, N., Veillette, M., Baray, J.-L., Deguillaume, L., Amato, P., and Duchaine, C.: Quantification of antibiotic resistance genes (ARGs) in clouds at a mountain site (puy de Dôme, central France), Sci. Total Environ., 865, 161264, https://doi.org/10.1016/j.scitotenv.2022.161264, 2023.
Rotthauwe, J. H., Witzel, K. P., and Liesack, W.: The ammonia monooxygenase structural gene amoA as a functional marker: molecular fine-scale analysis of natural ammonia-oxidizing populations., Appl. Environ. Microbiol., 63, 4704–4712, 1997.
Ruiz de Morales, J., Calbó, J., González, J.-A., and Sola, Y.: A method to assess the cloud-aerosol transition zone from ceilometer measurements, Atmos. Res., 310, 107623, https://doi.org/10.1016/j.atmosres.2024.107623, 2024.
Rustad, T. R., Minch, K. J., Brabant, W., Winkler, J. K., Reiss, D. J., Baliga, N. S., and Sherman, D. R.: Global analysis of mRNA stability in Mycobacterium tuberculosis, Nucleic Acids Res., 41, 509–517, https://doi.org/10.1093/nar/gks1019, 2013.
Šantl-Temkiv, T., Amato, P., Gosewinkel, U., Thyrhaug, R., Charton, A., Chicot, B., Finster, K., Bratbak, G., and Löndahl, J.: High-Flow-Rate Impinger for the Study of Concentration, Viability, Metabolic Activity, and Ice-Nucleation Activity of Airborne Bacteria, Environ. Sci. Technol., 51, 11224–11234, https://doi.org/10.1021/acs.est.7b01480, 2017.
Šantl-Temkiv, T., Gosewinkel, U., Starnawski, P., Lever, M., and Finster, K.: Aeolian dispersal of bacteria in southwest Greenland: their sources, abundance, diversity and physiological states, FEMS Microbiol. Ecol., 94, fiy031, https://doi.org/10.1093/femsec/fiy031, 2018.
Scala, D. J. and Kerkhof, L. J.: Diversity of Nitrous Oxide Reductase (nosZ) Genes in Continental Shelf Sediments, Appl. Environ. Microbiol., 65, 1681–1687, https://doi.org/10.1128/AEM.65.4.1681-1687.1999, 1999.
Schostag, M. D., Albers, C. N., Jacobsen, C. S., and Priemé, A.: Low Turnover of Soil Bacterial rRNA at Low Temperatures, Front. Microbiol., 11, https://doi.org/10.3389/fmicb.2020.00962, 2020.
Sohaskey, C. D. and Wayne, L. G.: Role of narK2X and narGHJI inHypoxic Upregulation of Nitrate Reduction byMycobacteriumtuberculosis, J. Bacteriol., 185, 7247–7256, https://doi.org/10.1128/jb.185.24.7247-7256.2003, 2003.
Song, W., Liu, J., Qin, W., Huang, J., Yu, X., Xu, M., Stahl, D., Jiao, N., Zhou, J., and Tu, Q.: Functional Traits Resolve Mechanisms Governing the Assembly and Distribution of Nitrogen-Cycling Microbial Communities in the Global Ocean, mBio, 13, e03832-21, https://doi.org/10.1128/mbio.03832-21, 2022.
Sorai, M., Yoshida, N., and Ishikawa, M.: Biogeochemical simulation of nitrous oxide cycle based on the major nitrogen processes, J. Geophys. Res.-Biogeo., 112, https://doi.org/10.1029/2005JG000109, 2007.
Stolz, J. F. and Basu, P.: Evolution of Nitrate Reductase: Molecular and Structural Variations on a Common Function, ChemBioChem, 3, 198–206, 2002.
Sutton, M. A., Reis, S., Riddick, S. N., Dragosits, U., Nemitz, E., Theobald, M. R., Tang, Y. S., Braban, C. F., Vieno, M., Dore, A. J., Mitchell, R. F., Wanless, S., Daunt, F., Fowler, D., Blackall, T. D., Milford, C., Flechard, C. R., Loubet, B., Massad, R., Cellier, P., Personne, E., Coheur, P. F., Clarisse, L., Van Damme, M., Ngadi, Y., Clerbaux, C., Skjøth, C. A., Geels, C., Hertel, O., Wichink Kruit, R. J., Pinder, R. W., Bash, J. O., Walker, J. T., Simpson, D., Horváth, L., Misselbrook, T. H., Bleeker, A., Dentener, F., and de Vries, W.: Towards a climate-dependent paradigm of ammonia emission and deposition, Philos. Trans. R. Soc. B Biol. Sci., 368, 20130166, https://doi.org/10.1098/rstb.2013.0166, 2013.
Suzek, B. E., Wang, Y., Huang, H., McGarvey, P. B., Wu, C. H., and the UniProt Consortium: UniRef clusters: a comprehensive and scalable alternative for improving sequence similarity searches, Bioinformatics, 31, 926–932, https://doi.org/10.1093/bioinformatics/btu739, 2015.
Suzuki, M., ARAI, H., ISHII, M., and IGARASHI, Y.: Gene Structure and Expression Profile of Cytochrome bc Nitric Oxide Reductase from Hydrogenobacter thermophilus TK-6, Biosci. Biotechnol. Biochem., 70, 1666–1671, https://doi.org/10.1271/bbb.60018, 2006.
Syakila, A. and Kroeze, C.: The global nitrous oxide budget revisited, Greenh. Gas Meas. Manag., 1, 17–26, https://doi.org/10.3763/ghgmm.2010.0007, 2011.
Trüper, H. G. and Pfennig, N.: Characterization and Identification of the Anoxygenic Phototrophic Bacteria, in: The Prokaryotes: A Handbook on Habitats, Isolation, and Identification of Bacteria, edited by: Starr, M. P., Stolp, H., Trüper, H. G., Balows, A., and Schlegel, H. G., Springer, Berlin, Heidelberg, 299–312, https://doi.org/10.1007/978-3-662-13187-9_18, 1981.
Tu, Q., He, Z., Wu, L., Xue, K., Xie, G., Chain, P., Reich, P. B., Hobbie, S. E., and Zhou, J.: Metagenomic reconstruction of nitrogen cycling pathways in a CO2-enriched grassland ecosystem, Soil Biol. Biochem., 106, 99–108, https://doi.org/10.1016/j.soilbio.2016.12.017, 2017.
Tu, Q., Lin, L., Cheng, L., Deng, Y., and He, Z.: NCycDB: a curated integrative database for fast and accurate metagenomic profiling of nitrogen cycling genes, Bioinformatics, 35, 1040–1048, https://doi.org/10.1093/bioinformatics/bty741, 2019.
Vaïtilingom, M., Amato, P., Sancelme, M., Laj, P., Leriche, M., and Delort, A.-M.: Contribution of Microbial Activity to Carbon Chemistry in Clouds, Appl. Environ. Microbiol., 76, 23–29, https://doi.org/10.1128/AEM.01127-09, 2010.
Vaïtilingom, M., Attard, E., Gaiani, N., Sancelme, M., Deguillaume, L., Flossmann, A. I., Amato, P., and Delort, A.-M.: Long-term features of cloud microbiology at the puy de Dôme (France), Atmos. Environ., 56, 88–100, https://doi.org/10.1016/j.atmosenv.2012.03.072, 2012.
Vaïtilingom, M., Deguillaume, L., Vinatier, V., Sancelme, M., Amato, P., Chaumerliac, N., and Delort, A.-M.: Potential impact of microbial activity on the oxidant capacity and organic carbon budget in clouds, P. Natl. Acad. Sci. USA, 110, 559–564, https://doi.org/10.1073/pnas.1205743110, 2013.
Vergne, A., Darbot, V., Bardot, C., Enault, F., Le Jeune, A.-H., Carrias, J.-F., Corbara, B., Céréghino, R., Leroy, C., Jeanthon, C., Giraud, E., Mary, I., and Lehours, A.-C.: Assemblages of anoxygenic phototrophic bacteria in tank bromeliads exhibit a host-specific signature, J. Ecol., 109, 2550–2565, https://doi.org/10.1111/1365-2745.13657, 2021.
Vernocchi, V., Abd El, E., Brunoldi, M., Danelli, S. G., Gatta, E., Isolabella, T., Mazzei, F., Parodi, F., Prati, P., and Massabò, D.: Airborne bacteria viability and air quality: a protocol to quantitatively investigate the possible correlation by an atmospheric simulation chamber, Atmos. Meas. Tech., 16, 5479–5493, https://doi.org/10.5194/amt-16-5479-2023, 2023.
Vinatier, V., Wirgot, N., Joly, M., Sancelme, M., Abrantes, M., Deguillaume, L., and Delort, A.-M.: Siderophores in Cloud Waters and Potential Impact on Atmospheric Chemistry: Production by Microorganisms Isolated at the Puy de Dôme Station, Environ. Sci. Technol., 50, 9315–9323, https://doi.org/10.1021/acs.est.6b02335, 2016.
Voss, M., Baker, A., Bange, H. W., Conley, D., Cornell, S., Deutsch, B., Engel, A., Ganeshram, R., Garnier, J., Heiskanen, A.-S., Jickells, T., Lancelot, C., McQuatters-Gollop, A., Middelburg, J., Schiedek, D., Slomp, C. P., and Conley, D. P.: Nitrogen processes in coastal and marine ecosystems, in: The European Nitrogen Assessment, edited by: Sutton, M. A., Howard, C. M., Erisman, J. W., Billen, G., Bleeker, A., Grennfelt, P., Van Grinsven, H., and Grizzetti, B., Cambridge University Press, 147–176, https://doi.org/10.1017/CBO9780511976988.011, 2011.
Voss, M., Bange, H. W., Dippner, J. W., Middelburg, J. J., Montoya, J. P., and Ward, B.: The marine nitrogen cycle: recent discoveries, uncertainties and the potential relevance of climate change, Philos. Trans. R. Soc. B Biol. Sci., 368, 20130121, https://doi.org/10.1098/rstb.2013.0121, 2013.
Walker, M. C. and van der Donk, W. A.: The many roles of glutamate in metabolism, J. Ind. Microbiol. Biotechnol., 43, 419–430, https://doi.org/10.1007/s10295-015-1665-y, 2016.
Wang, B., Zheng, X., Zhang, H., Yu, X., Lian, Y., Yang, X., Yu, H., Hu, R., He, Z., Xiao, F., and Yan, Q.: Metagenomic insights into the effects of submerged plants on functional potential of microbial communities in wetland sediments, Mar. Life Sci. Technol., 3, 405–415, https://doi.org/10.1007/s42995-021-00100-3, 2021.
Whelan, J. A., Russell, N. B., and Whelan, M. A.: A method for the absolute quantification of cDNA using real-time PCR, J. Immunol. Methods, 278, 261–269, https://doi.org/10.1016/S0022-1759(03)00223-0, 2003.
Whitman, W. B., Coleman, D. C., and Wiebe, W. J.: Prokaryotes: The unseen majority, P. Natl. Acad. Sci. USA, 95, 6578–6583, https://doi.org/10.1073/pnas.95.12.6578, 1998.
Wilson, L. P. and Bouwer, E. J.: Biodegradation of aromatic compounds under mixed oxygen/denitrifying conditions: a review, J. Ind. Microbiol. Biotechnol., 18, 116–130, https://doi.org/10.1038/sj.jim.2900288, 1997.
Wright, D. N., Bailey, G. D., and Goldberg, L. J.: Effect of Temperature on Survival of Airborne Mycoplasma pneumoniae, J. Bacteriol., 99, 491–495, 1969.
Xiang, S.-R., Doyle, A., Holden, P. A., and Schimel, J. P.: Drying and rewetting effects on C and N mineralization and microbial activity in surface and subsurface California grassland soils, Soil Biol. Biochem., 40, 2281–2289, https://doi.org/10.1016/j.soilbio.2008.05.004, 2008.
Yang, J., Feng, L., Pi, S., Cui, D., Ma, F., Zhao, H., and Li, A.: A critical review of aerobic denitrification: Insights into the intracellular electron transfer, Sci. Total Environ., 731, 139080, https://doi.org/10.1016/j.scitotenv.2020.139080, 2020.
Zeng, J., Liao, S., Qiu, M., Chen, M., Ye, J., Zeng, J., and Wang, A.: Effects of carbon sources on the removal of ammonium, nitrite and nitrate nitrogen by the red yeast Sporidiobolus pararoseus Y1, Bioresour. Technol., 312, 123593, https://doi.org/10.1016/j.biortech.2020.123593, 2020.
Zhang, J., Wu, P., Hao, B., and Yu, Z.: Heterotrophic nitrification and aerobic denitrification by the bacterium Pseudomonas stutzeri YZN-001, Bioresour. Technol., 102, 9866–9869, https://doi.org/10.1016/j.biortech.2011.07.118, 2011.
Zhang, Q. and Anastasio, C.: Chemistry of fog waters in California's Central Valley – Part 3: concentrations and speciation of organic and inorganic nitrogen, Atmos. Environ., 35, 5629–5643, https://doi.org/10.1016/S1352-2310(01)00337-5, 2001.
Zumft, W. G.: Cell biology and molecular basis of denitrification, Microbiol. Mol. Biol. Rev., 61, 533–616, https://doi.org/10.1128/mmbr.61.4.533-616.1997, 1997.
Zumft, W. G. and Kroneck, P. M. H.: Respiratory Transformation of Nitrous Oxide (N2O) to Dinitrogen by Bacteria and Archaea, in: Advances in Microbial Physiology, vol. 52, edited by: Poole, R. K., Academic Press, 107–227, https://doi.org/10.1016/S0065-2911(06)52003-X, 2006.