the Creative Commons Attribution 4.0 License.
the Creative Commons Attribution 4.0 License.
Vertebrate impact on bacterial community structure of coastal Arctic spring snowpacks
Sławomir Sułowicz
Krzysztof Zawierucha
Anna Markowicz
Krystyna Kozioł
Wiktoria Zientak
Adam Nawrot
Krzesimir Tomaszewski
Christoph Keuschnig
Bartłomiej Luks
Catherine Larose
Snow covers up to 35 % of the Earth's surface seasonally, impacts biogeochemical cycling, and forms a microbial habitat despite harsh and variable conditions. While atmospheric deposition is a well-known source of microbial input, the role of vertebrates in shaping snow microbiomes remains underexplored. In Arctic ecosystems, seabirds and terrestrial mammals influence the concentration and distribution of nutrients, but their effects on microbial communities in snow remain poorly understood. Here, we explore the role of vertebrates in shaping snow microbial biodiversity of Arctic terrestrial snowpacks. The study was conducted on the northern coast of Hornsund Fjord on Spitsbergen. Forty snow samples were collected in four transects, two established along the gradient from the centre of a seabird (Alle alle) colony towards non-impacted areas and two transects along the coast. We identified 854 bacterial ASVs using short-read sequencing of the 16S rRNA gene. Samples clustered into four groups based on community composition, but were not linked to spatial factors such as distance to colonies. Bird and terrestrial mammal faecal indicators like Catellicoccus or Streptococcus were detected in 17 out of the 40 samples and drove the formation of two distinct clusters. These findings suggest that coastal Arctic snow microbiomes are strongly shaped by biological activity, with wildlife acting as key microbial vectors.
- Article
(3021 KB) - Full-text XML
-
Supplement
(1259 KB) - BibTeX
- EndNote
Snow is an important component of the cryosphere, hydrosphere and biosphere, regulating albedo, water resources, and biogeochemical cycles (Déry and Brown, 2007; Dong et al., 2023; Hudson, 2006). Although seasonal snow covers up to half of the Northern Hemisphere land surface, most studies on the biological diversity and ecological processes on the snow surface are focused on seasonal snow patches during summer, specifically on snow algae blooms in lower latitudes (Hoham and Remias, 2020; Lemke et al., 2007). Arctic seasonal snowpacks are now recognized as viable habitats for diverse microbial communities (Maccario et al., 2015), even outside the melting period (Amato et al., 2007; Malard et al., 2021). To colonize and thrive in cold environments like snowpacks, microorganisms must adapt to physiological stressors such as low temperatures (below 0 °C), high solar radiation, low nutrient and water availability and freeze-thaw cycles (Larose et al., 2013). In the Arctic, high latitude leads to pronounced seasonality, causing gradual yet extreme shifts in photoperiod, irradiance, and temperature (Larose et al., 2013), which can induce oxidative stress (Sanchez-Cid et al., 2023). These temporal ecosystems provide an opportunity for studying the drivers of microbial community structure and assembly mechanisms, as they form new habitats seeded by microbes from snow-free ground or the atmosphere (Keuschnig et al., 2023).
When considering potential colonization of microorganisms to snowpacks, seeding by abiotic transport factors has generally been the focus. One of the most recognized sources for freshly developing snow habitats is the atmosphere; with microorganisms entering the snowpack via wet and dry deposition (Maccario et al., 2019). Colonization from the terrestrial surface upon which the snowpack develops might also be a factor, e.g. from nunataks (Monteil et al., 2012). Keuschnig et al., (2023) revealed that microbial abundance in surface snow, but not diversity, was tightly linked to sea spray, although few marine taxa were detected in snowpacks. In contrast, biotic vectors, such as migrating animals, have been less studied as colonization routes. Vertebrates that inhabit the Arctic, such as birds, reindeers, bears and foxes, can transport and spread significant quantities of biological and chemical material with the potential to influence community structure, diversity and ecosystem function in resident habitats (Bauer and Hoye, 2014). Seabirds, feeding in the sea and breeding on land, deliver nutrients in the form of guano, feathers and egg-shells to terrestrial ecosystems, thereby linking nutrient-rich marine ecosystems with nutrient-poor ecosystems on land (Zmudczyńska-Skarbek et al., 2024). Depending on their diet and nesting sites, these birds can shape the diversity of plant cover and other organisms (Zawierucha et al., 2016, 2019; Zmudczyńska-Skarbek et al., 2024; Zwolicki et al., 2013). Bird colonies reestablish during the late snow season when migrating seabirds return to the Arctic. These colonies attract other vertebrates that feed on lichens (reindeer, Rangifer tarandus) or for hunting (polar fox, Vulpes lagopus) (Jakubas et al., 2008). Unlike abiotic vectors, these organisms are not limited by environmental gradients, allowing them to transport materials in diverse directions and against prevailing abiotic flow patterns (McInturf et al., 2019). Animal-origin nutrient additions might also impact snow microbial community structure since organic inputs have been shown to be the main drivers of bacterial diversity (Keuschnig et al., 2023) and microbial interactions (Bergk Pinto et al., 2019). For example, seabird colonies have been shown to affect nutrients (), trace elements and organic matter concentration (Souza-Kasprzyk et al., 2022). In addition to contributing nutrients, vertebrates can directly introduce microorganisms, including multidrug-resistant bacteria, parasites, and propagules into snowpacks through excretions, facilitating their transfer and dispersal (Hayashi et al., 2018). For instance, migratory birds have been shown to play an important role in the spread of antibiotic resistant bacteria to snow ecosystems in remote localities (Segawa et al., 2013). Whether these organisms are able to survive and colonize once they have been deposited into snowpacks has yet to be determined. The adverse abiotic factors encountered in the snow habitat, like cold temperatures and limited nutrient availability, likely hinder the survival of animal-associated microorganisms, while established snow communities might outcompete newcomers for resources.
While the influence of aerosols and snow/ice-free areas on bacterial communities in cryospheric ecosystems (e.g., snow, ice) has been well documented (Franzetti et al., 2017), the role of vertebrate activity in shaping microbial composition and function on snow remains unexplored. In this study, we assess how marine vertebrates (seabirds) and terrestrial animals (polar foxes, reindeer) alter microbial communities in coastal spring snowpacks in the vicinity of Polish Polar Station Hornsund (PPSH), Svalbard. Hornsund is a fjord in the southwestern part of Spitsbergen, the largest island of the Svalbard archipelago. Climate at Hornsund shores is strongly shaped by Atlantic waters, and by less saline, coastal Arctic waters from the Barents Sea. Ocean currents, geomorphology and the presence of both marine and land-terminating glaciers sustain a high productivity and animal species richness in the area (Skagseth et al., 2008; Wesławski et al., 2006). Hornsund, an All Taxon Biodiversity Inventory (ATBI) site (Warwick et al., 2003), is inhabited by one of the largest worldwide concentrations of little auks (Alle alle) during breeding season (Keslinka et al., 2019). The little auk colony supplied ca. 60 t of dry guano per km2 of colony area during the breeding season, which shapes tundra plants, algae, invertebrates and bacterial communities (Stempniewicz, 1990; Zawierucha et al., 2019; Zielińska et al., 2016b; Zwolicki et al., 2013). Apart from birds, Hornsund is an important route for migration of polar bears Ursus maritimus, and habitat for two other mammals in Svalbard, reindeers Rangifer tarandus platyrhynchus and polar foxes Vulpes lagopus (Stempniewicz, 2017). Reindeers and foxes are known to disseminate parasites in this Arctic region (Myšková et al., 2019; Popiołek et al., 2007).
In our study, we analyzed bacterial biodiversity along four transects: two adjacent to a little auk colony and two distanced from the direct avian influence. Sampling areas exhibited clear signs of vertebrate activity, including observations of reindeer herd presence, polar fox tracks, and biological remnants (feathers, bones, excrements). We hypothesized that seabirds and terrestrial vertebrates serve as a source of bacteria and nutrients on snowpacks. We assume such nutrients could reshape the microbial community on snow. Understanding such microbial diversity and dynamics is urgent given the rapid decline of Arctic snowpacks.
2.1 Sample collection, DNA isolation and NGS sequencing
To study the effect of biotic vectors on the colonization of non-melting snowpacks, we conducted a field experiment near the PPSH in May 2024, on the northern coast of Hornsund fjord (south-west Spitsbergen), on the slopes of the Fugleberget (569 m a.s.l.) and Ariekammen (517 m a.s.l.) mountains. They are located ca. 1 km north from the station, and the little auk colonies on these slopes are inhabited during summer season by ca. 25 000 pairs (Keslinka et al., 2019). In winter 2023/2024, the snow season started in late October. Between November and February, snowpack in the catchment was stable, with minor depth changes due to wind redeposition and occasional thaws. Maximum depth depended on the measurement point within the catchment and reached 111–127 cm at the feet of Fugleberget (sample lines A and B), 70–80 cm in the Fuglebergsletta plain (sample line C), and up to 58 cm at the coast (sampling line D). In mid-May snowmelt has started, with snow patches lasting at the bottom of the slope until the end of June, and snow disappearing at the coast as early as first week of June 2024. The total recorded precipitation in the period 1 October 2023–31 May 2024 amounted to 209.5 mm. Of this, 152.4 mm occurred on days when the mean daily air temperature was at or below 0 °C, while 57.1 mm was recorded on days with mean temperatures above 0 °C. Notably, 15 d within the observation period experienced precipitation under positive temperature conditions.
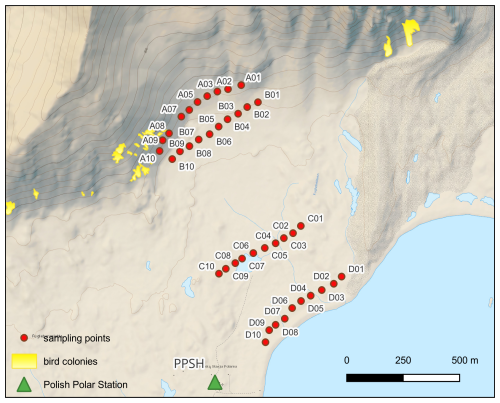
A total of 40 surface snow samples (top 10 cm) were collected into sterile Whirl-Pak bags (equivalent to 2–3 L of melted snow) using a sterilized Teflon shovel from four transects (Line A, B, C, D) (Fig. 1). Two transects were established from the centre of a seabird colony towards non-impacted areas along an altitudinal gradient, while two other parallel transects were established closer to the coast. The measured snow density of this top layer showed low variability (255 ± 26 g dm−3; average ± 1 SD). To reduce contamination, Tyvex® body suits, face masks and latex gloves were used during sample handling (Larose et al., 2010). Samples were immediately transported to the laboratory, and were left to melt at room temperature. A total of 1.5 L of melted snow were filtered onto sterile MCE (Mixed Cellulose Esters) 0.22 µm pore size, 47 mm diameter white gridded filters (Millipore) using a sterile filtration unit (Nalge Nunc International Corporation). The samples for chemical analyses were measured on site or transported in cool storage for further analysis. Similarly, filters with microbial biomass for DNA isolation were stored in sterile Eppendorf tubes at −20 °C. Procedural blanks were carried out by filtering MilliQ water (sample MQB2 and MQB3 in Supplementary Excel 1 metadata table) and treated as samples for downstream analysis.
Environmental DNA was extracted from filters using the PowerWater® DNA Isolation Kit (MoBio, Carlsbad, CA, US) according to the manufacturer's instructions. The DNA purity was checked spectrophotometrically and the concentration of DNA measured using the NanoPhotometer NP80 was in the range of 2.7–9.3 ng µL−1. The 16S rRNA fragment was amplified using bacterial primers 341F (CCTACGGGNGGCWGCAG) and 805R (GACTACHVGGGTATCTAATCC) spanning the V3–V4 hypervariable regions (Herlemann et al., 2011). Sequencing was performed on the MiSeq™ platform (Illumina, San Diego, USA) using paired-end reads (2 × 300 bp) by the Macrogen company (Macrogen Europe, Amsterdam, The Netherlands). Amplicon dataset, including blanks, was deposited at the Sequence Read Archive (ID: PRJNA1277323).
2.2 Bioinformatics and statistical analysis
Paired-end 16S rRNA gene amplicon reads were processed in R using DADA2 (Callahan et al., 2016) to generate amplicon sequence variants (ASVs). Briefly, primers targeting the V3-V4 region of the bacterial 16S rRNA gene were cut from raw reads with cutadapt (Martin, 2011) in paired-end mode with a minimum post-trim length of 50 bp. Forward and reverse reads were cut to 270 and 260 bp based on observation of quality profiles and the reads were further filtered using truncQ = 12 and maxEE = 1, generating in total 1.4 million high quality reads. Error models, dereplication, ASV inference (pool = FALSE) and merging were done with default options. Merged reads with a size between 400–435 bp based on the expected amplicon size were retained which corresponded to 8611 ASVs, then chimeras were removed using the consensus method, resulting in 7640 ASVs. Taxonomy was assigned to these ASVs using SILVA NR99 training set v138.2 (Chuvochina et al., 2025). Contaminants were identified using the R package decontam in prevalence mode and a threshold of 0.2. We performed a stringent prevalence (at least present in 3 samples) and abundance (at least 5 counts per ASV over all samples) filtering to further reduce the influence of spurious low abundance ASVs (Bokulich et al., 2013). Script details are attached in Supplementary materials. After removal of ASVs annotated to chloroplast and mitochondrial sequences, this analysis resulted in a final set of 854 ASVs.
Next, Estimate Alpha and Beta Diversities workflow (Qiagen, 2024) was used to estimate relative abundance, alpha and beta diversity. Relative abundance at the family and genus level was presented as a stacked area chart. Alpha-diversity richness metrics (total number of ASVs, Faith's phylogenetic diversity PD) were calculated based on ASV counts (presence–absence), whereas diversity indices accounting for evenness (Shannon entropy and Simpson's index) were calculated using ASV read abundances. (Faith and Baker, 2006; Qiagen, 2024). Significant changes in the alpha diversity indices between snow clusters were evaluated using pairwise Mann-Whitney U tests (P<0.05).
Beta-diversity analyses were performed using generalized UniFrac distances d(0.5) calculated from ASV tables based on read abundances. Principal Coordinates Analysis (PCoA) was then applied to the resulting distance matrix to visualize differences in community composition among samples (Chen et al., 2012). Significant changes between samples based on their transect localization (line as a main factor) or created snow clusters (cluster as factors) were detected using the PERMANOVA analysis tool with 99 999 permutations. Among the 52 genera present at >1 % relative abundance in one of the clusters, 46 significantly differentiated snow clusters (FDR-corrected P<0.05). These genera were visualized using a Euclidean distance-based heatmap with TMM normalized (CPM), log-transformed and Z-score-scaled read abundances. Their overlap among clusters was visualized using Venn diagram (Qiagen, 2024).
2.3 Physicochemical analysis of snow properties
Samples for the determination of physicochemical parameters and major ions were collected into sterile Whirl-Pak bags using face masks, gloves and a special protective suit against contamination. Samples were transported to the Polish Polar Station Hornsund. After melting at room temperature, pH and conductivity were measured using calibrated EPP-1 pH and EFC-1t conductivity probes and pH/conductivity meter CPC-505 (Elmetron), and HCO was determined by titration with 0.02 M HCl to pH 4.4 using a Titrino 702SM automatic titrator (Metrohm) and a pH probe (Metrohm). Then, a sample aliquot for inorganic ion analysis was filtered with sterile MCE (Mixed Cellulose Esters) 0.45 µm pore size, 47 mm diameter white gridded filters (Millipore). Inorganic ion concentrations were determined on a Metrohm 930 Compact IC Flex ion chromatograph equipped with an autosampler (Metrohm, Herisau, Switzerland). Cation samples were acidified with 2 µL of 2 mM HNO3 per 10 mL sample prior to analysis. Cations NH, Ca2+, Mg2+ were determined (PN EN ISO 14911, 2002) without suppression using column Metrohm 930 Compact IC Flex and eluent HNO3 34 mM/dipicolinic acid 14 mM in water (from IC eluent concentrate (20×), Supelco 61905). Anions (Br−, Cl−, NO, PO, SO) were determined (PN EN ISO 10304-1, 2009) using chemical suppression on column Metrosep A Supp 5 – 250/4.0 and eluent Na2CO3 64 mM/NaHCO3 20 mM in water (from IC eluent concentrate (20×), Supelco 62414). The injection volume was 20 µL in the anion system and 100 µL in the cation system.
Snow samples for TOC (Total Organic Carbon) were collected in the field into HDPE 500 mL bottles, melted in them and transferred directly after melting into 60 mL amber glass vials with a PTFE septum. These vials were closed airtight with no headspace, stored in cool storage (+4 °C) and transported to Poland. TOC (total organic carbon) was determined with a TOC-L Analyser (Shimadzu, Japan), utilizing catalytic oxidation to CO2 by combustion at 680 °C with a high-sensitivity platinum catalyst. CO2 was then detected by NDIR (non-dispersive infra-red detection). TOC was determined as NPOC (non-purgeable organic carbon), which is a method that may lead to an underestimation of volatile organic compounds. Calibration curves achieved R2≥0.995, with blank offset (deionized water <30 ppb TOC was used for all blanks). The analytical range was 0.09–10 mg L−1 (for concentrations >10 mg L−1, dilutions were applied); limit of detection (LOD) and limit of quantification (LOQ) were 0.03 and 0.09 mg L−1, respectively. Other measurement settings were: 3 out of 4 repeats accepted, CV < 2 %, sparging time (with HCl) 90 s. Field container blanks were performed in 5 repeats and yielded results < LOQ. Comparisons of two sample storage types, i.e., samples stored frozen in polypropylene vials with the samples in amber glass, yielded consistent results.
2.4 Statistical analysis of physicochemical variables
Canonical correspondence analysis (CCA) was used to identify the most important environmental factors that correlated with the snow cluster microbial communities. The CCA was performed using PAST software (Hammer et al., 2001). Among 20 measured environmental variables describing snow physicochemical properties, a subset of 12 (pH, conductivity, non-purgeable organic carbon [NPOC], cations NH, Ca2+, Mg2+ and anions HCO, Br−, Cl−, NO, PO, SO) was selected following preliminary screening to remove redundant and derived variables (e.g., N-NH4, N-NO3) and to avoid overrepresentation of closely related parameters (e.g., Na+ and Cl−, total and inorganic nitrogen). This approach ensured that the retained variables represented distinct environmental gradients while reducing collinearity. Additionally, Spearman's rank correlation analysis was applied to assess relationships among the environmental variables and significance of the correlations (P<0.05, Bonferroni corrected). Despite this selection, some degree of correlation between variables is expected in natural environmental datasets (Table S2). Microbial community data were analyzed at the genus level and restricted to 46 bacterial genera exceeding 1 % relative abundance in at least one cluster and significantly differentiated snow clusters (the false discovery rate corrected, FDR-corrected P<0.05), in order to focus on the most informative taxa and reduce the impact of stochastic variation on clustering patterns. In addition, two sequencing-derived technical variables (the total number of sequencing reads per sample and the total number of detected ASVs per sample) were included as auxiliary variables to account for variation related to sequencing depth and observed richness; these variables were not interpreted as ecological drivers. Prior to analysis, all variables except pH (which is inherently logarithmic) were log10-transformed to stabilize variances and reduce the influence of extreme values on the ordination (Supplementary materials Excel 2).
3.1 Bacterial community structure – beta diversity and taxa distribution
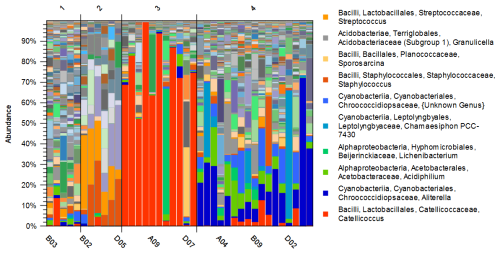
Principal Coordinate Analysis was used to compare beta-diversity profiles across different sampling sites. When the transect line was considered as a grouping factor, PERMANOVA did not reveal significant overall multivariate differences in bacterial community composition among lines (P=0.089) (Fig. S1). However, based on the 2D and 3D view (Fig. 2) of principal coordinate analysis (PCoA) of bacterial 16S rRNA profile, samples formed four clusters characterized by distinct multivariate community profiles, a pattern supported by PERMANOVA results (P<0.001 for each pair of clusters, except for the comparison of Clusters 1 and 2, where P<0.01). This observation was additionally supported by taxa distribution pattern in each cluster (Fig. 3). The obtained 16S rRNA sequences were assigned to a total of 854 bacterial ASVs that were mainly classified into Bacilli, Cyanobacteriia, and Alpha- and Gammaproteobacteria classes (Table S1). Distribution varied across samples, and the bacterial markers responsible for their grouping were identified. At the genus level (Fig. 3), Catellicoccus (Bacilli class) dominated samples A05–A10, B10, C07, and D08, comprising between 52.3 %–99.5 % of reads. Other abundant genera included Streptococcus (Bacilli; up to 35.9 %) and Aliterella (Cyanobacteriia class; up to 72.2 % in D06). Samples with Catellicoccus marker created cluster number three (marked red in Fig. 2). Cluster number two was distinguished based on the higher Streptococcus content (orange one). These two clusters contained 17 snow samples. Other samples were classified into two additional clusters. ASVs affiliated with Aliterella characterized cluster number four (blue). Additionally, Cluster 1 (Fig. 2, marked in gold) separated from Cluster 4 along PCo2 and was characterized by high bacterial diversity (Fig. S2, Sect. S1, Fig. S3).
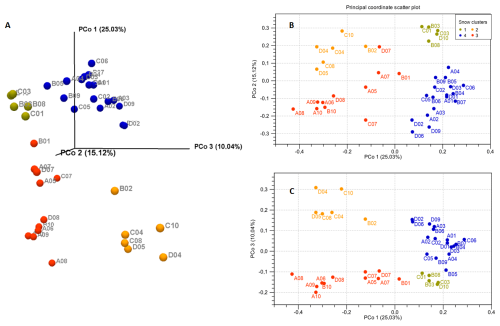
Figure 2Principal coordinates analysis (PCoA) plots of bacterial taxonomic profiles based on the generalized UniFrac distance d(0.5) matrix. (A) 3D view; (B) 2D view – PCo1 and PCo2 axes; (C) 2D view – PCo1 and PCo3 axes.
3.2 Microbial taxa that differentiate snow clusters
To identify additional marker taxa that were not clearly apparent from percentage-based bar charts, statistical analyses were performed to detect genera significantly differentiating the samples, and the resulting 46 genera were subsequently visualized using a heatmap to highlight key taxa across clusters (Fig. 4). In Cluster 3, ASV reads affiliated with Catellicoccus and genus Candidatus Arthromitus (Clostridia class) were significantly higher (P<0.001) than in other clusters. Abundance of ASVs classified as Psychrobacter, Sporosarcina, Aequorivita and Gelidibacter were significantly higher in Cluster 3 than in Clusters 2 and 4, and Lichenibacterium sequences were more abundant in Cluster 3 than in Clusters 1 and 2. Cluster 2 contained significantly more ASV reads affiliated with Streptococcus, Veillonella (class Negativicutes) and Rothia genera (class Actinobacteria) than other clusters. Additionally, the higher abundance of Haemophilus and Lactobacillus distinguished Cluster 2 from 3 and 4, and the Escherichia-Shigella genus from Clusters 1 and 4. A significantly higher abundance of two cyanobacterial taxa, Aliterella and Chamaesiphon PCC-7430, was detected in Cluster 4.The higher abundances of cyanobacterial CENA359 and Acidothermus genera differentiated Cluster 4 from 1 and 2, and Polymorphobacter from 2 and 3. As for Cluster 1, two sequences from Gammaproteobacteria class, affiliated with the unknown genus from the Lysobacteraceae family and the Oxalicibacterium genus, as well as Cryomorpha and Pricia genera (Bacteroidia class), showed higher sequence read counts than the others.
Based on Venn diagram visualization (Fig. S4), the most significant differences at genera level were observed between Cluster 2 and the other snow clusters. Among 46 genera considered, 41 significantly differentiated this cluster from Cluster 3, and 39 and 30 with Clusters 4 and 1, respectively.
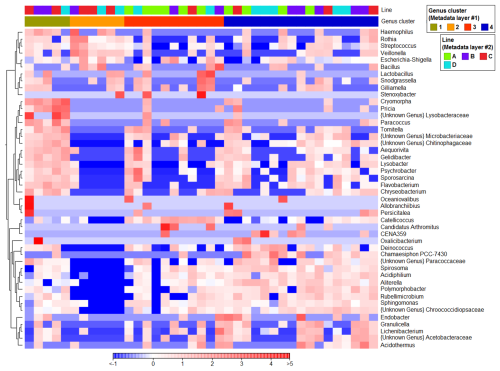
Figure 4Normalized distribution of the 46 most abundant genera (based on read abundances) that significantly (P<0.05) differentiated snow clusters. Samples are ordered based on clusters derived from PCoA, while transect information is provided in the top bar (“Line”). Colors represent scaled (normalized) relative abundances, where red indicates higher and blue lower abundance relative to the mean across samples (z-score scaling).
3.3 Exploring relationships between bacterial community structure and snow environmental variables – canonical correspondence analysis (CCA)
Canonical correspondence analysis (CCA) was used to relate variation in bacterial community structure with snow physicochemical variables (Figs. 5, S5). The top two axes (CCA 1 and 2) were included and accounted for 31.03 % and 16.18 % of microbial genera sequence structure variation, respectively.
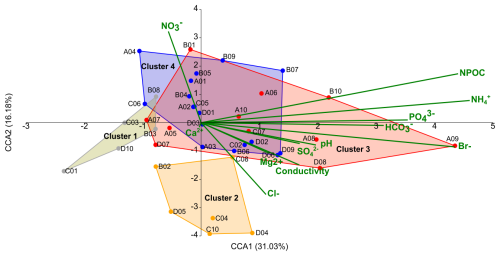
Figure 5Canonical correspondence analysis (CCA) of bacterial profiles at the genus level in snow clusters. The biplot presents the correlation between physicochemical properties of snow and the microbial community structure. Bold green lines represent the physicochemical parameters; the line length indicates which physicochemical parameters determine the cluster distribution the most. Four snow clusters are marked by gold (Cluster 1), orange (Cluster 2), red (Cluster 3) and blue (Cluster 4) colours.
In Cluster 1, marker genera such as Pricia, Cryomorpha, Oxalicibacterium and members of the Lysobacteraceae (Fig. S5, marked in gold) covaried with low nutrient availability and generally low concentrations of measured physicochemical parameters (Fig. 5). The abundance of markers of Cluster 2: Streptococcus, Veillonella, Rothia, Haemophilus, Lactobacillus and Escherichia–Shigella (Fig. S5, marked in orange), was positively correlated with higher salinity-related parameters (represented in the analysis by Cl−, SO and conductivity) and lower NO concentrations. In Cluster 3, the presence of Catellicoccus and Candidatus Arthromitus markers (Fig. S5, in red) was correlated with nutrient-enriched snow conditions, including higher concentrations of organic carbon (non-purgeable organic carbon, NPOC), NH, and anions PO, HCO, and Br−, as well as elevated pH values, representing a set of interrelated variables linked to nutrient enrichment. In Cluster 4, marker genera including Aliterella, Chamaesiphon PCC-7430, CENA359 and Polymorphobacter were associated with a broader nitrogen-related environmental gradient, including inorganic nutrient availability. Additionally, selected marker genera, such as Oxalicibacterium, Lichenibacterium, and Acidothermus, characteristic of Clusters 1, 3, and 4, respectively, showed a positive correlation with nitrate (NO) concentrations.
Snowpacks are complex ecosystems populated by diverse microorganisms, and their community composition is linked to the specific physicochemical properties of snow and the nutrients that snow provides (Maccario et al., 2015). As newly formed habitats, seasonal terrestrial snowpacks are colonized from the terrestrial surface upon which the snowpack develops or microorganisms from aerosols (Keuschnig et al., 2023; Maccario et al., 2019). In this study, we focus on the effects of vertebrates on the snow surface microbiome during early spring (pre-melt) in the High Arctic. As our approach is DNA-based, we do not infer activity.
4.1 Vertebrates as Key Contributors to Snow Microbial Communities
We showed that wild animals contribute substantially to the surface snow microbiomes, with bacteria of animal origin identified in almost half of our samples (Clusters 2 and 3). Successful colonization into snow communities by these microorganisms is unlikely due to the significant differences in ecosystem characteristics between snow and the host environment that may limit their survival. However, studies have shown that organisms that fail to establish long-term populations can still impact the recipient community regardless of their ability to proliferate or spread: for example, the genetic pool of ecosystems can be impacted through the release of free DNA, which can then be taken up by the local community (Amor et al., 2020; Collins and Deming, 2011). Furthermore, it is possible that these bacteria persist until the tundra surface becomes exposed, at which point they may re-enter the digestive tracts of grazing animals.
Cluster 3 was dominated by the Gram-positive bacteria Catellicoccus of the Lactobacillales order, a ubiquitous organism in the gut microbiome of many avian species that is used as a bird faecal indicator (Grond et al., 2018; Kreisinger et al., 2017). Catellicoccus genus is also a dominant bacteria in the gut microbiome of thick-billed murres (Uria lomvia) (Góngora et al., 2021), and was detected in the microbiome of ruddy turnstone (Arenaria interpres) (Grond et al., 2014, 2017). Similarly, the detection of Candidatus Arthromitus (Grond et al., 2017), despite its uneven distribution among samples, suggests that Arctic birds were the main source of microorganisms on the snow surface in Cluster 3. Psychrobacter, a cold-adapted Gram-negative aerobic bacteria commonly identified in ice, snow and frozen soils (Rodrigues et al., 2009; Zeng et al., 2013) was also detected in higher abundance in Cluster 3. These cold-adapted bacteria are commonly found in Arctic and Antarctic ornithogenic soils, which are derived from the deposition of the faecal matter of birds (Bowman et al., 1996; Lasek et al., 2017). Interestingly, Psychrobacter strains were also isolated from the guano of little auks collected in the vicinity of the Polish Polar Station Hornsund on Spitsbergen (Dziewit et al., 2013), which is consistent with our hypothesis. This interpretation is further supported by the physicochemical data showing correlation of organic carbon, ammonium ions (forming the majority of total and inorganic nitrogen in the samples), and phosphate concentrations and pH with the relative abundance of Catellicoccus, likely showing their joint origin in bird droppings, as Catellicoccus has been found in faecal matter of e.g. gull (Koskey et al., 2014; Yao et al., 2023), and the chemical constituents are also characteristic for guano. This bird-related nutrient enrichment of the Arctic tundra has been shown in multiple locations, including the little auk colony at the study location (Finne et al., 2024; Skrzypek et al., 2015; Zwolicki et al., 2016).
Cluster 2 had the lowest diversity of all clusters, which could be explained by the presence of a few dominant organisms. Genera detected in higher abundance in this cluster were markers of the bird or animal microbiome (including human). The prevalence of animal-derived genera, corroborated by a heatmap and a Venn diagram, confirmed that Cluster 2 is the most unique grouping within the snow microbiome. For instance, Streptococcus, another Lactobacillales member common in migratory bird guts (Kreisinger et al., 2017) as well as animals like reindeer (Aagnes et al., 1995; Sundset et al., 2007) or fish (Ringø et al., 2018), was correlated with a strict anaerobic bacteria Veillonella (Fig. S5). Both bacterial genera were described as frequently co-occurring members of the human and animal gut microbiome (van den Bogert et al., 2013; Lu et al., 2020) or were detected in birds that live in Svalbard during summer like snow bunting (Plectrophenax nivalis), sanderling (Calidris alba) (Cho and Young, 2020) or all-season rock ptarmigans (Lagopus muta hyperborea) (Ushida et al., 2016). Veillonella were also observed in a small proportion in the tundra river system in Svalbard (Kosek et al., 2018), glacial snow and soil (Yang et al., 2016) or in the inner fjord Kongsfjorden (Jain and Krishnan, 2017). Another significantly more abundant genus was Escherichia-Shigella (these two genera cannot be distinguished solely on the basis of the 16S rRNA gene sequence), often identified in mammal, avian and human microbiomes (Franz et al., 2022; Góngora et al., 2021; Kamenova et al., 2023; Nekrasova et al., 2024). A similar situation was noted for Rothia, previously observed in samples from the oral and gut microbiomes of birds (Abolnik et al., 2021; Cho and Young, 2020) or humans (Buetas et al., 2024; Yamane et al., 2010). Based on our analysis, these genera were mainly associated with higher salinity indices (multiple correlated ionic concentrations, represented in the CCA graph by conductivity, Cl−, Mg2+, Ca2+, Br− and SO concentrations) and negatively correlated with NO anions, suggesting a high tolerance for saline stress, which might support their survival in harsh environments (Zhang et al., 2021). These salinity-related parameters were strongly correlated with each other, suggesting a shared marine aerosol signal rather than independent effects of individual ions. The elevated salinity of snow may be due to the sea spray impact, which is very typical for snowpacks in the Hornsund area of Svalbard (Barbaro et al., 2021).
While the community composition of these two groups was strongly associated with vertebrate-derived inputs, other colonization sources were also likely. For example, in Cluster 3, the high relative abundance of Lichenibacterium genus (Alphaproteobacteria class), which is often linked to lichen-associated Rhizobiales, suggests a lichen origin (Pankratov et al., 2020) that may have been sourced from the snow free, lichen covered rocks observed near our study sites. Other bacteria, like Gelidibacter or Aequorivita, are also often found in marine and polar environments (Amores-Arrocha et al., 2023; Groudieva et al., 2004).
4.2 Bacterial taxa distribution in non-animal impacted snow
Clusters 1 and 4 exhibited minimal animal influence, instead reflecting microbial communities typical of Arctic snow environments. These clusters displayed the highest species diversity (Fig. S3), significantly exceeding values reported for other Spitsbergen snow samples (Keuschnig et al., 2023; Thomas et al., 2020). Dominant taxa in Cluster 4 included cold-adapted genera such as Aliterella (Cluster 4 key indicator), cyanobacteria previously identified in Antarctic seawater (Rigonato et al., 2016), Atacama Desert rocks (Jung et al., 2020), and lichen photobionts (Jung et al., 2021). Additional cyanobacteria in Cluster 4, Chamaesiphon, is known from polar lake microbial mats (Pessi et al., 2023) and riverine rocks (Aigner et al., 2018; Nemergut et al., 2007), suggesting wind dispersal from local terrestrial or aquatic habitats (Harding et al., 2011; Jensen et al., 2022). Other sequences characteristic for Cluster 4 were classified to CENA359, an undescribed cyanobacterial strain from environmental 16S rRNA gene surveys, previously detected in diverse aquatic and extreme environments and affiliated with the family Leptolyngbyaceae (Kamran et al., 2021), to Acidothermus, detected in Arctic soil (Christiansen et al., 2025; Semenova et al., 2025), and to psychrotolerant bacteria Polymorphobacter, previously isolated from glacial ice, (Thomas et al., 2020) as well as from Arctic lithic niches (Choe et al., 2021).
Cluster 4 was associated with elevated nitrate concentrations and dominated by cyanobacterial taxa. Free-living cyanobacteria are known to actively fix atmospheric N2 in High Arctic tundra ecosystems, including coastal sites on Spitsbergen, with fixed nitrogen initially incorporated into organic biomass (Jungblut et al., 2010). The observed nitrate in snow may therefore reflect indirect processes such as mineralization and subsequent nitrification, or alternative sources including atmospheric deposition or oxidized ammonium of fecal origin (Rousk et al., 2017). Disentangling these potential sources would require additional analyses, such as stable nitrogen isotope measurements (δ15N) and transcriptomic studies to establish microbial activity, which were beyond the scope of the present study. As key nitrogen fixers in Arctic ecosystems (Liengen and Olsen, 1997; Solheim et al., 1996), cyanobacteria play critical roles in soil formation, biocrust development, and aquatic primary production (Patova et al., 2016; Rousk et al., 2017), while also serving as ecosystem engineers through formation of bioaggregates from organic and mineral matter on glaciers (Rozwalak et al., 2022; Wejnerowski et al., 2023). Collectively, these patterns suggest Cluster 4 represents a characteristic snow microbiome shaped by wind-dispersed tundra materials containing nitrogen-cycling taxa.
Cluster 1 differed from Cluster 4, not only by a significantly lower content of Chamaesiphon PCC-7430, CENA359, Aliterella, and Acidothermus, but also by a significantly higher relative abundance of the unknown genus from the Lysobacteraceae. Members of this family, belonging to Gammaproteobacteria class, were isolated from Arctic permafrost soils (Höfle et al., 2015). Another member of this class, Oxalicibacterium, included strain isolates from different environments like soil or plant litter (Cavanaugh et al., 2006). Furthermore, Cryomorpha and Pricia members (Bacteroidia) were isolated from Antarctic intertidal sediments or South Ocean samples particulates and from quartz stone subliths (Bowman et al., 2003; Yu et al., 2012). These cold-adapted taxa suggest airborne dispersal as the primary colonization mechanism, mirroring Cluster 4. The abundance of Gammaproteobacteria and Bacteroidia correlated with lower nutrient and pH levels (Figs. 5, S5), suggesting tolerance to oligotrophic conditions. These phyla are well-documented in Arctic tundra soils (Kosek et al., 2017; Nissinen et al., 2012) and Hornsund river systems (Kosek et al., 2019). For instance, genus Pricia thrives in nutrient-poor, oxygen-limited settings (Liu et al., 2021). Moreover, reindeer feces may represent an additional source, as several genera (Cryobacterium, and Bacteroidia members) have been identified in fecal samples from Ny-Ålesund (Fang et al., 2024) and Hornsund (Zielińska et al., 2016a).
Overall, the strong correlations observed among multiple environmental variables indicate that microbial community patterns should be interpreted as responses to combined environmental gradients (e.g., faecal nutrient enrichment and marine aerosol input) rather than to individual physicochemical parameters.
Our study shows that vertebrates influence the bacterial community structure in springtime Arctic snowpacks in coastal areas. In addition to identifying cold-adapted organisms typically associated with psychrophilic environments, we also detected microbes of vertebrate origin. These were mainly linked to the gut microbiome of birds and terrestrial mammals, including humans. Although further studies are needed to determine whether these organisms are active and able to survive in the snow, sequences classified to Catellicoccus and Streptococcus genera indicated vertebrate activity and nutrient enrichment. Physicochemical properties of snow, especially salinity, pH, and nutrient concentrations, were impacted by both vertebrates and marine aerosols, and they likely influenced the snow microbiome. Our study indicates that animals are an important source of microorganisms to the snow surface. Overall, our findings indicate that in addition to airborne transport factors such as sea spray and wind transport from snow-free terrestrial environments, microbial composition in Arctic coastal snowpacks is more dependent on wildlife than previously assumed. Therefore, both abiotic and biotic transport factors should be considered when assessing microbial ecosystem dynamics in polar regions.
The bioinformatics workflow is implemented in a custom R script based on the DADA2 pipeline. The script can be accessed directly as part of the Supplementary Information associated with this article and enables full reproducibility of the sequence processing steps.
The amplicon sequencing dataset, including blank controls, has been deposited in the NCBI Sequence Read Archive (SRA) under accession number PRJNA1277323. Processed data, including the ASV table and associated sample metadata, as well as physicochemical measurements of snow samples, are provided in the Supplement as Supplementary Excel 1 (NGS data) and Supplementary Excel 2 (CCA data), respectively. All data are available in formats that allow full reproduction of the analyses presented.
The supplement related to this article is available online at https://doi.org/10.5194/bg-23-3023-2026-supplement.
SS: Conceptualization, Formal analysis, Investigation, Resources, Data Curation, Writing – Original Draft, Writing – Review & Editing, Visualization; KZ: Conceptualization, Investigation, Resources, Writing – Original Draft, Writing – Review & Editing; AM: Investigation, Writing – Review & Editing; KK: Conceptualization, Investigation, Resources, Writing – Review & Editing; WZ: Investigation, Writing – Review & Editing; AN: Conceptualization, Investigation; KT: Investigation; CK: Formal analysis, Methodology, Software, Validation; BL: Conceptualization, Visualization, Resources, Writing – Review & Editing, Supervision, Project administration, Funding acquisition; CL: Conceptualization, Formal analysis, Investigation, Resources, Writing – Original Draft, Writing – Review & Editing.
The contact author has declared that none of the authors has any competing interests.
Publisher's note: Copernicus Publications remains neutral with regard to jurisdictional claims made in the text, published maps, institutional affiliations, or any other geographical representation in this paper. The authors bear the ultimate responsibility for providing appropriate place names. Views expressed in the text are those of the authors and do not necessarily reflect the views of the publisher.
This article is part of the special issue “Cryospheric ecosystems: climate feedback loops, threatened ecosystems, and consequences of climate change”. It is a result of the Cryospheric ecosystems conference, Poznań, Poland, 1–4 September 2025.
We gratefully acknowledge the team of the Polish Polar Station Hornsund for their logistical support during fieldwork campaign. We also thank the reviewers for their constructive and valuable comments, which helped improve the quality of this manuscript.
This research has been supported by the EEA Grants/Norway Grants (project: HarSval – Bilateral initiative aiming at Harmonisation of the Svalbard cooperation, grant no. UMO-2023/43/7/ST10/00001).
This paper was edited by Helge Niemann and reviewed by Julia Engelmann and one anonymous referee.
Aagnes, T. H., Sormo, W., and Mathiesen, S. D.: Ruminal microbial digestion in free-living, in captive lichen-fed, and in starved reindeer (Rangifer tarandus tarandus) in winter, Appl. Environ. Microbiol., 61, 583–591, https://doi.org/10.1128/aem.61.2.583-591.1995, 1995.
Abolnik, C., Strydom, C., Landman, D., and Pieterse, R.: Identification of bacteria in the tracheal swabs of farmed ostriches and their effect on the viability of influenza A virus, J. Vet. Diagnostic Investig., 33, 1089–1095, https://doi.org/10.1177/10406387211034483, 2021.
Aigner, S., Herburger, K., Holzinger, A., and Karsten, U.: Epilithic Chamaesiphon (Synechococcales, Cyanobacteria) species in mountain streams of the Alps – interspecific differences in photo-physiological traits, J. Appl. Phycol., 30, 1125–1134, https://doi.org/10.1007/s10811-017-1328-7, 2018.
Amato, P., Hennebelle, R., Magand, O., Sancelme, M., Delort, A. M., Barbante, C., Boutron, C., and Ferrari, C.: Bacterial characterization of the snow cover at Spitzberg, Svalbard, FEMS Microbiol. Ecol., 59, 255–264, https://doi.org/10.1111/j.1574-6941.2006.00198.x, 2007.
Amor, D. R., Ratzke, C., and Gore, J.: Transient invaders can induce shifts between alternative stable states of microbial communities, Sci. Adv., 6, 1–9, https://doi.org/10.1126/sciadv.aay8676, 2020.
Amores-Arrocha, H., Asamoah-Asare, A. K. B., Opio, J., Martin, A., Cuthbertson, L., Bradford, H. R., Avila-Jimenez, M. L., and Pearce, D. A.: Analysis of Bacterial Communities around the Adventdalen Landfill Site in Svalbard, Microorganisms, 11, https://doi.org/10.3390/microorganisms11041093, 2023.
Barbaro, E., Koziol, K., Björkman, M. P., Vega, C. P., Zdanowicz, C., Martma, T., Gallet, J.-C., Kępski, D., Larose, C., Luks, B., Tolle, F., Schuler, T. V., Uszczyk, A., and Spolaor, A.: Measurement report: Spatial variations in ionic chemistry and water-stable isotopes in the snowpack on glaciers across Svalbard during the 2015–2016 snow accumulation season, Atmos. Chem. Phys., 21, 3163–3180, https://doi.org/10.5194/acp-21-3163-2021, 2021.
Bauer, S. and Hoye, B. J.: Migratory animals couple biodiversity and ecosystem functioning worldwide, Science, 344, https://doi.org/10.1126/science.1242552, 2014.
Bergk Pinto, B., Maccario, L., Dommergue, A., Vogel, T. M., and Larose, C.: Do Organic Substrates Drive Microbial Community Interactions in Arctic Snow?, Front. Microbiol., 10, 1–13, https://doi.org/10.3389/fmicb.2019.02492, 2019.
Bokulich, N. A., Subramanian, S., Faith, J. J., Gevers, D., Gordon, J. I., Knight, R., Mills, D. A., and Caporaso, J. G.: Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing, Nat. Methods, 10, 57–59, https://doi.org/10.1038/nmeth.2276, 2013.
Bowman, J. P., Cavanagh, J., Austin, J. J., and Sanderson, K.: Novel Psychrobacter species from Antarctic ornithogenic soils, Int. J. Syst. Bacteriol., 46, 841–848, https://doi.org/10.1099/00207713-46-4-841, 1996.
Bowman, J. P., Nichols, C. M., and Gibson, J. A. E.: Algoriphagus ratkowskyi gen. nov., sp. nov., Brumimicrobium glaciale gen. nov., sp. nov., Cryomorpha ignava gen. nov., sp. nov. and Crocinitomix catalasitica gen. nov., sp. nov., novel flavobacteria isolated from various polar habitats, Int. J. Syst. Evol. Microbiol., 53, 1343–1355, https://doi.org/10.1099/ijs.0.02553-0, 2003.
Buetas, E., Jordán-López, M., López-Roldán, A., D'Auria, G., Martínez-Priego, L., De Marco, G., Carda-Diéguez, M., and Mira, A.: Full-length 16S rRNA gene sequencing by PacBio improves taxonomic resolution in human microbiome samples, BMC Genomics, 25, 1–19, https://doi.org/10.1186/s12864-024-10213-5, 2024.
Callahan, B. J., McMurdie, P. J., Rosen, M. J., Han, A. W., Johnson, A. J. A., and Holmes, S. P.: DADA2: High resolution sample inference from Illumina amplicon data, Nat. Methods, 13, 581–583, https://doi.org/10.1038/nmeth.3869, 2016.
Cavanaugh, C. M., Mckiness, Z. O. E. P., Newton, I. L. G., and Stewart, F. J.: The Prokaryotes, 297–321, https://doi.org/10.1007/0-387-30745-1, 2006.
Chen, J., Bittinger, K., Charlson, E. S., Hoffmann, C., Lewis, J., Wu, G. D., Collman, R. G., Bushman, F. D., and Li, H.: Associating microbiome composition with environmental covariates using generalized UniFrac distances, Bioinformatics, 28, 2106–2113, https://doi.org/10.1093/bioinformatics/bts342, 2012.
Cho, H. and Young, W.: Interspecific comparison of the fecal microbiota structure in three Arctic migratory bird species, Ecol. Evol., 10, 5582–5594, https://doi.org/10.1002/ece3.6299, 2020.
Choe, Y. H., Kim, M., and Lee, Y. K.: Distinct Microbial Communities in Adjacent Rock and Soil Substrates on a High Arctic Polar Desert, Front. Microbiol., 11, 1–15, https://doi.org/10.3389/fmicb.2020.607396, 2021.
Christiansen, C. T., Engel, K., Hall, M., Neufeld, J. D., Walker, V. K., and Grogan, P.: Arctic tundra soil depth, more than seasonality, determines active layer bacterial community variation down to the permafrost transition, Soil Biol. Biochem., 200, 109624, https://doi.org/10.1016/j.soilbio.2024.109624, 2025.
Chuvochina, M., Gerken, J., Frentrup, M., Sandikci, Y., Goldmann, R., Freese, H. M., Göker, M., Sikorski, J., Yarza, P., Quast, C., Peplies, J., Glöckner, F. O., and Reimer, L. C.: SILVA in 2026: a global core biodata resource for rRNA within the DSMZ digital diversity, Nucleic Acids Res., 334–341, https://doi.org/10.1093/nar/gkaf1247, 2025.
Collins, R. E. and Deming, J. W.: Abundant dissolved genetic material in Arctic sea ice Part I: Extracellular DNA, Polar Biol., 34, 1819–1830, https://doi.org/10.1007/s00300-011-1041-y, 2011.
Déry, S. J. and Brown, R. D.: Recent Northern Hemisphere snow cover extent trends and implications for the snow-albedo feedback, Geophys. Res. Lett., 34, 2–7, https://doi.org/10.1029/2007GL031474, 2007.
Dong, Z., Jiang, H., Baccolo, G., Di Mauro, B., and Zawierucha, K.: Biological and Pollution Aerosols on Snow and Ice – Interplay between the Atmosphere and the Cryosphere, J. Earth Sci., 34, 1951–1956, https://doi.org/10.1007/s12583-023-2004-2, 2023.
Dziewit, L., Cegielski, A., Romaniuk, K., Uhrynowski, W., Szych, A., Niesiobedzki, P., Zmuda-Baranowska, M. J., Zdanowski, M. K., and Bartosik, D.: Plasmid diversity in arctic strains of Psychrobacter spp., Extremophiles, 17, 433–444, https://doi.org/10.1007/s00792-013-0521-0, 2013.
Faith, D. P. and Baker, A. M.: Phylogenetic Diversity (PD) and Biodiversity Conservation: Some Bioinformatics Challenges, Evol. Bioinforma., 2, 117693430600200, https://doi.org/10.1177/117693430600200007, 2006.
Fang, X. M., Li, J., Wang, N. F., Zhang, T., and Yu, L. Y.: Fziel, Environ. Res., 262, 119788, https://doi.org/10.1016/j.envres.2024.119788, 2024.
Finne, E. A., Varpe, Ø., Durant, J. M., Gabrielsen, G. W., and Poste, A. E.: Nutrient fluxes from an Arctic seabird colony to the adjacent coastal marine ecosystem, Polar Biol., 47, 859–872, https://doi.org/10.1007/s00300-022-03024-5, 2024.
Franz, M., Whyte, L., Atwood, T. C., Laidre, K. L., Roy, D., Watson, S. E., Góngora, E., and McKinney, M. A.: Distinct gut microbiomes in two polar bear subpopulations inhabiting different sea ice ecoregions, Sci. Rep., 12, 1–15, https://doi.org/10.1038/s41598-021-04340-2, 2022.
Franzetti, A., Navarra, F., Tagliaferri, I., Gandolfi, I., Bestetti, G., Minora, U., Azzoni, R. S., Diolaiuti, G., Smiraglia, C., and Ambrosini, R.: Potential sources of bacteria colonizing the cryoconite of an Alpine glacier, PLoS One, 12, 1–13, https://doi.org/10.1371/journal.pone.0174786, 2017.
Góngora, E., Elliott, K. H., and Whyte, L.: Gut microbiome is affected by inter-sexual and inter-seasonal variation in diet for thick-billed murres (Uria lomvia), Sci. Rep., 11, 1–12, https://doi.org/10.1038/s41598-020-80557-x, 2021.
Grond, K., Ryu, H., Baker, A. J., Santo, J. W., and Deborah, D.: Gastro-intestinal microbiota of two migratory shorebird species during spring migration staging in Delaware Bay , USA, J. Ornithol., 155, 969–977, https://doi.org/10.1007/s10336-014-1083-3, 2014.
Grond, K., Lanctot, R. B., Jumpponen, A., and Sandercock, B. K.: Recruitment and establishment of the gut microbiome in arctic shorebirds, FEMS Microbiol. Ecol., 93, 1–9, https://doi.org/10.1093/femsec/fix142, 2017.
Grond, K., Sandercock, B. K., Jumpponen, A., and Zeglin, L. H.: The avian gut microbiota: community, physiology and function in wild birds, J. Avian Biol., 49, 1–19, https://doi.org/10.1111/jav.01788, 2018.
Groudieva, T., Kambourova, M., Yusef, H., Royter, M., Grote, R., Trinks, H., and Antranikian, G.: Diversity and cold-active hydrolytic enzymes of culturable bacteria associated with Arctic sea ice, Spitzbergen, Extremophiles, 8, 475–488, https://doi.org/10.1007/s00792-004-0409-0, 2004.
Hammer, Ø., Harper, D. A. T., and Ryan, P. D.: PAST: Paleontological statistics software package for education and data analysis, 4, 9 pp., http://palaeo-electronica.org/2001_1/past/issue1_01.htm (last access: 27 March 2026), 2001.
Harding, T., Jungblut, A. D., Lovejoy, C., and Vincent, W. F.: Microbes in high arctic snow and implications for the cold biosphere, Appl. Environ. Microbiol., 77, 3234–3243, https://doi.org/10.1128/AEM.02611-10, 2011.
Hayashi, K., Tanabe, Y., Ono, K., Loonen, M. J. J. E., Asano, M., Fujitani, H., Tokida, T., Uchida, M., and Hayatsu, M.: Seabird-affected taluses are denitrification hotspots and potential N2O emitters in the High Arctic, Sci. Rep., 8, 1–11, https://doi.org/10.1038/s41598-018-35669-w, 2018.
Herlemann, D. P. R., Labrenz, M., Jürgens, K., Bertilsson, S., Waniek, J. J., and Andersson, A. F.: Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea, ISME J., 5, 1571–1579, https://doi.org/10.1038/ismej.2011.41, 2011.
Höfle, S. T., Kusch, S., Talbot, H. M., Mollenhauer, G., Zubrzycki, S., Burghardt, S., and Rethemeyer, J.: Characterisation of bacterial populations in Arctic permafrost soils using bacteriohopanepolyols, Org. Geochem., 88, 1–16, https://doi.org/10.1016/j.orggeochem.2015.08.002, 2015.
Hoham, R. W. and Remias, D.: Snow and Glacial Algae: A Review, J. Phycol., 56, 264–282, https://doi.org/10.1111/jpy.12952, 2020.
Hudson, A.: Biogeochemistry of snowmelt in an Antarctic glacial ecosystem, Water Resour. Res., 42, 1–15, https://doi.org/10.1029/2005WR004311, 2006.
Jain, A. and Krishnan, K. P.: Differences in free-living and particle-associated bacterial communities and their spatial variation in Kongsfjorden, Arctic, J. Basic Microbiol., 57, 827–838, https://doi.org/10.1002/jobm.201700216, 2017.
Jakubas, D., Zmudczyńska, K., Wojczulanis-Jakubas, K., and Stempniewicz, L.: Faeces deposition and numbers of vertebrate herbivores in the vicinity of planktivorous and piscivorous seabird colonies in Hornsund, Spitsbergen, Polish Polar Res., 29, 45–58, 2008.
Jensen, L. Z., Glasius, M., Gryning, S. E., Massling, A., Finster, K., and Šantl-Temkiv, T.: Seasonal Variation of the Atmospheric Bacterial Community in the Greenlandic High Arctic Is Influenced by Weather Events and Local and Distant Sources, Front. Microbiol., 13, 1–13, https://doi.org/10.3389/fmicb.2022.909980, 2022.
Jung, P., Mikhailyuk, T., Emrich, D., Baumann, K., and Dultz, S.: Shifting Boundaries: Ecological and Geographical Range extension Based on Three New Species in the Cyanobacterial Genera Cyanocohniella, Oculatella, and, Aliterella, J. Phycol., 56, 1216–1231, https://doi.org/10.1111/jpy.13025, 2020.
Jung, P., Brust, K., Schultz, M., Büdel, B., Donner, A., Lakatos, M., Ruprecht, U., Beck, A., and Orlando, J.: Opening the Gap: Rare Lichens With Rare Cyanobionts – Unexpected Cyanobiont Diversity in Cyanobacterial Lichens of the Order Lichinales, Front. Microbiol., 12, 728378, https://doi.org/10.3389/fmicb.2021.728378, 2021.
Jungblut, A. D., Lovejoy, C., and Vincent, W. F.: Global distribution of cyanobacterial ecotypes in the cold biosphere, ISME J., 4, 191–202, https://doi.org/10.1038/ismej.2009.113, 2010.
Kamenova, S., de Muinck, E. J., Veiberg, V., Utsi, T. A., Steyaert, S. M. J. G., Albon, S. D., Loe, L. E., and Trosvik, P.: Gut microbiome biogeography in reindeer supersedes millennia of ecological and evolutionary separation, FEMS Microbiol. Ecol., 99, 1–12, https://doi.org/10.1093/femsec/fiad157, 2023.
Kamran, A., Sauter, K., Reimer, A., Wacker, T., Reitner, J., and Hoppert, M.: Cyanobacterial mats in calcite-precipitating serpentinite-hosted alkaline springs of the Voltri massif, Italy, Microorganisms, 9, 1–20, https://doi.org/10.3390/microorganisms9010062, 2021.
Keslinka, L. K., Wojczulanis-Jakubas, K., Jakubas, D., and Neubauer, G.: Determinants of the little auk (Alle alle) breeding colony location and size in W and NW coast of Spitsbergen, PLoS One, 14, 1–20, https://doi.org/10.1371/journal.pone.0212668, 2019.
Keuschnig, C., Vogel, T. M., Barbaro, E., Spolaor, A., Koziol, K., Björkman, M. P., Zdanowicz, C., Gallet, J. C., Luks, B., Layton, R., and Larose, C.: Selection processes of Arctic seasonal glacier snowpack bacterial communities, Microbiome, 11, 35, https://doi.org/10.1186/s40168-023-01473-6, 2023.
Kosek, K., Jankowska, K., and Polkowska, Ż. A.: Bacterial presence in polar regions associated with environment modification by chemical compounds including contaminants, Environ. Rev., 25, 481–491, 2017.
Kosek, K., Luczkiewicz, A., Kozioł, K., Jankowska, K., Ruman, M., and Polkowska, Ż.: Environmental characteristics of a tundra river system in Svalbard. Part 1: Bacterial abundance, community structure and nutrient levels, Sci. Total Environ., 653, 1571–1584, https://doi.org/10.1016/j.scitotenv.2018.11.378, 2018.
Kosek, K., Kozioł, K., Luczkiewicz, A., Jankowska, K., Chmiel, S., and Polkowska, Ż.: Environmental characteristics of a tundra river system in Svalbard. Part 2: Chemical stress factors, Sci. Total Environ., 653, 1585–1596, https://doi.org/10.1016/j.scitotenv.2018.11.012, 2019.
Koskey, A. M., Fisher, J. C., Traudt, M. F., Newton, R. J., and McLellan, S. L.: Analysis of the Gull Fecal Microbial Community Reveals the Dominance of Catellicoccus marimammalium in Relation to Culturable Enterococci, Appl. Environ. Microbiol., 80, 757–765, https://doi.org/10.1128/AEM.02414-13, 2014.
Kreisinger, J., Kropáčková, L., Petrželková, A., Adámková, M., Tomášek, O., Martin, J. F., Michálková, R., and Albrecht, T.: Temporal stability and the effect of transgenerational transfer on fecal microbiota structure in a long distance migratory bird, Front. Microbiol., 8, 1–19, https://doi.org/10.3389/fmicb.2017.00050, 2017.
Larose, C., Berger, S., Ferrari, C., Navarro, E., Dommergue, A., Schneider, D., and Vogel, T. M.: Microbial sequences retrieved from environmental samples from seasonal Arctic snow and meltwater from Svalbard, Norway, Extremophiles, 14, 205–212, https://doi.org/10.1007/s00792-009-0299-2, 2010.
Larose, C., Dommergue, A., and Vogel, T. M.: The dynamic Arctic snow pack: An unexplored environment for microbial diversity and activity, Biology (Basel)., 2, 317–330, https://doi.org/10.3390/biology2010317, 2013.
Lasek, R., Dziewit, L., Ciok, A., Decewicz, P., Romaniuk, K., Jedrys, Z., Wibberg, D., Schlüter, A., Pühler, A., and Bartosik, D.: Genome content, metabolic pathways and biotechnological potential of the psychrophilic Arctic bacterium Psychrobacter sp. DAB_AL43B, a source and a host of novel Psychrobacter-specific vectors, J. Biotechnol., 263, 64–74, https://doi.org/10.1016/j.jbiotec.2017.09.011, 2017.
Lemke, P., Ren, J., Alley, R. B., Allison, I., Carrasco, J., Flato, G., Fujii, Y., Kaser, G., Mote, P., Thomas, R. H., and Zhang, T.: Observations: Changes in Snow, Ice and Frozen Ground, in: Climate Change 2007: The Physical Science Basis. Contribution of Working Group I to the Fourth Assessment Report of the Intergovernmental Panel on Climate Change, edited by: Solomon, S., Qin, D., Manning, M., Chen, Z., Marquis, M., Averyt, K., Tignor, M., and Miller, H. L., Cambridge University Press, Cambridge, United Kingdom and New York, NY, USA, 337–383, https://doi.org/10.1016/j.jmb.2004.10.032, 2007.
Liengen, T. and Olsen, R. A.: Nitrogen fixation by free-living cyanobacteria from different coastal sites in a high archc tundra, Spitsbergen, Arct. Alp. Res., 29, 470–477, https://doi.org/10.2307/1551994, 1997.
Liu, Q., Li, W., Liu, D., Li, L., Li, J., Lv, N., Liu, F., Zhu, B., Zhou, Y., Xin, Y., and Dong, X.: Light stimulates anoxic and oligotrophic growth of glacial Flavobacterium strains that produce zeaxanthin, ISME J., 15, 1844–1857, https://doi.org/10.1038/s41396-020-00891-w, 2021.
Lu, H., Yan, H., Masey O'Neill, H. M., Bradley, C., Bedford, M. R., Wilcock, P., Nakatsu, C. H., Adeola, O., and Ajuwon, K. M.: Effect of timing of postweaning xylanase supplementation on growth performance, nutrient digestibility, and fecal microbial composition in weanling pigs, Can. J. Anim. Sci., 100, 27–36, https://doi.org/10.1139/cjas-2019-0021, 2020.
Maccario, L., Sanguino, L., Vogel, T. M., and Larose, C.: Snow and ice ecosystems: Not so extreme, Res. Microbiol., 166, 782–795, https://doi.org/10.1016/j.resmic.2015.09.002, 2015.
Maccario, L., Carpenter, S. D., Deming, J. W., Vogel, T. M., and Larose, C.: Sources and selection of snow-specific microbial communities in a Greenlandic sea ice snow cover, Sci. Rep., 9, 1–14, https://doi.org/10.1038/s41598-019-38744-y, 2019.
Malard, L. A., Anwar, M. Z., Jacobsen, C. S., and Pearce, D. A.: Influence of Spatial Scale on Structure of Soil Bacterial Communities across an Arctic Landscape, Appl. Environ. Microbiol., 87, 1–16, https://doi.org/10.1128/AEM.02220-20, 2021.
Martin, M.: Cutadapt removes adapter sequences from high-throughput sequencing reads, EMBnet.journal Bioinforma. Action, 17, 10–12, 2011.
McInturf, A. G., Pollack, L., Yang, L. H., and Spiegel, O.: Vectors with autonomy: what distinguishes animal-mediated nutrient transport from abiotic vectors?, Biol. Rev., 94, 1761–1773, https://doi.org/10.1111/brv.12525, 2019.
Monteil, C. L., Guilbaud, C., Glaux, C., Lafolie, F., Soubeyrand, S., and Morris, C. E.: Emigration of the plant pathogen Pseudomonas syringae from leaf litter contributes to its population dynamics in alpine snowpack, Environ. Microbiol., 14, 2099–2112, https://doi.org/10.1111/j.1462-2920.2011.02680.x, 2012.
Myšková, E., Brož, M., Fuglei, E., Kvičerová, J., Mácová, A., Sak, B., Kváč, M., and Ditrich, O.: Gastrointestinal parasites of arctic foxes (Vulpes lagopus) and sibling voles (Microtus levis) in Spitsbergen, Svalbard, Parasitol. Res., 118, 3409–3418, https://doi.org/10.1007/s00436-019-06502-8, 2019.
Nekrasova, A. I., Kalashnikova, I. G., Korobeynikova, A. V., Ashniev, G. A., Bobrova, M. M., Bakoev, S. Y., Petryaikina, E. S., Nekrasov, A. S., Zagainova, A. V., Lukashina, M. V., Tolkacheva, L. R., Bobrovnitskii, I. P., Yudin, V. S., Keskinov, A. A., Makarov, V. V., and Yudin, S. M.: Characteristics of the Gut Microbiota Composition of the Arctic Zone Residents in the Far Eastern Region, Biomedicines, 12, https://doi.org/10.3390/biomedicines12112472, 2024.
Nemergut, D. R., Anderson, S. P., Cleveland, C. C., Martin, A. P., Miller, A. E., Seimon, A., and Schmidt, S. K.: Microbial community succession in an unvegetated, recently deglaciated soil, Microb. Ecol., 53, 110–122, https://doi.org/10.1007/s00248-006-9144-7, 2007.
Nissinen, R. M., Männistö, M. K., and van Elsas, J. D.: Endophytic bacterial communities in three arctic plants from low arctic fell tundra are cold-adapted and host-plant specific, FEMS Microbiol. Ecol., 82, 510–522, https://doi.org/10.1111/j.1574-6941.2012.01464.x, 2012.
Norwegian Polar Institute: Kartdata Svalbard 1:100 000 (S100 Kartdata)/Map Data, Norwegian Polar Institute [data set], https://doi.org/10.21334/NPOLAR.2014.645336C7, 2014.
Pankratov, T. A., Grouzdev, D. S., Patutina, E. O., Kolganova, T. V., Suzina, N. E., and Berestovskaya, J. J.: Lichenibacterium ramalinae gen. nov, sp. nov., Lichenibacterium minor sp. nov., the first endophytic, beta-carotene producing bacterial representatives from lichen thalli and the proposal of the new family Lichenibacteriaceae within the order Rhizobiales, Antonie van Leeuwenhoek, Int. J. Gen. Mol. Microbiol., 113, 477–489, https://doi.org/10.1007/s10482-019-01357-6, 2020.
Patova, E., Sivkov, M., and Patova, A.: Nitrogen fixation activity in biological soil crusts dominated by cyanobacteria in the Subpolar Urals (European North-East Russia), FEMS Microbiol. Ecol., 92, 1–9, https://doi.org/10.1093/femsec/fiw131, 2016.
Pessi, I. S., Popin, R. V., Durieu, B., Lara, Y., Tytgat, B., Savaglia, V., Roncero-Ramos, B., Hultman, J., Verleyen, E., Vyverman, W., and Wilmotte, A.: Novel diversity of polar Cyanobacteria revealed by genome-resolved metagenomics, Microb. Genomics, 9, 1–14, https://doi.org/10.1099/mgen.0.001056, 2023.
PN EN ISO 10304-1: Water quality – Determination of dissolved anions by liquid chromatography of ions Part 1: Determination of bromide, chloride, fluoride, nitrate, nitrite, phosphate and sulfate, 2009.
PN EN ISO 14911: Water quality – Determination of dissolved Li+, Na+, NH, K+, Mn2+, Ca2+, Mg2+, Sr2+ and Ba2+ using ion chromatography – Method for water and waste water, 2002.
Popiołek, M., Szczesna, J., Kotusz, J., Kusznierz, J., and Witkowski, A.: The level of infection with gastro-intestinal nematodes in Svalbard reindeers from Hornsund area, Spitsbergen, Polish Polar Res., 28, 277–282, 2007.
Qiagen: User Manual for CLC Microbial Genomics Module 24.0.1, 293 pp., https://resources.qiagenbioinformatics.com/manuals/clcmgm/2401/index.php?manual=Introduction.html (last access: 27 March 2026), 2024.
Rigonato, J., Gama, W. A., Alvarenga, D. O., Henrique, L., Branco, Z., Brandini, F. P., and Genu, D. B.: Aliterella atlantica gen. nov., sp. nov., and Aliterella antarctica sp. nov., novel members of coccoid Cyanobacteria, Int. J. Syst. Evol. Microbiol., 66, 2853–2861, https://doi.org/10.1099/ijsem.0.001066, 2016.
Ringø, E., Hoseinifar, S. H., Ghosh, K., Doan, H. Van, Beck, B. R., and Song, S. K.: Lactic acid bacteria in finfish-An update, Front. Microbiol., 9, 1–37, https://doi.org/10.3389/fmicb.2018.01818, 2018.
Rodrigues, D. F., Da C Jesus, E., Ayala-Del-Río, H. L., Pellizari, V. H., Gilichinsky, D., Sepulveda-Torres, L., and Tiedje, J. M.: Biogeography of two cold-adapted genera: Psychrobacter and Exiguobacterium, ISME J., 3, 658–665, https://doi.org/10.1038/ismej.2009.25, 2009.
Rousk, K., Sorensen, P. L., and Michelsen, A.: Nitrogen fixation in the High Arctic: a source of “new” nitrogen?, Biogeochemistry, 136, 213–222, https://doi.org/10.1007/s10533-017-0393-y, 2017.
Rozwalak, P., Podkowa, P., Buda, J., Niedzielski, P., Kawecki, S., Ambrosini, R., Azzoni, R. S., Baccolo, G., Ceballos, J. L., Cook, J., Di Mauro, B., Ficetola, G. F., Franzetti, A., Ignatiuk, D., Klimaszyk, P., Łokas, E., Ono, M., Parnikoza, I., Pietryka, M., Pittino, F., Poniecka, E., Porazinska, D. L., Richter, D., Schmidt, S. K., Sommers, P., Souza-Kasprzyk, J., Stibal, M., Szczuciński, W., Uetake, J., Wejnerowski, Ł., Yde, J. C., Takeuchi, N., and Zawierucha, K.: Cryoconite – From minerals and organic matter to bioengineered sediments on glacier's surfaces, Sci. Total Environ., 807, https://doi.org/10.1016/j.scitotenv.2021.150874, 2022.
Sanchez-Cid, C., Keuschnig, C., Vogel, T. M., and Larose, C.: Impact of in situ solar irradiation on snow bacterial communities and functional potential, FEMS Microbiol. Ecol., 99, 1–7, https://doi.org/10.1093/femsec/fiad042, 2023.
Segawa, T., Takeuchi, N., Rivera, A., Yamada, A., Yoshimura, Y., Barcaza, G., Shinbori, K., Motoyama, H., Kohshima, S., and Ushida, K.: Distribution of antibiotic resistance genes in glacier environments, Environ. Microbiol. Rep., 5, 127–134, https://doi.org/10.1111/1758-2229.12011, 2013.
Semenova, E. M., Babich, T. L., Sokolova, D. S., Myazin, V. A., Korneykova, M. V., and Nazina, T. N.: Bacterial Diversity of Arctic Soils with Long-Standing Pollution by Petroleum Products and Heavy Metals, Microorganisms, 14, 55, https://doi.org/10.3390/microorganisms14010055, 2025.
Skagseth, Ø., Furevik, T., Ingvaldsen, R., Loeng, H., Mork, K. A., Orvik, K. A., and Ozhigin, V.: Volume and heat transports to the Arctic Ocean via the Norwegian and Barents seas, in: Arctic–Subarctic Ocean Fluxes, edited by: Dickson, R. R., Meincke, J., and Rhines, P., Springer, Dordrecht, 45–64, https://doi.org/10.1007/978-1-4020-6774-7_3, 2008.
Skrzypek, G., Wojtuń, B., Richter, D., Jakubas, D., Wojczulanis-Jakubas, K., and Samecka-Cymerman, A.: Diversification of nitrogen sources in various tundra vegetation types in the high arctic, PLoS One, 10, 1–21, https://doi.org/10.1371/journal.pone.0136536, 2015.
Solheim, B., Endal, A., and Vigstad, H.: Nitrogen fixation in Arctic vegetation and soils from Svalbard, Norway, Polar Biol., 16, 35–40, https://doi.org/10.1007/BF02388733, 1996.
Souza-Kasprzyk, J., Paiva, T. de C., Convey, P., da Cunha, L. S. T., Soares, T. A., Zawierucha, K., Costa, E. S., Niedzielski, P., and Torres, J. P. M.: Influence of marine vertebrates on organic matter, phosphorus and other chemical element levels in Antarctic soils, Polar Biol., 45, 1571–1580, https://doi.org/10.1007/s00300-022-03091-8, 2022.
Stempniewicz, L.: Biomass of Dovekie Excreta in the Vicinity of a Breeding Colony, Colon. Waterbirds, 13, 62–66, 1990.
Stempniewicz, L.: Polar bears observed climbing steep slopes to graze on scurvy grass in Svalbard, Polar Res., 36, https://doi.org/10.1080/17518369.2017.1326453, 2017.
Sundset, M. A., Præsteng, K. E., Cann, I. K. O., Mathiesen, S. D., and MacKie, R. I.: Novel rumen bacterial diversity in two geographically separated sub-species of reindeer, Microb. Ecol., 54, 424–438, https://doi.org/10.1007/s00248-007-9254-x, 2007.
Thomas, F. A., Sinha, R. K., and Krishnan, K. P.: Bacterial community structure of a glacio-marine system in the Arctic (Ny-Ålesund, Svalbard), Sci. Total Environ., 718, 135264, https://doi.org/10.1016/j.scitotenv.2019.135264, 2020.
Ushida, K., Segawa, T., Tsuchida, S., and Murata, K.: Cecal bacterial communities in wild Japanese rock ptarmigans and captive Svalbard rock ptarmigans, J. Vet. Med. Sci., 78, 251–257, https://doi.org/10.1292/jvms.15-0313, 2016.
van den Bogert, B., Erkus, O., Boekhorst, J., de Goffau, M., Smid, E. J., Zoetendal, E. G., and Kleerebezem, M.: Diversity of human small intestinal Streptococcus and Veillonella populations, FEMS Microbiol. Ecol., 85, 376–388, https://doi.org/10.1111/1574-6941.12127, 2013.
Warwick, R. M., Emblow, C., Féral, J.-P., Hummel, H., van Avesaath, P., and Heip, C.: Report of the European Concerted Action: European Marine Biodiversity Research Sites. Report of the European Concerted Action: BIOMARE (Implementation and networking of large scale, long term marine biodiversity research in Europe), NIOO-CEME, https://researchportal.murdoch.edu.au/esploro/outputs/report/European-Marine-Biodiversity-Research-Sites-Report/991005544677807891 (last acccess: 27 March 2026), 2003.
Wejnerowski, Ł., Poniecka, E., Buda, J., Klimaszyk, P., Piasecka, A., Dziuba, M. K., Mugnai, G., Takeuchi, N., and Zawierucha, K.: Empirical testing of cryoconite granulation: Role of cyanobacteria in the formation of key biogenic structure darkening glaciers in polar regions, J. Phycol., 59, 939–949, https://doi.org/10.1111/jpy.13372, 2023.
Wesławski, J. M., Kwaśniewski, S., Stempniewicz, L., and Błachowiak-Samołyk, K.: Biodiversity and energy transfer to top trophic levels in two contrasting Arctic fjords, Polish Polar Res., 27, 259–278, 2006.
Yamane, K., Nambu, T., Yamanaka, T., Mashimo, C., Sugimori, C., Leung, K.-P., and Fukushima, H.: Complete Genome Sequence of Rothia mucilaginosa DY-18: A Clinical Isolate with Dense Meshwork-Like Structures from a Persistent Apical Periodontitis Lesion, Sequencing, 2010, 1–6, https://doi.org/10.1155/2010/457236, 2010.
Yang, G. L., Hou, S. G., Le Baoge, R., Li, Z. G., Xu, H., Liu, Y. P., Du, W. T., and Liu, Y. Q.: Differences in Bacterial Diversity and Communities between Glacial Snow and Glacial Soil on the Chongce Ice Cap, West Kunlun Mountains, Sci. Rep., 6, 1–8, https://doi.org/10.1038/srep36548, 2016.
Yao, H., Zhang, Z., Wu, N., Wang, M., Wu, Q., Wu, H., and Zhao, D.: Comparative analysis of intestinal flora at different overwintering periods in wild relict gulls (Larus relictus): first evidence from Northern China, Front. Microbiomes, 2, 1–11, https://doi.org/10.3389/frmbi.2023.1218281, 2023.
Yu, Y., Li, H., Zeng, Y., Sun, K., and Chen, B.: Pricia antarctica gen. nov., sp. nov., a member of the family Flavobacteriaceae, isolated from Antarctic intertidal sediment, Int. J. Syst. Evol. Microbiol., 62, 2218–2223, https://doi.org/10.1099/ijs.0.037515-0, 2012.
Zawierucha, K., Zmudczyńska-Skarbek, K., Kaczmarek, Ł., and Wojczulanis-Jakubas, K.: The influence of a seabird colony on abundance and species composition of water bears (Tardigrada) in Hornsund (Spitsbergen, Arctic), Polar Biol., 39, 713–723, https://doi.org/10.1007/s00300-015-1827-4, 2016.
Zawierucha, K., Zmudczyńska-Skarbek, K., Guil, N., and Bogdziewicz, M.: Seabirds modify trophic groups, while altitude promotes xeric-tolerant species of Tardigrada in the high Arctic tundra (Svalbard archipelago), Acta Oecologica, 98, 50–58, https://doi.org/10.1016/j.actao.2019.05.007, 2019.
Zeng, Y. X., Yan, M., Yu, Y., Li, H. R., He, J. F., Sun, K., and Zhang, F.: Diversity of bacteria in surface ice of Austre Lovénbreen glacier, Svalbard., Arch. Microbiol., 195, 313–322, https://doi.org/10.1007/s00203-013-0880-z, 2013.
Zhang, G., Bai, J., Tebbe, C. C., Zhao, Q., Jia, J., Wang, W., Wang, X., and Yu, L.: Salinity controls soil microbial community structure and function in coastal estuarine wetlands, Environ. Microbiol., 23, 1020–1037, https://doi.org/10.1111/1462-2920.15281, 2021.
Zielińska, S., Kidawa, D., Stempniewicz, L., Łos, M., and Łos, J. M.: New insights into the microbiota of the Svalbard reindeer Rangifer tarandus platyrhynchus, Front. Microbiol., 7, 1–9, https://doi.org/10.3389/fmicb.2016.00170, 2016a.
Zielińska, S., Kidawa, D., Stempniewicz, L., Łos, M., and Łos, J. M.: The arctic soil bacterial communities in the vicinity of a little auk colony, Front. Microbiol., 7, 1–9, https://doi.org/10.3389/fmicb.2016.01298, 2016b.
Zmudczyńska-Skarbek, K., Bokhorst, S., Convey, P., Gwiazdowicz, D. J., Skubała, P., Zawierucha, K., and Zwolicki, A.: The impact of marine vertebrates on polar terrestrial invertebrate communities, Polar Biol., 47, 805–820, https://doi.org/10.1007/s00300-023-03134-8, 2024.
Zwolicki, A., Zmudczyńska-Skarbek, K. M., Iliszko, L., and Stempniewicz, L.: Guano deposition and nutrient enrichment in the vicinity of planktivorous and piscivorous seabird colonies in Spitsbergen, Polar Biol., 36, 363–372, https://doi.org/10.1007/s00300-012-1265-5, 2013.
Zwolicki, A., Zmudczyńska-Skarbek, K., Richard, P., and Stempniewicz, L.: Importance of marine-derived nutrients supplied by planktivorous seabirds to high arctic tundra plant communities, PLoS One, 11, 1–16, https://doi.org/10.1371/journal.pone.0154950, 2016.